Abstract
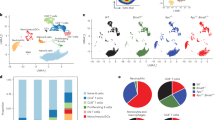
Although neutrophils have been linked to the formation of the pre-metastatic niche, the mechanism of their migration to distant, uninvolved tissues has remained elusive. We report that bone marrow neutrophils from mice with early-stage cancer exhibited much more spontaneous migration than that of control neutrophils from tumor-free mice. These cells lacked immunosuppressive activity but had elevated rates of oxidative phosphorylation and glycolysis, and increased production of ATP, relative to that of control neutrophils. Their enhanced spontaneous migration was mediated by autocrine ATP signaling through purinergic receptors. In ectopic tumor models and late stages of cancer, bone marrow neutrophils demonstrated potent immunosuppressive activity. However, these cells had metabolic and migratory activity indistinguishable from that of control neutrophils. A similar pattern of migration was observed for neutrophils and polymorphonuclear myeloid-derived suppressor cells from patients with cancer. These results elucidate the dynamic changes that neutrophils undergo in cancer and demonstrate the mechanism of neutrophils’ contribution to early tumor dissemination.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Fridlender, Z. G. et al. Polarization of tumor-associated neutrophil phenotype by TGF-β: “N1” versus “N2” TAN. Cancer Cell 16, 183–194 (2009).
Eruslanov, E. B. et al. Tumor-associated neutrophils stimulate T cell responses in early-stage human lung cancer. J. Clin. Invest. 124, 5466–5480 (2014).
Veglia, F., Perego, M. & Gabrilovich, D. Myeloid-derived suppressor cells coming of age. Nat. Immunol. 19, 108–119 (2018).
Coffelt, S. B., Wellenstein, M. D. & de Visser, K. E. Neutrophils in cancer: neutral no more. Nat. Rev. Cancer 16, 431–446 (2016).
Zhou, J., Nefedova, Y., Lei, A. & Gabrilovich, D. Neutrophils and PMN-MDSC: Their biological role and interaction with stromal cells. Semin. Immunol. 35, 19–28 (2018).
Ortiz, M. L. et al. Immature myeloid cells directly contribute to skin tumor development by recruiting IL-17-producing CD4+ T cells. J. Exp. Med. 212, 351–367 (2015).
Ortiz, M. L., Lu, L., Ramachandran, I. & Gabrilovich, D. I. Myeloid-derived suppressor cells in the development of lung cancer. Cancer Immunol. Res. 2, 50–58 (2014).
Bronte, V. et al. Recommendations for myeloid-derived suppressor cell nomenclature and characterization standards. Nat. Commun. 7, 12150 (2016).
Seyfried, T. N. & Huysentruyt, L. C. On the origin of cancer metastasis. Crit. Rev. Oncog. 18, 43–73 (2013).
Achberger, S. et al. Circulating immune cell and microRNA in patients with uveal melanoma developing metastatic disease. Mol. Immunol. 58, 182–186 (2014).
Weide, B. et al. Myeloid-derived suppressor cells predict survival of patients with advanced melanoma: comparison with regulatory T cells and NY-ESO-1- or melan-A-specific T cells. Clin. Cancer Res. 20, 1601–1609 (2014).
Condamine, T., Ramachandran, I., Youn, J. I. & Gabrilovich, D. I. Regulation of tumor metastasis by myeloid-derived suppressor cells. Annu. Rev. Med. 66, 97–110 (2015).
Coffelt, S. B. et al. IL-17-producing γδ T cells and neutrophils conspire to promote breast cancer metastasis. Nature 522, 345–348 (2015).
Toh, B. et al. Mesenchymal transition and dissemination of cancer cells is driven by myeloid-derived suppressor cells infiltrating the primary tumor. PLoS Biol. 9, e1001162 (2011).
Liu, Y. et al. MicroRNA-494 is required for the accumulation and functions of tumor-expanded myeloid-derived suppressor cells via targeting of PTEN. J. Immunol. 188, 5500–5510 (2012).
Yang, L. et al. Abrogation of TGF β signaling in mammary carcinomas recruits Gr-1+CD11b+ myeloid cells that promote metastasis. Cancer Cell 13, 23–35 (2008).
Huh, S. J., Liang, S., Sharma, A., Dong, C. & Robertson, G. P. Transiently entrapped circulating tumor cells interact with neutrophils to facilitate lung metastasis development. Cancer Res. 70, 6071–6082 (2010).
Ichikawa, M., Williams, R., Wang, L., Vogl, T. & Srikrishna, G. S100A8/A9 activate key genes and pathways in colon tumor progression. Mol. Cancer Res. 9, 133–148 (2011).
Kowanetz, M. et al. Granulocyte-colony stimulating factor promotes lung metastasis through mobilization of Ly6G+Ly6C+ granulocytes. Proc. Natl. Acad. Sci. USA 107, 21248–21255 (2010).
Kolaczkowska, E. & Kubes, P. Neutrophil recruitment and function in health and inflammation. Nat. Rev. Immunol. 13, 159–175 (2013).
Mócsai, A., Walzog, B. & Lowell, C. A. Intracellular signalling during neutrophil recruitment. Cardiovasc. Res. 107, 373–385 (2015).
Kumar, V., Patel, S., Tcyganov, E. & Gabrilovich, D. I. The nature of myeloid-derived suppressor cells in the tumor microenvironment. Trends Immunol. 37, 208–220 (2016).
Brandau, S. et al. Myeloid-derived suppressor cells in the peripheral blood of cancer patients contain a subset of immature neutrophils with impaired migratory properties. J. Leukoc. Biol. 89, 311–317 (2011).
Kato, M. et al. Transgenic mouse model for skin malignant melanoma. Oncogene 17, 1885–1888 (1998).
Hingorani, S. R. et al. Trp53R172H and KrasG12D cooperate to promote chromosomal instability and widely metastatic pancreatic ductal adenocarcinoma in mice. Cancer Cell 7, 469–483 (2005).
Greenberg, N. M. et al. Prostate cancer in a transgenic mouse. Proc. Natl. Acad. Sci. USA 92, 3439–3443 (1995).
Evans, R.A. et al. Lack of immunoediting in murine pancreatic cancer reversed with neoantigen. JCI Insight 1, e88328 (2016).
Casbon, A. J. et al. Invasive breast cancer reprograms early myeloid differentiation in the bone marrow to generate immunosuppressive neutrophils. Proc. Natl. Acad. Sci. USA 112, E566–E575 (2015).
Biswas, S. K. Metabolic Reprogramming of immune cells in cancer progression. Immunity 43, 435–449 (2015).
Chen, Y. et al. ATP release guides neutrophil chemotaxis via P2Y2 and A3 receptors. Science 314, 1792–1795 (2006).
Chen, Y. et al. Purinergic signaling: a fundamental mechanism in neutrophil activation. Sci. Signal. 3, ra45 (2010).
Junger, W. G. Purinergic regulation of neutrophil chemotaxis. Cell. Mol. Life Sci. 65, 2528–2540 (2008).
Junger, W. G. Immune cell regulation by autocrine purinergic signalling. Nat. Rev. Immunol. 11, 201–212 (2011).
Lecut, C. et al. P2X1 ion channels promote neutrophil chemotaxis through Rho kinase activation. J. Immunol. 183, 2801–2809 (2009).
Hind, L. E., Vincent, W. J. & Huttenlocher, A. Leading from the back: the role of the uropod in neutrophil polarization and migration. Dev. Cell 38, 161–169 (2016).
Yang, H. W., Collins, S. R. & Meyer, T. Locally excitable Cdc42 signals steer cells during chemotaxis. Nat. Cell Biol. 18, 191–201 (2016).
Condamine, T. et al. Lectin-type oxidized LDL receptor-1 distinguishes population of human polymorphonuclear myeloid-derived suppressor cells in cancer patients. Sci. Immunol. 1, aaf8943 (2016).
Condamine, T. et al. ER stress regulates myeloid-derived suppressor cell fate through TRAIL-R-mediated apoptosis. J. Clin. Invest. 124, 2626–2639 (2014).
Tybulewicz, V. L. & Henderson, R. B. Rho family GTPases and their regulators in lymphocytes. Nat. Rev. Immunol. 9, 630–644 (2009).
Hammami, I. et al. Immunosuppressive activity enhances central carbon metabolism and bioenergetics in myeloid-derived suppressor cells in vitro models. BMC Cell Biol. 13, 18 (2012).
Hossain, F. et al. Inhibition of fatty acid oxidation modulates immunosuppressive functions of myeloid-derived suppressor cells and enhances cancer therapies. Cancer Immunol. Res. 3, 1236–1247 (2015).
Schug, Z. T. et al. Acetyl-CoA synthetase 2 promotes acetate utilization and maintains cancer cell growth under metabolic stress. Cancer Cell 27, 57–71 (2015).
Li, B. & Dewey, C. N. RSEM: accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinformatics 12, 323 (2011).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Othmer, H. G., Dunbar, S. R. & Alt, W. Models of dispersal in biological systems. J. Math. Biol. 26, 263–298 (1988).
Dunn, G. A. Characterising a kinesis response: time averaged measures of cell speed and directional persistence. Agents Actions Suppl. 12, 14–33 (1983).
Farrell, B. E., Daniele, R. P. & Lauffenburger, D. A. Quantitative relationships between single-cell and cell-population model parameters for chemosensory migration responses of alveolar macrophages to C5a. Cell Motil. Cytoskeleton 16, 279–293 (1990).
Acknowledgements
We thank R. Ramakrishnan (H. Lee Moffitt Cancer Center) for the luciferase expressing LL2 tumor cell line; and D. Laxminarasimha for help in animal experiments. This work was supported by the Wistar Institute Animal and Bioinformatics core facilities and by the US National Institutes of Health (P01 CA140043 and T32 CA09171). S.Y. was supported by International Program for Ph.D Candidates, Sun Yat-Sen University, China.
Author information
Authors and Affiliations
Contributions
Investigation, S.P., S.F., J.M., G.D., A.P., C.L., K.A.-T. and M.S.; formal analysis, A.K. and Z.S.; resources, Y.N., L.R.L., C.C., R.H.V., C.M., B.N., N.H., G.M. and M.G.; writing (original draft), S.P. and G.D.; writing (review and editing), J.Z., D.C.A. and D.I.G.; funding acquisition, D.C.A. and D.I.G.; conceptualization and supervision, D.I.G.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figures 1–8 and Supplementary Tables 1 and 2
Supplementary Video 1
Time lapse video demonstrating spontaneous movement of neutrophils from naïve and RET TB mice. Scale bar = 50 µm
Rights and permissions
About this article
Cite this article
Patel, S., Fu, S., Mastio, J. et al. Unique pattern of neutrophil migration and function during tumor progression. Nat Immunol 19, 1236–1247 (2018). https://doi.org/10.1038/s41590-018-0229-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41590-018-0229-5
This article is cited by
-
Multi-model analysis of gallbladder cancer reveals the role of OxLDL-absorbing neutrophils in promoting liver invasion
Experimental Hematology & Oncology (2024)
-
Understanding the immunosuppressive microenvironment of glioma: mechanistic insights and clinical perspectives
Journal of Hematology & Oncology (2024)
-
Incorporating the inflammation-related parameters enhances the performance of the nomogram for predicting local control in lung cancer patients treated with stereotactic body radiation therapy
Journal of Cancer Research and Clinical Oncology (2024)
-
Single-nucleus RNA sequencing and deep tissue proteomics reveal distinct tumour microenvironment in stage-I and II cervical cancer
Journal of Experimental & Clinical Cancer Research (2023)
-
The role and metabolic adaptations of neutrophils in premetastatic niches
Biomarker Research (2023)