Abstract
Nucleoside analogs have broad clinical utility as antiviral drugs. Key to their systemic distribution and cellular entry are human nucleoside transporters. Here, we establish that the human concentrative nucleoside transporter 3 (CNT3) interacts with antiviral drugs used in the treatment of coronavirus infections. We report high-resolution single-particle cryo-electron microscopy structures of bovine CNT3 complexed with antiviral nucleosides N4-hydroxycytidine, PSI-6206, GS-441524 and ribavirin, all in inward-facing states. Notably, we found that the orally bioavailable antiviral molnupiravir arrests CNT3 in four distinct conformations, allowing us to capture cryo-electron microscopy structures of drug-loaded outward-facing and drug-loaded intermediate states. Our studies uncover the conformational trajectory of CNT3 during membrane transport of a nucleoside analog antiviral drug, yield new insights into the role of interactions between the transport and the scaffold domains in elevator-like domain movements during drug translocation, and provide insights into the design of nucleoside analog antiviral prodrugs with improved oral bioavailability.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Atomic coordinates have been deposited in the PDB with the following ID numbers: 8TZ2 (apo), 8TZ5 (NHC), 8TZ6 (PSI-6206), 8TZ1 (ribavirin), 8TZ3 (GS4 consensus), 8TZ4 (GS4 subset), 8TZD (INT1–INT1–OFS; MPV condition 1), 8TZ7 (INT1 homotrimer; MPV condition 1 ensemble), 8TZ8 (INT1–INT1–INT3; MPV condition 1 ensemble), 8TZA (INT1–INT1–INT2; MPV condition 2 ensemble) and 8TZ9 (INT2 homotrimer; MPV condition 2 ensemble). The reconstructed cryo-EM maps have been deposited in the Electron Microscopy Data Bank with the following ID numbers: EMD-41731 (apo), EMD-41734 (NHC), EMD-41735 (PSI-6206), EMD-41730 (ribavirin), EMD-41732 (GS4 consensus), EMD-41733 (GS4 subset), EMD-41755 (INT1–INT1–OFS; MPV condition 1), EMD-41736 (INT1 homotrimer; MPV condition 1 ensemble), EMD-41737 (INT1–INT1–INT3; MPV condition 1 ensemble), EMD-41738 (INT2 homotrimer; MPV condition 2 ensemble), EMD-41739 (INT1–INT1–INT2; MPV condition 2 ensemble), EMD-41740 (consensus; MPV condition 1), EMD-41752 (consensus; MPV condition 2), EMD-41747 (INT3 homotrimer; MPV condition 1 ensemble), EMD-41746 (INT3–INT3–INT1; MPV condition 1 ensemble), EMD-41745 (INT3–INT3–OFS; MPV condition 1 ensemble), EMD-41744 (INT1–INT1–OFS; MPV condition 1 ensemble), EMD-41741 (OFS–OFS–OFS; MPV condition 1 ensemble), EMD-41742 (OFS–OFS–INT1; MPV condition 1 ensemble), EMD-41743 (OFS–OFS–INT3; MPV condition 1 ensemble), EMD-41748 (OFS–INT1–INT3 clockwise; MPV condition 1 ensemble), EMD-41749 (OFS–INT1–INT3 counterclockwise; MPV condition 1 ensemble), EMD-41751 (INT1 homotrimer; MPV condition 2 ensemble) and EMD-41750 (INT1–INT2–INT2; MPV condition 2 ensemble). Source data are provided with this paper. Additional data relevant to this paper are available upon reasonable request to S.-Y. L.
References
Yates, M. K. & Seley-Radtke, K. L. The evolution of antiviral nucleoside analogues: a review for chemists and non-chemists. Part II: complex modifications to the nucleoside scaffold. Antivir. Res. 162, 5–21 (2019).
Seley-Radtke, K. L. & Yates, M. K. The evolution of nucleoside analogue antivirals: a review for chemists and non-chemists. Part 1: early structural modifications to the nucleoside scaffold. Antivir. Res. 154, 66–86 (2018).
Pruijssers, A. J. & Denison, M. R. Nucleoside analogues for the treatment of coronavirus infections. Curr. Opin. Virol. 35, 57–62 (2019).
Graci, J. D. & Cameron, C. E. Mechanisms of action of ribavirin against distinct viruses. Rev. Med. Virol. 16, 37–48 (2006).
Beigel, J. H. et al. Remdesivir for the treatment of Covid-19—final report. N. Engl. J. Med. 383, 1813–1826 (2020).
Lawitz, E. et al. Sofosbuvir for previously untreated chronic hepatitis C infection. N. Engl. J. Med. 368, 1878–1887 (2013).
Jayk Bernal, A. et al. Molnupiravir for oral treatment of Covid-19 in nonhospitalized patients. N. Engl. J. Med. 386, 509–520 (2022).
Mehellou, Y., Rattan, H. S. & Balzarini, J. The ProTide prodrug technology: from the concept to the clinic. J. Med. Chem. 61, 2211–2226 (2018).
Warren, T. K. et al. Therapeutic efficacy of the small molecule GS-5734 against Ebola virus in rhesus monkeys. Nature 531, 381–385 (2016).
Sheahan, T. P. et al. An orally bioavailable broad-spectrum antiviral inhibits SARS-CoV-2 in human airway epithelial cell cultures and multiple coronaviruses in mice. Sci. Transl. Med. 12, eabb5883 (2020).
Yan, V. C. & Muller, F. L. Advantages of the parent nucleoside GS-441524 over remdesivir for Covid-19 treatment. ACS Med. Chem. Lett. 11, 1361–1366 (2020).
Williamson, B. N. et al. Clinical benefit of remdesivir in rhesus macaques infected with SARS-CoV-2. Nature 585, 273–276 (2020).
Feld, J. J. & Hoofnagle, J. H. Mechanism of action of interferon and ribavirin in treatment of hepatitis C. Nature 436, 967–972 (2005).
Murphy, B. G. et al. The nucleoside analog GS-441524 strongly inhibits feline infectious peritonitis (FIP) virus in tissue culture and experimental cat infection studies. Vet. Microbiol. 219, 226–233 (2018).
Agostini, M. L. et al. Small-molecule antiviral β-d-N4-hydroxycytidine inhibits a proofreading-intact coronavirus with a high genetic barrier to resistance. J. Virol. 93, e01348-19 (2019).
Toots, M. et al. Characterization of orally efficacious influenza drug with high resistance barrier in ferrets and human airway epithelia. Sci. Transl. Med. 11, eaax5866 (2019).
Yoon, J. J. et al. Orally efficacious broad-spectrum ribonucleoside analog inhibitor of influenza and respiratory syncytial viruses. Antimicrob. Agents Chemother. 62, e00766-18 (2018).
Hirschi, M., Johnson, Z. L. & Lee, S. Y. Visualizing multistep elevator-like transitions of a nucleoside transporter. Nature 545, 66–70 (2017).
Johnson, Z. L. et al. Structural basis of nucleoside and nucleoside drug selectivity by concentrative nucleoside transporters. eLife 3, e03604 (2014).
Johnson, Z. L., Cheong, C. G. & Lee, S. Y. Crystal structure of a concentrative nucleoside transporter from Vibrio cholerae at 2.4 Å. Nature 483, 489–493 (2012).
Zhou, Y. et al. Cryo-EM structure of the human concentrative nucleoside transporter CNT3. PLoS Biol. 18, e3000790 (2020).
Young, J. D., Yao, S. Y., Baldwin, J. M., Cass, C. E. & Baldwin, S. A. The human concentrative and equilibrative nucleoside transporter families, SLC28 and SLC29. Mol. Asp. Med. 34, 529–547 (2013).
Wright, N. J. & Lee, S. Y. Structures of human ENT1 in complex with adenosine reuptake inhibitors. Nat. Struct. Mol. Biol. 26, 599–606 (2019).
Wang, C. et al. Structural basis of the substrate recognition and inhibition mechanism of Plasmodium falciparum nucleoside transporter PfENT1. Nat. Commun. 14, 1727 (2023).
Tsubota, A. et al. Contribution of ribavirin transporter gene polymorphism to treatment response in peginterferon plus ribavirin therapy for HCV genotype 1b patients. Liver Int. 32, 826–836 (2012).
Doehring, A. et al. Role of nucleoside transporters SLC28A2/3 and SLC29A1/2 genetics in ribavirin therapy: protection against anemia in patients with chronic hepatitis C. Pharmacogenet. Genomics 21, 289–296 (2011).
Rau, M. et al. Impact of genetic SLC28 transporter and ITPA variants on ribavirin serum level, hemoglobin drop and therapeutic response in patients with HCV infection. J. Hepatol. 58, 669–675 (2013).
Iikura, M. et al. ENT1, a ribavirin transporter, plays a pivotal role in antiviral efficacy of ribavirin in a hepatitis C virus replication cell system. Antimicrob. Agents Chemother. 56, 1407–1413 (2012).
Morello, J. et al. Influence of a single nucleotide polymorphism at the main ribavirin transporter gene on the rapid virological response to pegylated interferon–ribavirin therapy in patients with chronic hepatitis C virus infection. J. Infect. Dis. 202, 1185–1191 (2010).
Rasmussen, H. B. et al. Cellular uptake and intracellular phosphorylation of GS-441524: implications for its effectiveness against COVID-19. Viruses 13, 1369 (2021).
Wang, A. Q. et al. Preclinical pharmacokinetics and in vitro properties of GS-441524, a potential oral drug candidate for COVID-19 treatment. Front. Pharmacol. 13, 918083 (2022).
Wright, N. J. & Lee, S. Y. Toward a molecular basis of cellular nucleoside transport in humans. Chem. Rev. 121, 5336–5358 (2021).
Smith, K. M. et al. Cation coupling properties of human concentrative nucleoside transporters hCNT1, hCNT2 and hCNT3. Mol. Membr. Biol. 24, 53–64 (2007).
Hamilton, S. R. et al. Subcellular distribution and membrane topology of the mammalian concentrative Na+–nucleoside cotransporter rCNT1. J. Biol. Chem. 276, 27981–27988 (2001).
Fernandez-Calotti, P., Casulleras, O., Antolin, M., Guarner, F. & Pastor-Anglada, M. Galectin-4 interacts with the drug transporter human concentrative nucleoside transporter 3 to regulate its function. FASEB J. 30, 544–554 (2016).
Damaraju, V. L. et al. Influence of sugar ring conformation on the transportability of nucleosides by human nucleoside transporters. ChemBioChem 12, 2774–2778 (2011).
Arimany-Nardi, C. et al. Identification and characterization of a secondary sodium-binding site and the main selectivity determinants in the human concentrative nucleoside transporter 3. Mol. Pharm. 14, 1980–1987 (2017).
Gucwa, M. et al. CMM—an enhanced platform for interactive validation of metal binding sites. Protein Sci. 32, e4525 (2023).
Errasti-Murugarren, E., Molina-Arcas, M., Casado, F. J. & Pastor-Anglada, M. The human concentrative nucleoside transporter-3 C602R variant shows impaired sorting to lipid rafts and altered specificity for nucleoside-derived drugs. Mol. Pharmacol. 78, 157–165 (2010).
Zhang, J. et al. Uridine binding and transportability determinants of human concentrative nucleoside transporters. Mol. Pharmacol. 68, 830–839 (2005).
Cao, L. et al. The adenosine analog prodrug ATV006 is orally bioavailable and has preclinical efficacy against parental SARS-CoV-2 and variants. Sci. Transl. Med. 14, eabm7621 (2022).
Schafer, A. et al. Therapeutic treatment with an oral prodrug of the remdesivir parental nucleoside is protective against SARS-CoV-2 pathogenesis in mice. Sci. Transl. Med. 14, eabm3410 (2022).
NCT05603143: Study of obeldesivir in participants with COVID-19 who have a high risk of developing serious or severe illness (BIRCH). https://clinicaltrials.gov/show/NCT05603143 (2022).
Qiu, B., Matthies, D., Fortea, E., Yu, Z. & Boudker, O. Cryo-EM structures of excitatory amino acid transporter 3 visualize coupled substrate, sodium, and proton binding and transport. Sci. Adv. 7, eabf5814 (2021).
Erkens, G. B., Hänelt, I., Goudsmits, J. M., Slotboom, D. J. & van Oijen, A. M. Unsynchronised subunit motion in single trimeric sodium-coupled aspartate transporters. Nature 502, 119–123 (2013).
Georgieva, E. R., Borbat, P. P., Ginter, C., Freed, J. H. & Boudker, O. Conformational ensemble of the sodium-coupled aspartate transporter. Nat. Struct. Mol. Biol. 20, 215–221 (2013).
Arkhipova, V., Guskov, A. & Slotboom, D. J. Structural ensemble of a glutamate transporter homologue in lipid nanodisc environment. Nat. Commun. 11, 998 (2020).
Garaeva, A. A. & Slotboom, D. J. Elevator-type mechanisms of membrane transport. Biochem. Soc. Trans. 48, 1227–1241 (2020).
Drew, D., North, R. A., Nagarathinam, K. & Tanabe, M. Structures and general transport mechanisms by the major facilitator superfamily (MFS). Chem. Rev. 121, 5289–5335 (2021).
Qureshi, A. A. et al. The molecular basis for sugar import in malaria parasites. Nature 578, 321–325 (2020).
Wright, N. J. et al. Methotrexate recognition by the human reduced folate carrier SLC19A1. Nature 609, 1056–1062 (2022).
Kimanius, D., Dong, L., Sharov, G., Nakane, T. & Scheres, S. H. W. New tools for automated cryo-EM single-particle analysis in RELION-4.0. Biochem. J. 478, 4169–4185 (2021).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Liebschner, D. et al. Macromolecular structure determination using X-rays, neutrons and electrons: recent developments in Phenix. Acta Crystallogr. D Struct. Biol. 75, 861–877 (2019).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Williams, C. J. et al. MolProbity: More and better reference data for improved all‐atom structure validation. Protein Sci. 27, 293–315 (2018).
Pettersen, E. F. et al. UCSF ChimeraX: structure visualization for researchers, educators, and developers. Protein Sci. 30, 70–82 (2021).
Wu, E. L. et al. CHARMM—GUI membrane builder toward realistic biological membrane simulations. J. Comput. Chem. 35, 1997–2004 (2014).
Suh, D. et al. CHARMM—GUI enhanced sampler for various collective variables and enhanced sampling methods. Protein Sci. 31, e4446 (2022).
Huang, J. et al. CHARMM36m: an improved force field for folded and intrinsically disordered proteins. Nat. Methods 14, 71–73 (2017).
Jo, S., Kim, T. & Im, W. Automated builder and database of protein/membrane complexes for molecular dynamics simulations. PLoS ONE 2, e880 (2007).
Eastman, P. et al. OpenMM 7: rapid development of high performance algorithms for molecular dynamics. PLoS Comput. Biol. 13, e1005659 (2017).
Acknowledgements
Cryo-EM data were screened and collected at the Duke University SMIF, the National Cancer Institute’s NCEF at the Frederick National Laboratory for Cancer Research, the Pacific Northwest Center for Cryo-EM (PNCC) at Oregon Health and Science University and the National Institute of Environmental Health Sciences. We thank J. Myers at PNCC, A. Wier, T. Fox and U. Baxa at NCEF and N. Bhattacharya at SMIF for assistance with microscope operation. We thank M. Hirschi for initial biochemistry of bCNT3 and K. Tsolova for help with part of the radioactive tracer uptake assay and manuscript reading. This research was supported by a National Institutes of Health grant R21AI166134 (S.-Y.L.) and the National Institute of Health Intramural Research Program, the US National Institutes of Environmental Health Science (ZIC ES103326 to M.J.B.) and the National Science Foundation (MCB-2111728 to W.I.). A portion of this research was supported by National Institutes of Health grant U24GM129547, was performed at the PNCC at Oregon Health and Science University and was accessed through EMSL (grid.436923.9), a Department of Energy Office of Science User Facility sponsored by the Office of Biological and Environmental Research. The Duke University SMIF is affiliated with the North Carolina Research Triangle Nanotechnology Network, which is, in part, supported by the National Science Foundation (ECCS-2025064).
Author information
Authors and Affiliations
Contributions
N.J.W. conducted biochemical preparation, sample freezing, single-particle 3D reconstruction, model building and radiotracer uptake assays. F.Z. performed electrophysiological recordings. Y.S. collected data and performed part of structural analysis. Y.Y. performed biochemical characterization of bCNT3, and J.F. performed part of the radiotracer uptake assay, all under the guidance of S.-Y.L. L.K. performed all MD simulations under the guidance of W.I. K.S. performed part of the cryo-EM sample screening under the guidance of M.J.B. N.J.W. and S.-Y.L. wrote the paper with input from the rest of authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Chemical Biology thanks Raimund Dutzler and the other, anonymous, reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Functional characterization, protein biochemistry, and cryo-EM sample preparation of bCNT3.
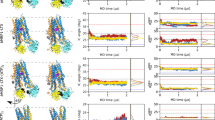
a, b, c, Time-dependent uptake of 1.0 µM [3H]-ribavirin at room temperature is linear within 30 min for hCNT1 (a, background corrected uptake rate of 0.0036 ± 0.0002 pmole/min), hCNT2 (b, background corrected uptake rate of 0.41 ± 0.01 pmole/min), hCNT3 (b, background corrected uptake rate of 0.87 ± 0.04 pmole/min), hENT1 (c, background corrected uptake rate of 0.056 ± 0.004 pmole/min) and hENT2 (c, background corrected uptake rate of 0.098 ± 0.007 pmole/min). Data shown in panels a, b, c are from the same series of experiments (n = 3 biological replicates, individual replicates shown) with the water-injected control measurements re-shown in each figure panel to convey signal-to-background for each transporter subtype. d, Cold-competition of WT hCNT3 or WT bCNT3 mediated [3H]-ribavirin uptake by cold NHC or GS4. (ChoCl – sodium-free negative control condition with 96 mM choline chloride; 15-minute uptake with 0.1 μM [3H]-ribavirin; n = 3 biological replicates with individual replicates and mean ± s.e.m. shown). e, Representative size-exclusion chromatography profile and corresponding SDS-PAGE analysis of purified nanodisc reconstituted bCNT3. Fractions pooled for cryo-EM analysis indicated with an asterisk (*). Purification and nanodisc reconstitution of bCNT3 was repeated routinely with similar results for each cryo-EM structure reported in this study. f, Representative cryo-EM micrograph (top) and 2D class averages (bottom) of nanodisc reconstituted bCNT3. Micrographs of similar quality were recorded for each cryo-EM dataset.
Extended Data Fig. 2 Cryo-EM image processing of drug-bound and drug-free IFS states of bCNT3.
a, Cryo-EM image processing procedures. b, Fourier-shell correlation plots (top) and angular orientation distributions (bottom) from the final reconstructions. c, Local resolution of the final reconstructions (local resolution determined in cryoSPARC).
Extended Data Fig. 3 Local cryo-EM map quality.
Local cryo-EM maps for secondary structure segments. From top to bottom are NHC, PSI-6206, Ribavirin, GS441524 (subset), GS441524 (consensus) and apo, respectively. Cryo-EM densities are shown at map threshold values of 6σ for NHC, PSI-6206, ribavirin, GS4 sub., apo, or 7σ for GS4 cons.
Extended Data Fig. 4 Sodium ion coordination in the inward-facing state.
a, Location of the two sodium ion binding sites relative to the nucleoside binding site within the NHC bound IFS bCNT3 structure. b, Architecture and coordination geometries of the two sodium ion binding sites in the 2.31 Å cryo-EM map (map thresholds used denoted in panel insets).
Extended Data Fig. 5 TEVC studies of molnupiravir interaction with CNT3.
a, TEVC recordings of inward currents elicited by application of 100 μM uridine, or current block by co-application of 100 μM uridine + 1 mM MPV in WT hCNT3 expressing oocytes (representative trace at left, summary of peak currents from n = 6 biological replicates at right with individual measurements and mean ± s.e.m. shown; values are baseline corrected to leak current per oocyte). b, TEVC recordings of inward currents elicited by application of 100 μM uridine, or current block by co-application of 100 μM uridine + 1 mM MPV in WT bCNT3 expressing oocytes (representative trace at left, summary of peak currents from n = 4 biological replicates at right with individual measurements and mean ± s.e.m. shown; values are baseline corrected to leak current per oocyte).
Extended Data Fig. 6 Cryo-EM image processing of molnupiravir-bound OFS, INT1, INT2, INT3 conformational states of bCNT3.
a, Initial cryo-EM image processing procedure for the ‘MPV condition 1’ dataset. b, Cryo-EM ensemble analysis of ‘MPV condition 1’. Final Fourier shell correlation plots and angular orientation distribution plots shown next to every reconstruction obtained.
Extended Data Fig. 7 Cryo-EM image processing of molnupiravir-bound INT1 and INT2 conformational states of bCNT3.
a, Initial cryo-EM image processing procedure for the ‘MPV condition 2’ dataset. b, Cryo-EM ensemble analysis of ‘MPV condition 2’. Final Fourier shell correlation plots and angular orientation distribution plots shown next to every reconstruction obtained.
Extended Data Fig. 8 MPV ligand densities in representative OFS, INT1, INT2, and INT3 cryo-EM reconstructions.
Cryo-EM densities around the nucleoside binding site from representative reconstructions containing OFS, INT1, INT2, or INT3 protomers (reconstructions used: OFS-INT1-INT1 highest res; INT1-INT1-INT1 ‘condition 1’ ensemble analysis; INT2-INT2-INT2 ‘condition 2’ ensemble analysis; INT1-INT1-INT3 ‘condition 1’ ensemble analysis). Sharpened maps shown (threshold = 6σ).
Extended Data Fig. 9 Comparison of conformational trajectories in MPV-loaded bCNT3 and apo nwCNT.
a, Structural superposition of bCNT3 (top left) and nwCNT (bottom left) conformers. Alignment based on the trimerization interface, TM3 and TM6. Relative positions of HP1 (middle) and HP2 (right), with lines denoting a marker residue for each conformational state. b, Transport domain movement quantified by its tilt angle relative to a representative IFS structure (X-axis) and distance displacement of nucleoside binding site center of mass (Y-axis). For bCNT3, coordinates corresponding to the highest resolution instance of each conformer used for the analysis (OFS-INT1-INT1 highest res; INT1-INT1-INT1 ‘condition 1’ ensemble analysis; INT2-INT2-INT2 ‘condition 2’ ensemble analysis; INT1-INT1-INT3 ‘condition 1’ ensemble analysis; all relative to the GS4 consensus IFS structure). c, Structural superposition of the transport domain for OFS, INT1, INT3, INT3 and IFS (NHC) structures. d, [3H]-ribavirin uptake (1.0 μM) in 10 minutes into oocytes expressing WT or mutant bCNT3 (n = 3; mean ± s.e.m. and individual replicates shown).
Extended Data Fig. 10 Map-to-model analysis for reconstructions obtained from the cryo-EM ensemble analysis.
a, Map-to-model of every trimer reconstruction obtained from MPV ‘condition 1’ with the ensemble analysis. Map-to-model correlation analysis shown below each reconstruction (for details see Methods). b, Map to model of every trimer reconstruction obtained from MPV ‘condition 2’ with the ensemble analysis. Map-model correlation analysis shown below each reconstruction (for details see Methods).
Supplementary information
Supplementary Information
Supplementary Figs. 1–8 and Tables 1–5.
Source data
Source Data Fig. 1
Source data.
Source Data Fig. 4
Source data.
Source Data Fig. 5
Source data.
Source Data Extended Data Fig. 1
Source data.
Source Data Extended Data Fig. 1e
Uncropped SDS–PAGE image.
Source Data Extended Data Fig. 5
Source data.
Source Data Extended Data Fig. 9
Source data.
Source Data Extended Data Fig. 10
Source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wright, N.J., Zhang, F., Suo, Y. et al. Antiviral drug recognition and elevator-type transport motions of CNT3. Nat Chem Biol (2024). https://doi.org/10.1038/s41589-024-01559-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41589-024-01559-8