Abstract
Glycolysis is a universal metabolic process that breaks down glucose to produce adenosine triphosphate (ATP) and biomass precursors. The Entner–Doudoroff (ED) pathway is a glycolytic pathway that parallels textbook glycolysis but yields half as much ATP. Accordingly, in organisms that possess both glycolytic pathways (for example, Escherichia coli), its raison dʼêtre remains a mystery. In this study, we found that the ED pathway provides a selective advantage during growth acceleration. Upon carbon and nitrogen upshifts, E. coli accelerates growth faster with than without the ED pathway. Concurrent isotope tracing reveals that the ED pathway flux increases faster than that of textbook glycolysis. We attribute the fast response time of the ED pathway to its strong thermodynamic driving force and streamlining of glucose import. Intermittent nutrient supply manifests the evolutionary advantage of the parallel glycolysis; thus, the dynamic nature of an ostensibly redundant pathway’s role in promoting rapid adaptation constitutes a metabolic design principle.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Source data for Figs. 1–6 are provided in Supplementary Tables 1–17 and the GitHub public repository: https://github.com/richardlaw517/Parallel_Glycolysis.
LTEE source data are available at https://barricklab.org/shiny/LTEE-Ecoli/. Source data are provided with this paper.
Code availability
The code for the analysis of metabolic fluxes and LTEE is available on the GitHub public repository: https://github.com/richardlaw517/Parallel_Glycolysis.
References
Basan, M. et al. A universal trade-off between growth and lag in fluctuating environments. Nature 584, 470–474 (2020).
Romano, A. H. & Conway, T. Evolution of carbohydrate metabolic pathways. Res. Microbiol. 147, 448–455 (1996).
Bar-Even, A., Flamholz, A., Noor, E. & Milo, R. Rethinking glycolysis: on the biochemical logic of metabolic pathways. Nat. Chem. Biol. 8, 509–517 (2012).
Park, J. O. et al. Near-equilibrium glycolysis supports metabolic homeostasis and energy yield. Nat. Chem. Biol. 15, 1001–1008 (2019).
Xiong, W. et al. Phosphoketolase pathway contributes to carbon metabolism in cyanobacteria. Nat. Plants 2, 15187 (2015).
Chen, X. et al. The Entner–Doudoroff pathway is an overlooked glycolytic route in cyanobacteria and plants. Proc. Natl Acad. Sci. USA 113, 5441–5446 (2016).
Flamholz, A., Noor, E., Bar-Even, A., Liebermeister, W. & Milo, R. Glycolytic strategy as a tradeoff between energy yield and protein cost. Proc. Natl Acad. Sci. USA 110, 10039–10044 (2013).
Basan, M. et al. Overflow metabolism in Escherichia coli results from efficient proteome allocation. Nature 528, 99–104 (2015).
Noor, E. et al. The protein cost of metabolic fluxes: prediction from enzymatic rate laws and cost minimization. PLoS Comput. Biol. 12, e1005167 (2016).
Peekhaus, N. & Conway, T. What’s for dinner?: Entner–Doudoroff metabolism in Escherichia coli. J. Bacteriol. 180, 3495–3502 (1998).
Mahadevan, R. & Lovley, D. R. The degree of redundancy in metabolic genes is linked to mode of metabolism. Biophys. J. 94, 1216–1220 (2008).
Güell, O., Sagués, F. & Serrano, M. Á. Essential plasticity and redundancy of metabolism unveiled by synthetic lethality analysis. PLoS Comput. Biol. 10, e1003637 (2014).
Nowak, M. A., Boerlijst, M. C., Cooke, J. & Smith, J. M. Evolution of genetic redundancy. Nature 388, 167–171 (1997).
Murray, E. L. & Conway, T. Multiple regulators control expression of the Entner–Doudoroff aldolase (Eda) of Escherichia coli. J. Bacteriol. 187, 991–1000 (2005).
Parisutham, V. & Lee, S. K. Novel functions and regulation of cryptic cellobiose operons in Escherichia coli. PLoS ONE 10, e0131928 (2015).
Grüning, N. M. & Ralser, M. Glycolysis: how a 300yr long research journey that started with the desire to improve alcoholic beverages kept revolutionizing biochemistry. Curr. Opin. Syst. Biol. 28, 100380 (2021).
Feist, A. M. et al. A genome-scale metabolic reconstruction for Escherichia coli K-12 MG1655 that accounts for 1260 ORFs and thermodynamic information. Mol. Syst. Biol. 3, 121 (2007).
Fuhrman, L. K., Wanken, A., Nickerson, K. W. & Conway, T. Rapid accumulation of intracellular 2-keto-3-deoxy-6-phosphogluconate in an Entner–Doudoroff aldolase mutant results in bacteriostasis. FEBS Microbiol. Lett. 159, 261–266 (1998).
Chantranupong, L., Wolfson, R. L. & Sabatini, D. M. Nutrient-sensing mechanisms across evolution. Cell 161, 67–83 (2015).
Korem Kohanim, Y. et al. A bacterial growth law out of steady state. Cell Rep. 23, 2891–2900 (2018).
Towbin, B. D. et al. Optimality and sub-optimality in a bacterial growth law. Nat. Commun. 8, 14123 (2017).
Fischer, E. & Sauer, U. A novel metabolic cycle catalyzes glucose oxidation and anaplerosis in hungry Escherichia coli. J. Biol. Chem. 278, 46446–46451 (2003).
Bren, A. et al. Glucose becomes one of the worst carbon sources for E. coli on poor nitrogen sources due to suboptimal levels of cAMP. Sci. Rep. 6, 24834 (2016).
Taylor, B. R. et al. A metabolic sum rule dictates bacterial response to short-chain fatty acid stress. Preprint at bioRxiv https://doi.org/10.1101/2022.08.31.506075 (2022).
Hermsen, R., Okano, H., You, C., Werner, N. & Hwa, T. A growth-rate composition formula for the growth of E. coli on co-utilized carbon substrates. Mol. Syst. Biol. 11, 801 (2015).
Xu, Y. F., Amador-Noguez, D., Reaves, M. L., Feng, X. J. & Rabinowitz, J. D. Ultrasensitive regulation of anapleurosis via allosteric activation of PEP carboxylase. Nat. Chem. Biol. 8, 562–568 (2012).
Doucette, C. D., Schwab, D. J., Wingreen, N. S. & Rabinowitz, J. D. α-ketoglutarate coordinates carbon and nitrogen utilization via enzyme I inhibition. Nat. Chem. Biol. 7, 894–901 (2011).
Kleijn, R. J., Van Winden, W. A., Van Gulik, W. M. & Heijnen, J. J. Revisiting the 13C-label distribution of the non-oxidative branch of the pentose phosphate pathway based upon kinetic and genetic evidence. FEBS J. 272, 4970–4982 (2005).
Stincone, A. et al. The return of metabolism: biochemistry and physiology of the pentose phosphate pathway. Biol. Rev. 90, 927–963 (2015).
Pereira, F. C. & Berry, D. Microbial nutrient niches in the gut. Environ. Microbiol. 19, 1366–1378 (2017).
Lenski, R. E., Rose, M. R., Simpson, S. C. & Tadler, S. C. Long-term experimental evolution in Escherichia coli. I. Adaptation and divergence during 2,000 generations. Am. Nat. 138, 1315–1341 (1991).
Wünsche, A. et al. Diminishing-returns epistasis decreases adaptability along an evolutionary trajectory. Nat. Ecol. Evol. 1, 61 (2017).
Lenski, R. E. Experimental evolution and the dynamics of adaptation and genome evolution in microbial populations. ISME J. 11, 2181–2194 (2017).
Emmerling, M. et al. Metabolic flux responses to pyruvate kinase knockout in Escherichia coli. J. Bacteriol. 184, 152–164 (2002).
Burgos-Barragan, G. et al. Mammals divert endogenous genotoxic formaldehyde into one-carbon metabolism. Nature 548, 549–554 (2017).
Tibbetts, A. S. & Appling, D. R. Compartmentalization of mammalian folate-mediated one-carbon metabolism. Annu. Rev. Nutr. 30, 57–81 (2010).
Corbet, C. & Feron, O. Cancer cell metabolism and mitochondria: nutrient plasticity for TCA cycle fueling. Biochim. Biophys. Acta Rev. Cancer 1868, 7–15 (2017).
Kiparissides, A. & Hatzimanikatis, V. Thermodynamics-based metabolite sensitivity analysis in metabolic networks. Metab. Eng. 39, 117–127 (2017).
Kee, J. M., Oslund, R. C., Perlman, D. H. & Muir, T. W. A pan-specific antibody for direct detection of protein histidine phosphorylation. Nat. Chem. Biol. 9, 416–421 (2013).
Nguyen, J. et al. A distinct growth physiology enhances bacterial growth under rapid nutrient fluctuations. Nat. Commun. 12, 3662 (2021).
Sekar, K. et al. Bacterial glycogen provides short-term benefits in changing environments. Appl. Environ. Microbiol. 86, e00049–20 (2020).
Jenior, M. L., Moutinho, T. J., Dougherty, B. V. & Papin, J. A. Transcriptome-guided parsimonious flux analysis improves predictions with metabolic networks in complex environments. PLoS Comput. Biol. 16, e1007099 (2020).
Wang, C. Y. et al. Metabolome and proteome analyses reveal transcriptional misregulation in glycolysis of engineered E. coli. Nat. Commun. 12, 4929 (2021).
Yan, Y., Liu, N. & Tang, Y. Recent developments in self-resistance gene directed natural product discovery. Nat. Prod. Rep. 37, 879–892 (2020).
Almabruk, K. H., Dinh, L. K. & Philmus, B. Self-resistance of natural product producers: past, present, and future focusing on self-resistant protein variants. ACS Chem. Biol. 13, 1426–1437 (2018).
Hofmeyr, J. H. S. & Cornish-Bowden, A. Quantitative assessment of regulation in metabolic systems. Eur. J. Biochem. 200, 223–236 (1991).
Kochanowski, K. et al. Global coordination of metabolic pathways in Escherichia coli by active and passive regulation. Mol. Syst. Biol. 17, e10064 (2021).
Vander Heiden, M. G. et al. Evidence for an alternative glycolytic pathway in rapidly proliferating cells. Science 329, 1492–1499 (2010).
Saragliadis, A., Trunk, T. & Leo, J. C. Producing gene deletions in Escherichia coli by P1 transduction with excisable antibiotic resistance cassettes. J. Vis. Exp. 2018, 58267 (2018).
Baba, T. et al. Construction of Escherichia coli K-12 in-frame, single-gene knockout mutants: the Keio collection. Mol. Syst. Biol. 2, 2006.0008 (2006).
Gutnick, D., Calvo, J. M., Klopotowski, T. & Ames, B. N. Compounds which serve as the sole source of carbon or nitrogen for Salmonella typhimurium LT-2. J. Bacteriol. 100, 215–219 (1969).
Orth, J. D. et al. A comprehensive genome-scale reconstruction of Escherichia coli metabolism—2011. Mol. Syst. Biol. 7, 535 (2011).
Seitzer, P., Bennett, B. & Melamud, E. MAVEN2: an updated open-source mass spectrometry exploration platform. Metabolites 12, 684 (2022).
Wang, L. et al. Peak annotation and verification engine for untargeted LC–MS metabolomics. Anal. Chem. 91, 1838–1846 (2019).
Yuan, J. et al. Metabolomics-driven quantitative analysis of ammonia assimilation in E. coli. Mol. Syst. Biol. 5, 302 (2009).
Park, J. O. et al. Metabolite concentrations, fluxes and free energies imply efficient enzyme usage. Nat. Chem. Biol. 12, 482–489 (2016).
Bennett, B. D., Yuan, J., Kimball, E. H. & Rabinowitz, J. D. Absolute quantitation of intracellular metabolite concentrations by an isotope ratio-based approach. Nat. Protoc. 3, 1299–1311 (2008).
Acknowledgements
The authors would like to thank the members of the Park laboratory, the UCLA Metabolomics Center and the UCLA Molecular Instrumentation Center for helpful discussion and J. E. Barrick, R. E. Lenski and colleagues for the E. coli strains from the long-term evolution experiment. This work was supported by the National Institute of General Medical Sciences of the National Institutes of Health under award number R35GM143127 (J.O.P.); the National Institutes of Health Instrumentation Grant 1S10OD016387-01; the BioPACIFIC Materials Innovation Platform of the National Science Foundation under award number DMR-1933487 (R.C.L., G.N. and J.O.P.); the National Science Foundation Research Traineeship in Integrated Urban Solutions for Food, Energy, and Water Management (NSF-INFEWS) under award number DGE-1735325 (G.N.); the Hellman Fellowship (J.O.P.); and the Center for Clean Technology Fellowship (R.C.L.). The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health or the National Science Foundation.
Author information
Authors and Affiliations
Contributions
R.C.L. and J.O.P. designed the study and wrote the paper. R.C.L. and G.N. carried out the experiments. R.C.L. and J.O.P. analyzed the metabolomic and isotope-labeling results. R.C.L., G.N. and J.O.P. analyzed the sequencing data from the long-term evolution experiment.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Chemical Biology thanks Markus Basan and Hannes Link for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Parallel glycolysis bolsters growth acceleration upon carbon and nitrogen upshifts.
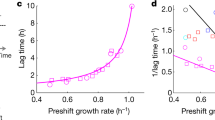
a, WT and ∆edd strains had been cultured on acetate as the sole carbon source and, at t = 0 min, were spiked with glucose. Growth rates were monitored until a steady growth rate of ~0.7 hr–1 was achieved 80 minutes after upshift. b, WT and ∆edd strains were cultured on arginine as the sole nitrogen source before ammonia was spiked in at t = 0 min. Growth rates were monitored until a stable growth rate of ~0.75 hr–1 was achieved. c, The growth rates of WT and ∆edd were measured in glucose minimal media with and without high acetate and compared to the stable growth rates of E. coli cultures following carbon upshift from acetate to glucose spiked in. Error bars represent the s.e.m. (n = 3 biological replicates).
Extended Data Fig. 2 The ED pathway improves growth acceleration upon carbon upshift from non-glycolytic to glycolytic substrates.
a, WT and ∆edd cultures had been grown on succinate or glycerol as the sole carbon source and, at t = 0 min, were spiked with glucose. Growth rates and growth acceleration, which is the time derivative of growth rate, were quantified. b, Cells had been grown on a PTS sugar mannose as the sole carbon source and, at t = 0 min, were spiked with glucose. Growth rates as well as growth acceleration over two time periods were quantified. c, Cells were subjected to carbon upshift from acetate to succinate, both gluconeogenic substrates at t = 0 min. Growth rates and growth acceleration were quantified. Error bars represent the s.e.m. (n = 3 biological replicates). P values are obtained by two-tailed t-tests.
Extended Data Fig. 3 [1,2-13C2]- and [5,6-13C2]-glucose reveal fluxes through different glycolytic routes.
a, [1,2-13C2]-glucose uniquely generates M + 1 labeled triose phosphates through the oxidative pentose phosphate pathway (OxPPP). For every three glucose molecules going through the PPP, three molecules of M + 0, one molecule of M + 1, and one molecule of M + 2 triose phosphate are generated. b, [5,6-13C2]-glucose introduces unlabeled lower glycolytic intermediates only through the EMP pathway. Unlike in the case of [1,2-13C2]-glucose, the EMP and the ED pathways produce the same positionally labeled pyruvate. The major fraction of M + 2 3PG indicated the ED pathway activity in the nutrient replete condition. c, Using the carbon balance model of central carbon metabolism with [5,6-13C2]-glucose, 3PG and pyruvate (PYR) labeling measurements are sufficient to solve for central carbon metabolism fluxes. This elegant solution is attributable to the fact that [5,6-13C2]-glucose tracing is not convoluted by carbon shuffling in the PPP. Mass balance of carbons over the EMP pathway, the ED pathway, and the PPP (the inner control volume) as well as lower glycolysis (the outer control volumes) revealed the relationship between 3PG and PYR labeling and the central carbon metabolism fluxes.
Extended Data Fig. 4 Rapid labeling and turnover of glycolytic intermediates.
a, E. coli grown on an unlabeled medium were rapidly switched to the same medium containing [U-13C6]-glucose. The labeled fractions of glycolytic intermediates were measured using LC-MS and normalized to the labeled fraction of glucose. Error bars represent the s.e.m. (n = 3 biological replicates). b, The turnover of individual metabolites was calculated as the flux through that metabolite divided by the metabolite pool size using previously reported fluxes and concentrations56.
Extended Data Fig. 5 The ED pathway flux jumps upon nutrient upshifts.
a,b, Nutrient upshift experiments analogous to those conducted with [1,2-13C2]-glucose were replicated using [5,6-13C2]-glucose and the resulting labeling of (a) 6PG and (b) 3PG was measured via LC-MS. The labeling of M + 0 and M + 2 isotopomers revealed both the ED pathway flux and recursive PPP usage (Supplementary Note 3). c, Cell lysates from E. coli cultures grown on acetate were collected for an enzyme activity assay and incubated with 6PG, the direct precursor to KDPG (m/z = 257.0066) in the ED pathway. The increase in KDPG over time indicated the presence of the ED pathway enzymes in cells operating gluconeogenesis. Error bars represent the s.e.m. (n = 2 biological replicates). d, E. coli in the carbon-limited acetate medium underwent carbon upshift by [U-13C6]-glucose addition. The labeling of glycolytic intermediates following the upshift informed us of the fractions of 3PG pool coming from the labeled glucose and from the pre-upshift unlabeled acetate. This information was used to accurately compute glycolytic fluxes from [1,2-13C2]-glucose and [5,6-13C2]-glucose tracing. Error bars represent the s.e.m. (n = 3 biological replicates).
Extended Data Fig. 6 Flow cytometry of WT-∆edd co-cultures.
At least 10,000 events were recorded per sample. Singlets were gated based on FSC-A (forward scattered area) and FSC-H (forward scatted height) intensities in a way to minimize the frequency of aggregates observed. Non-fluorescent (WT), GFP expressing (WT + GFP) and mCherry expressing (∆edd+mCherry) cells were used as controls and analyzed on the day of experiments to determine the compensation of spectral overlap between fluorophores and gating quadrants for the glucose upshift (top) and ammonia upshift (bottom) experiments. Samples at the beginning (left) and end (right) of co-culture intermittent nutrient supply experiments are shown as examples.
Extended Data Fig. 7 Mutations in the LTEE reflect selective pressure.
a,b, Mutation frequencies in genes deviated from the Poisson distribution. Genes with the same number of mutations were counted, and the normalized counts were fit to a Poisson distribution. Poor fitting for genes with 2 or more mutations suggested a driving force toward improved fitness. Error bars represent the standard deviation (n = 212 all possible sets of twelve independent clones). c,d, The total number of mutations in individual genes across all 24 strains at the 50,000th generation. For each gene, we counted the number of mutations in all 24 strains. A few genes were consistently and heavily mutated: malT is a regulator for the maltose regulon that is also involved in λ phage infection; yeeJ is an autotransporter protein; metH is the gene for methionine synthase; pykF is pyruvate kinase, the last step of glycolysis; thrA is a bifunctional aspartokinase and homoserine dehydrogenase; nadR is a regulator for NAD biosynthesis; hsIU is a putative ATP dependent protease; and iclR is a regulator for isocitrate lyase of the glyoxylate cycle.
Extended Data Fig. 8 Mutations in the EMP and the ED pathways over the LTEE improve growth acceleration.
a, Development of nonsynonymous mutations and indels in randomly sampled gene groups were compared to the mutation history of the parallel glycolytic pathways over 50,000 generations of the E. coli long-term evolution experiment (LTEE). b, The proportions of nonsynonymous mutations and indels were monitored over the LTEE. c, List of mutations in the ED and the EMP pathways. The 24 genome sequences from 12 populations were analyzed at generation 50,000 for mutations in the two glycolytic pathways. SNP denotes a single nucleotide polymorphism (only nonsynonymous mutations are shown) and FS denotes a frameshift indel. d, Seven strains from the LTEE with different numbers of overall and ED-pathway mutations were chosen for growth assays. e, The LTEE strains underwent carbon upshifts from no carbon source, acetate, mannose, or glycerol by spiking in glucose, and their growth acceleration was measured. Error bars represent the s.e.m. (n = 3 biological replicates). P values are obtained by two-tailed t-tests between the designated and ancestral strain. f, After power analysis, the ancestral strain and Ara+3 A, Ara+5 A, and Ara-2 A strains isolated in the 5,000th generation were subjected to acetate-to-glucose upshift, and their growth acceleration was measured. Each box shows the three quartiles, and whiskers extend to the minimum and maximum values within 1.5-fold of the interquartile range (n = 6 biological replicates). Statistical significance between strains was obtained by one-way ANOVA with Tukey’s honestly significant difference test for multiple comparison.
Extended Data Fig. 9 The ED pathway provides dual benefits of carbon-efficient NADPH production and seamless nitrogen-controlled glucose import.
a, The OxPPP produces two NADPH, but a carbon is lost as CO2 as 6PG enters the non-oxidative pentose phosphate pathway. On the other hand, the ED pathway produces NADPH without decarboxylation. b, Exponentially growing E. coli in unlabeled glucose minimal medium were rapidly switched to either [1-2H1]- or [3-2H2]-glucose. Labeling of the hydride of NADPH from [1-2H1]-glucose indicated flux through G6PDH, whereas labeling from [3-2H1]-glucose indicated flux through 6PGD. The ~3-fold labeling of NADPH from G6PDH compared to 6PGD indicated comparable contributions of the ED pathway and the OxPPP to NADPH generation. c, Concerted action of the phosphotransferase system (PTS), phosphoenolpyruvate synthase (PpsA), and ED pathway streamlines glucose import upon nitrogen upshift. While the EMP pathway takes 8 steps to generate PEP for glucose import, the concerted regulation and action of the ED pathway and PpsA supply PEP in 5 steps.
Supplementary information
Supplementary Information
Supplementary Notes 1–4, Tables 1–17 and References.
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 5
Statistical source data.
Source Data Fig. 6
Statistical source data.
Source Data Extended Data Fig./Table 1
Statistical source data.
Source Data Extended Data Fig./Table 2
Statistical source data.
Source Data Extended Data Fig./Table 3
Statistical source data.
Source Data Extended Data Fig./Table 4
Statistical source data.
Source Data Extended Data Fig./Table 5
Statistical source data.
Source Data Extended Data Fig./Table 6
Source flow cytometry .fcs files.
Source Data Extended Data Fig./Table 7
Statistical source data.
Source Data Extended Data Fig./Table 8
Statistical source data.
Source Data Extended Data Fig./Table 9
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Law, R.C., Nurwono, G. & Park, J.O. A parallel glycolysis provides a selective advantage through rapid growth acceleration. Nat Chem Biol 20, 314–322 (2024). https://doi.org/10.1038/s41589-023-01395-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-023-01395-2