Abstract
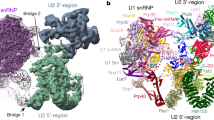
The spliceosome catalyses the excision of introns from pre-mRNA in two steps, branching and exon ligation, and is assembled from five small nuclear ribonucleoprotein particles (snRNPs; U1, U2, U4, U5, U6) and numerous non-snRNP factors1. For branching, the intron 5′ splice site and the branch point sequence are selected and brought by the U1 and U2 snRNPs into the prespliceosome1, which is a focal point for regulation by alternative splicing factors2. The U4/U6.U5 tri-snRNP subsequently joins the prespliceosome to form the complete pre-catalytic spliceosome. Recent studies have revealed the structural basis of the branching and exon-ligation reactions3, however, the structural basis of the early events in spliceosome assembly remains poorly understood4. Here we report the cryo-electron microscopy structure of the yeast Saccharomyces cerevisiae prespliceosome at near-atomic resolution. The structure reveals an induced stabilization of the 5′ splice site in the U1 snRNP, and provides structural insights into the functions of the human alternative splicing factors LUC7-like (yeast Luc7) and TIA-1 (yeast Nam8), both of which have been linked to human disease5,6. In the prespliceosome, the U1 snRNP associates with the U2 snRNP through a stable contact with the U2 3′ domain and a transient yeast-specific contact with the U2 SF3b-containing 5′ region, leaving its tri-snRNP-binding interface fully exposed. The results suggest mechanisms for 5′ splice site transfer to the U6 ACAGAGA region within the assembled spliceosome and for its subsequent conversion to the activation-competent B-complex spliceosome7,8. Taken together, the data provide a working model to investigate the early steps of spliceosome assembly.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Will, C. L. & Lührmann, R. Spliceosome structure and function. Cold Spring Harb. Perspect. Biol. 3, a003707 (2011).
Papasaikas, P. & Valcárcel, J. The spliceosome: the ultimate RNA chaperone and sculptor. Trends Biochem. Sci. 41, 33–45 (2016).
Wilkinson, M. E., Lin, P. C., Plaschka, C. & Nagai, K. Cryo-EM studies of pre-mRNA splicing: from sample preparation to model visualization. Annu. Rev. Biophys. 47, 175–199 (2018).
Behzadnia, N. et al. Composition and three-dimensional EM structure of double affinity-purified, human prespliceosomal A complexes. EMBO J. 26, 1737–1748 (2007).
Gao, G. & Dudley, S. C. Jr. RBM25/LUC7L3 function in cardiac sodium channel splicing regulation of human heart failure. Trends Cardiovasc. Med. 23, 5–8 (2013).
Hackman, P. et al. Welander distal myopathy is caused by a mutation in the RNA-binding protein TIA1. Ann. Neurol. 73, 500–509 (2013).
Bertram, K. et al. Cryo-EM structure of a pre-catalytic human spliceosome primed for activation. Cell 170, 701–713.e711 (2017).
Plaschka, C., Lin, P. C. & Nagai, K. Structure of a pre-catalytic spliceosome. Nature 546, 617–621 (2017).
Liang, W. W. & Cheng, S. C. A novel mechanism for Prp5 function in prespliceosome formation and proofreading the branch site sequence. Genes Dev. 29, 81–93 (2015).
Kondo, Y., Oubridge, C., van Roon, A. M. & Nagai, K. Crystal structure of human U1 snRNP, a small nuclear ribonucleoprotein particle, reveals the mechanism of 5′ splice site recognition. eLife 4, e04986 (2015).
Neubauer, G. et al. Identification of the proteins of the yeast U1 small nuclear ribonucleoprotein complex by mass spectrometry. Proc. Natl Acad. Sci. USA 94, 385–390 (1997).
Li, X. et al. CryoEM structure of Saccharomyces cerevisiae U1 snRNP offers insight into alternative splicing. Nat. Commun. 8, 1035 (2017).
Puig, O., Bragado-Nilsson, E., Koski, T. & Séraphin, B. The U1 snRNP-associated factor Luc7p affects 5′ splice site selection in yeast and human. Nucleic Acids Res. 35, 5874–5885 (2007).
Agarwal, R., Schwer, B. & Shuman, S. Structure-function analysis and genetic interactions of the Luc7 subunit of the Saccharomyces cerevisiae U1 snRNP. RNA 22, 1302–1310 (2016).
Schwer, B. & Shuman, S. Structure–function analysis of the Yhc1 subunit of yeast U1 snRNP and genetic interactions of Yhc1 with Mud2, Nam8, Mud1, Tgs1, U1 snRNA, SmD3 and Prp28. Nucleic Acids Res. 42, 4697–4711 (2014).
Chen, J. Y. et al. Specific alterations of U1-C protein or U1 small nuclear RNA can eliminate the requirement of Prp28p, an essential DEAD box splicing factor. Mol. Cell 7, 227–232 (2001).
Puig, O., Gottschalk, A., Fabrizio, P. & Séraphin, B. Interaction of the U1 snRNP with nonconserved intronic sequences affects 5′ splice site selection. Genes Dev. 13, 569–580 (1999).
Förch, P., Puig, O., Martínez, C., Séraphin, B. & Valcárcel, J. The splicing regulator TIA-1 interacts with U1-C to promote U1 snRNP recruitment to 5′ splice sites. EMBO J. 21, 6882–6892 (2002).
Spingola, M. & Ares, M. Jr. A yeast intronic splicing enhancer and Nam8p are required for Mer1p-activated splicing. Mol. Cell 6, 329–338 (2000).
Agafonov, D. E. et al. ATPγS stalls splicing after B complex formation but prior to spliceosome activation. RNA 22, 1329–1337 (2016).
Fica, S. M. & Nagai, K. Cryo-electron microscopy snapshots of the spliceosome: structural insights into a dynamic ribonucleoprotein machine. Nat. Struct. Mol. Biol. 24, 791–799 (2017).
Abovich, N. & Rosbash, M. Cross-intron bridging interactions in the yeast commitment complex are conserved in mammals. Cell 89, 403–412 (1997).
Tang, Q. et al. SF3B1/Hsh155 HEAT motif mutations affect interaction with the spliceosomal ATPase Prp5, resulting in altered branch site selectivity in pre-mRNA splicing. Genes Dev. 30, 2710–2723 (2016).
Staley, J. P. & Guthrie, C. An RNA switch at the 5′ splice site requires ATP and the DEAD box protein Prp28p. Mol. Cell 3, 55–64 (1999).
Agafonov, D. E. et al. Molecular architecture of the human U4/U6.U5 tri-snRNP. Science 351, 1416–1420 (2016).
Nguyen, T. H. D. et al. Cryo-EM structure of the yeast U4/U6.U5 tri-snRNP at 3.7 Å resolution. Nature 530, 298–302 (2016).
Huang, Y. H., Chung, C. S., Kao, D. I., Kao, T. C. & Cheng, S. C. Sad1 counteracts Brr2-mediated dissociation of U4/U6.U5 in tri-snRNP homeostasis. Mol. Cell. Biol. 34, 210–220 (2014).
Boesler, C. et al. A spliceosome intermediate with loosely associated tri-snRNP accumulates in the absence of Prp28 ATPase activity. Nat. Commun. 7, 11997 (2016).
Wan, R. et al. The 3.8 Å structure of the U4/U6.U5 tri-snRNP: Insights into spliceosome assembly and catalysis. Science 351, 466–475 (2016).
Ismaïli, N., Sha, M., Gustafson, E. H. & Konarska, M. M. The 100-kda U5 snRNP protein (hPrp28p) contacts the 5′ splice site through its ATPase site. RNA 7, 182–193 (2001).
Nguyen, T. H. D. et al. The architecture of the spliceosomal U4/U6.U5 tri-snRNP. Nature 523, 47–52 (2015).
Umen, J. G. & Guthrie, C. A novel role for a U5 snRNP protein in 3′ splice site selection. Genes Dev. 9, 855–868 (1995).
Galej, W. P. et al. Cryo-EM structure of the spliceosome immediately after branching. Nature 537, 197–201 (2016).
Wang, H., Ach, R. A. & Curry, B. Direct and sensitive miRNA profiling from low-input total RNA. RNA 13, 151–159 (2007).
Mastronarde, D. N. Automated electron microscope tomography using robust prediction of specimen movements. J. Struct. Biol. 152, 36–51 (2005).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Zhang, K. Gctf: Real-time CTF determination and correction. J. Struct. Biol. 193, 1–12 (2016).
Scheres, S. H. W. & Chen, S. Prevention of overfitting in cryo-EM structure determination. Nat. Methods 9, 853–854 (2012).
Kimanius, D., Forsberg, B. O., Scheres, S. H. & Lindahl, E. Accelerated cryo-EM structure determination with parallelisation using GPUs in RELION-2. eLife 5, e18722 (2016).
Scheres, S. H. RELION: implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Kucukelbir, A., Sigworth, F. J. & Tagare, H. D. Quantifying the local resolution of cryo-EM density maps. Nat. Methods 11, 63–65 (2014).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010).
Webb, B. & Sali, A. Comparative protein structure modeling using MODELLER. Curr. Protoc. Bioinformatics 5.6.1–5.6.32 (2014).
Yang, J. et al. The I-TASSER Suite: protein structure and function prediction. Nat. Methods 12, 7–8 (2015).
Kretzner, L., Krol, A. & Rosbash, M. Saccharomyces cerevisiae U1 small nuclear RNA secondary structure contains both universal and yeast-specific domains. Proc. Natl Acad. Sci. USA 87, 851–855 (1990).
Jacewicz, A., Schwer, B., Smith, P. & Shuman, S. Crystal structure, mutational analysis and RNA-dependent ATPase activity of the yeast DEAD-box pre-mRNA splicing factor Prp28. Nucleic Acids Res. 42, 12885–12898 (2014).
Hadjivassiliou, H., Rosenberg, O. S. & Guthrie, C. The crystal structure of S. cerevisiae Sad1, a catalytically inactive deubiquitinase that is broadly required for pre-mRNA splicing. RNA 20, 656–669 (2014).
Absmeier, E., Santos, K. F. & Wahl, M. C. Functions and regulation of the Brr2 RNA helicase during splicing. Cell Cycle 15, 3362–3377 (2016).
Ester, C. & Uetz, P. The FF domains of yeast U1 snRNP protein Prp40 mediate interactions with Luc7 and Snu71. BMC Biochem. 9, 29 (2008).
Kimura, E. et al. Serine-arginine-rich nuclear protein Luc7l regulates myogenesis in mice. Gene 341, 41–47 (2004).
Yan, C., Wan, R., Bai, R., Huang, G. & Shi, Y. Structure of a yeast activated spliceosome at 3.5 Å resolution. Science 353, 904–911 (2016).
Wilkinson, M. E. et al. Postcatalytic spliceosome structure reveals mechanism of 3′-splice site selection. Science 358, 1283–1288 (2017).
Robert, X. & Gouet, P. Deciphering key features in protein structures with the new ENDscript server. Nucleic Acids Res. 42, W320–W324 (2014).
Sievers, F. et al. Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol. Syst. Biol. 7, 539 (2011).
Buchan, D. W., Minneci, F., Nugent, T. C., Bryson, K. & Jones, D. T. Scalable web services for the PSIPRED Protein Analysis Workbench. Nucleic Acids Res. 41, W349–W357 (2013).
Acknowledgements
We thank C. Savva, S. Chen, G. Cannone, G. McMullan, J. Grimmett and T. Darling for maintaining electron microscopy and computing facilities; the mass spectrometry facility for protein identification; A. Murzin for discussions; E. Hesketh and R. Thompson for assistance with cryo-EM data collection of dataset three and S.-C. Cheng, A. Newman, L. Strittmatter, M. E. Wilkinson for critical reading of the manuscript. We thank J. Löwe, V. Ramakrishnan, D. Barford and R. Henderson for their continuing support. The project was supported by the Medical Research Council (MC_U105184330) and European Research Council Advanced Grant (693087-SPLICE3D). C.P. was supported by an EMBO Long-Term Fellowship (984-2015).
Author information
Authors and Affiliations
Contributions
C.P. and P.-C.L. established complex preparation, performed cryo-EM data analysis, model building, and refinement. C.C. assisted with cryo-EM analysis. C.P., P.-C.L. and K.N. analysed the structure and wrote the manuscript. C.P. and P.-C.L. designed the project and K.N. supervised the project.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Biochemical characterization and cryo-EM of the prespliceosome A-complex.
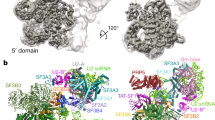
a, Mutation of the UBC4 pre-mRNA branch point sequence (UACUAAC to UACAAAC, in which A is the branch-point adenosine and A is the mutated nucleotide) stalls splicing before the first step, as described9. Splicing reactions were carried out for 30 min at 23 °C in yeast extract using wild-type (lane one) or mutant (U > A, lane two) pre-mRNA. This experiment was performed three times. The asterisk indicates a degradation product. For gel source data see Supplementary Fig. 1a. b, Protein analysis of purified A-complex (SDS–PAGE stained with Coomassie blue). The U2-associated Prp5 protein is sub-stoichiometric and not observed in the A-complex structure. The purification and analysis of protein compositions were performed at least five times with similar results. For gel source data see Supplementary Fig. 1b. c, Cryo-EM micrograph of the A-complex. Scale bar, 100 nm. d, 2D class averages of the A-complex were determined in RELION 2.139,40, and reveal a bipartite architecture, comprising the U1 snRNP and the U2 snRNP 3′ and 5′ regions, respectively. e, Composite cryo-EM density of the A-complex shown in two orthogonal views (compare to Fig. 1). The respective densities used for modelling the U1 snRNP (A2, grey), the U2 3′ region (A1, cyan), and the U2 5′ region (A3, green) are coloured and superimposed on a transparent outline of the full A3 map (Methods). The overall resolution of each map as well as the percentage from the cleaned dataset of 153,556 particles are shown in parentheses. Non-modelled regions are indicated and putatively assigned. f. Composite cryo-EM density with the final A-complex model superimposed in a cartoon representation. The path of 40 nucleotides of the disordered UBC4 pre-mRNA intron are indicated. A-complex components are coloured as in Fig. 1. Views as in e.
Extended Data Fig. 2 Cryo-EM image classification and refinement.
a, Image processing workflow for analysis of the A-complex cryo-EM dataset (see ‘Image processing’ in Methods). To visualize differences between the reconstructions, the U1 snRNP (grey), U2 3′ (cyan) and U2 5′ regions (green) are coloured. For each round of three-dimensional classification, the percentage of the data and the type of soft-edged mask are indicated. The type of mask and overall resolution are indicated for each 3D refinement (blue box). b, Orientation distribution plots for all particles that contribute to the respective A1, A2, and A3 cryo-EM reconstructions. c, Gold-standard Fourier shell correlation (FSC = 0.143) of the respective A1, A2 and A3 cryo-EM reconstructions. d, Two views of the composite A-complex cryo-EM density (maps A1, A2 and A3) coloured by local resolution as determined by ResMap43. e, As panel d, but for a central slice through the composite A-complex cryo-EM map.
Extended Data Fig. 3 Details of the U1 snRNP.
a, U1 snRNP structure with subunits coloured as in Fig. 1, except for Nam8 (orange), Snu56 (light blue), Snu71 (blue), Luc7 (dark purple), Mud1 (red) and the U1 snRNA (various). The pre-mRNA nucleotides are labelled relative to the first nucleotide (+1) of the intron. The Nam8 RRM1 and RRM2 domains are flexible and project downstream of the 5′SS. The protein attributed to Luc7 in the free U1 snRNP structure12 was re-assigned to Snu71. C-term, C terminus; N-term, N terminus; SL, stem loop. In the structure we do not observe any evidence that the C-terminal tails of SmB, SmD1, and SmD3 interact with the 5′SS, consistent with their absence in the human 5′SS–minimal U1 snRNP crystal structure10. b, Representative regions of the sharpened U1 snRNP density determined at 4 Å resolution (map A2) are superimposed on the refined coordinate model. The density reveals side-chain details, and here segments from the Prp42 N terminus (TPR repeat 1), the Sm ring subunit SmB, and the Snu56 α-helical domain are shown. c, The A2 cryo-EM density is shown superimposed on the coordinate models of a selection of U1 snRNP proteins: Luc7, Snu71, Yhc1 and Prp39. In the structure most of Snu71 is disordered, except for a small N-terminal domain (residues 2–43) that binds between the Prp42 N terminus and the Snu56 KH-like fold, consistent with protein crosslinking12. Functional regions and disordered domains are indicated. d, The U1 snRNA–pre-mRNA 5′ splice site (U1–5′SS) model is superimposed on its cryo-EM density (map A2). A secondary structure diagram of the U1–5′SS interaction is shown underneath the model. The register of the U1–5′SS is shifted by one nucleotide with respect to U1C (Yhc1) compared to the minimal human 5′SS–U1 snRNP crystal structure, owing to an additional nucleotide in the yeast U1 snRNA10 (U11). Lines indicate Watson–Crick base pairs and dots indicate pseudouridine (ψ)-containing base pairs. e, The Prp39–Prp42 heterodimer is coloured to indicate each of their respective TPR repeats. f, Cryo-EM density of U1 snRNA from maps A2 (dark grey) and A3 (light grey) without (top) and with the superimposed coordinate model of yeast U1 snRNA (bottom). The model is labelled and coloured according to functional regions of U1 snRNA (5′ end, pink; H helix, cyan; SL1, dark blue; SL2-1, green; SL3-1, light blue; SL2-2 and SL3-2 to -7, grey; 3′end and Sm site, yellow). g, Secondary-structure diagram of U1 snRNA. Bold letters indicate residues included in the model, lines indicate Watson–Crick base pairs, and dots G–U wobble and pseudouridine-containing base pairs. Compare to e. The conserved U1 snRNA ‘core’ is outlined with a grey box. The region of the putative phosphate backbone model of part of the U1 SL3-7 region is indicated with a grey box.
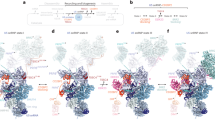
Extended Data Fig. 4 Comparisons of yeast and human U1 snRNPs and implications for alternative splicing.
a, Formation of the U1–5′SS helix induces stable binding of Luc7. In the absence of a pre-mRNA 5′SS in the free U1 snRNP density (left, EMD-8622), Luc7 and the U1 5′ end are disordered. Upon 5′SS recognition at the U1 5′ end (centre, map A2), Luc7 becomes ordered and stabilizes the U1–5′SS interaction, suggesting a mechanism for the selection of weak 5′SS sequences. The free U1 snRNP and the 5′SS-bound (map A2) cryo-EM densities are superimposed on the right. Although the long α-helical density next to Luc7 cannot be assigned with confidence, protein–protein crosslinking data12 and protein secondary structure prediction are consistent with the presence of either Prp40 or Snu71. On the basis of additional biochemical data on the interaction between the α-helical Prp40 FF1 domain and Luc7 ZnF252, we would speculate that the Prp40 FF1 domain is the most likely candidate for this density. b, Comparison of the yeast U1 snRNP ‘core’ with the human U1 snRNP crystal structure (PDB ID 3CW1). Protein and RNA (top) and RNA only (bottom) are shown side by side (left and centre) and superimposed by a global alignment in PyMOL (right). Coloured as in Extended Data Fig. 3a. c, The yeast U1 snRNP model suggests regulatory mechanisms for human alternative splicing factors. The human homologues of the peripheral yeast U1 proteins may function through stabilization of the U1–5′SS interaction (region 1), of the U1–U2 3′ region interface (region 2), or the U1–U2 5′ interface (region 3). The yeast U1 snRNP ‘core’ is shown superimposed on a surface representation of the U1 snRNP model (top), compared with the similarly coloured human U1 snRNP (below). Interaction sites with the U2 snRNP are labelled (top). d, The location of yeast U1 snRNP components with homology to human splicing factors are indicated in the U1 snRNP structure. The Prp39–Prp42 heterodimer (human PRPF39 homodimer), Nam818 (human TIA-1 and TIA-R), Luc753 (human LUC7L1–3), and the Yhc1 C terminus (human U1C) have clear counterparts in the human system. The yeast-specific U1 snRNA insertions may be replaced in the human system by alternative splicing factors that modulate interactions with the U2 5′ region. e, Model of the yeast E complex on the basis of the U1 snRNP structure and biochemical data22. Luc7, Snu71 and Prp40 form a heterotrimer in vitro52, and their interacting regions may be located near unassigned density (compare to Extended Data Fig. 1e) at the tip of an unassigned 40-residue α-helix next to Luc7 ZnF2. This helix is likely to belong to the U1 subunit Snu71 or Prp40, consistent with protein crosslinking12 and protein secondary-structure prediction. Prp40 could then bind the yeast branch point-binding protein (BBP, human SF1), which in turn interacts with Mud2 (human U2AF65) to tether the pre-mRNA branch-point sequence in the E complex22.
Extended Data Fig. 5 Conformational flexibility of the U2 snRNP.
a, Two defined positions of the U1 snRNP-U2 3′ region could be identified relative to the U2 5′ region. A-complex models were fitted into class two and four from round two of the 3D image classification (compare Extended Data Fig. 2a). The classes are aligned via their U2 5′ region, illustrating their relative flexibility. b, Cartoon schematic of observed positions of the U2 3′ region relative to the U2 5′ region in the A-complex (left), B-complex8 (centre), and activated B-complex (Bact) (right, modelled from previously published work54). Although in the B-complex the U2 3′ region is free, in the A- and Bact-complexes the position of the U2 3′ region is influenced by interactions with Prp39 as well as Syf1 and Clf1, respectively. c, The U2 snRNP subunit Lea1 (human U2A′) aids to position the U2 snRNP 3′ domain in different spliceosome states. In our A-complex structure, the Prp39 TPR repeat T1 contacts the helical C terminus of Lea1. In the yeast C-complex structure, the non-modelled density for the Syf1 N terminus binds a neighbouring but non-overlapping surface of Lea1 (PDB ID 5LJ5). In the C*/P-complex55 (PDB ID 6EXN), the Syf1 N terminus binds yet another Lea1 surface and the U2 3′ domain is repositioned relative to its C-complex location. Together, this suggests that the Lea1 provides multiple interfaces that can be used to position the U2 3′ domain in different spliceosomal complexes. d, Fit of the U2 3′ region coordinate model to the A1 cryo-EM density. The dashed black lineseparates the U2 3′ domain (Sm ring, Msl1 and Lea1 subunits and U2 snRNA, left) and the SF3a subcomplex (Prp9, Prp11 and Prp21, right). Two orthogonal views are shown (Supplementary Video 2). e, Fit of the U2 5′ region coordinate model to the A3 cryo-EM density. A density consistent with the U2 snRNA stem IIa/b and the branch helix is observed. Two density thresholds are shown side by side (left, 0.0163; right, 0.0121), and orthogonal views are shown underneath (Supplementary Video 2).
Extended Data Fig. 6 Luc7 and Nam8 sequence alignments.
a, The Luc7 (human LUC7-like) amino-acid sequence alignment comparing S. cerevisiae, Kluyveromyces lactis, Schizosaccharomyces pombe, Danio rerio, Xenopus tropicalis, Mus musculus, Bos taurus and Homo sapiens was generated with Clustal Omega and visualized with ESPript 356,57. For the human sequence, LUC7L1 was used. Secondary structure elements are indicated above the sequence and derive from the A-complex structure (purple) or PSIPRED58 secondary structure prediction (grey). Modelled regions (dashed line) and the Zn-coordinating residues of ZnF1 and ZnF2 (asterisks) are indicated. Invariant or conserved residues are highlighted with a red box or red letter font, respectively. b. As in panel a but for Nam8 (human TIA-1) comparing S. cerevisiae, K. lactis, S. pombe, Drosophila melanogaster, D. rerio, X. tropicalis, M. musculus, B. taurus, and H. sapiens amino acid sequences.
Extended Data Fig. 7 Details of the pre-B-complex model.
a, Multiple views of the pre-B-complex model, generated by combining functional and structural data from yeast and human systems8,25. The mobility of the U1 snRNP relative to the U2 snRNP in the A-complex (this study) as well as of the U2 snRNP relative to tri-snRNP in the B-complex structure8 are indicated (left). The pre-B model contained only minor clashes, and a clash between the highly flexible Prp28 C-terminal RecA-2 lobe (from the human tri-snRNP25) and the highly flexible U6 snRNA 5′ stem loop (from the yeast B-complex8) may be resolved by small movements of either domain. b, Structural comparisons of the yeast pre-B model (from this study) and the yeast B-complex structure (PDB ID 5NRL8) suggest the existence of a molecular checkpoint to couple 5′SS transfer to U1 snRNP release and formation of the activation-competent B-complex. In the pre-B model (left) Sad1 tethers Brr2 through its interaction with the conserved Brr2 PWI domain51, and the U1 snRNP and its U1–5′SS helix are positioned near the U6 ACAGAGA region and the helicase Prp28. Subsequent to Prp28-mediated 5′SS transfer, Brr2 is repositioned onto its U4 snRNA substrate, guided by the B-complex-specific proteins (right). In this conformation the Brr2 helicase and its associated factors would clash with the U1 snRNP, consistent with U1 snRNP destabilization and release yeast and human B-complexes7,8. Brr2 is now ready to initiate spliceosome activation and formation of the active site in the Bact-complex. Regions that are changed between pre-B- and B-complex models (black outline) and the clash between the Brr2-containing ‘helicase’ domain and the U1 snRNP in B-complex (red X) are indicated. The lower right panel would conform to the alternative ‘U1-B-complex’ model.
Extended Data Fig. 8 Model for early splicing events.
a, Cartoon schematic of proposed early splicing events, detailing (i) assembly of the pre-B-complex spliceosome from the A-complex and the U4/U6.U5 tri-snRNP and (ii) the subsequent conversion to the pre-catalytic B-complex spliceosome. In the pre-B model the mobile U1 snRNP is next to Prp28, which is bound at the Prp8N domain. To initiate 5′SS transfer, Prp28 could clamp the pre-mRNA at, or next to, the U1–5′SS helix to destabilize it and to hand over the 5′SS to the U6 ACAGAGA region of tri-snRNP, consistent with protein–RNA crosslinks30. Transfer of the 5′SS may induce the binding of the B-complex proteins to replace Prp28 at the Prp8N domain and induce the large movement of Brr2 to its B-complex location on U4 snRNA. The U1 snRNP, now loosely tethered to U2, may dissociate from the B-complex owing to the steric clash with the Brr2-containing ‘helicase’ domain8 (Extended Data Fig. 7b). Consistent with this, the human pre-B-complex converts to a B-complex-like state in the presence of a 5′SS oligonucleotide, which coincides with U1 snRNP release28. This model can explain how Brr2 is kept inactive to prevent premature U4/U6 duplex unwinding26. The model thereby implies the existence of a molecular checkpoint, coupling 5′SS transfer from U1 to U6 snRNA with Brr2 helicase repositioning and U1 snRNP release to generate the activation-competent B-complex spliceosome. b, Cartoon schematic of an alternative model for spliceosome assembly and 5′SS transfer that relies only on the yeast A-complex (from this work), tri-snRNP26,29 and B-complex structures8. In this model the tri-snRNP that associates with the A-complex already contains the Brr2 helicase bound to the U4 snRNA substrate and the yeast B-complex proteins at the Prp8 N-terminal domain. The tri-snRNP then binds the A-complex (transition I, ‘Assembly’), requiring a substantial readjustment to avoid a steric clash of the Brr2-containing ‘helicase’ domain and the U1 snRNP (‘U1-B-complex’). The Prp28 helicase is then recruited to the U1 snRNP directly as the Prp28-binding site on the Prp8 N-terminal domain in human tri-snRNP is occupied by B-complex proteins25. Prp28 then disrupts the U1–5′SS helix, leading to 5′SS transfer (transition II, ‘Transfer’). Similar to the ‘pre-B-complex’ assembly model in a, the U1 snRNP, now freed from the 5′SS, may then be released owing to a steric clash with the Brr2-containing ‘helicase’ domain. This model does not require Sad1. Compare to a.
Extended Data Fig. 9 Data collection, refinement statistics and validation.
a, Cryo-EM data collection and refinement statistics of the A-complex structure. Maps A1 and A3 were used to position the U2 snRNP 3′ and 5′ regions, respectively. b. FSC between the A2 cryo-EM density and the refined A-complex U1 snRNP coordinate model.
Supplementary information
Supplementary Figure 1
Uncropped gels shown in Extended Data Figure 1a (a) and 1b (b).
Supplementary Data
PyMol session of the A complex structure (PDB 6G90). Subunits are coloured as in Figure 1.
Video 1: 360° rotation of the A complex coordinate model (PDB 6G90).
Coloured as in Figure 1.
Video 2: 360° rotation of the composite A complex cryo-EM density.
A1, grey, EMD-4363; A2, cyan, EMD-4364; A3, green, EMD-4365. Coloured as in Extended Data Figure 1e.
Rights and permissions
About this article
Cite this article
Plaschka, C., Lin, PC., Charenton, C. et al. Prespliceosome structure provides insights into spliceosome assembly and regulation. Nature 559, 419–422 (2018). https://doi.org/10.1038/s41586-018-0323-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-018-0323-8
This article is cited by
-
Structural basis of human U5 snRNP late biogenesis and recycling
Nature Structural & Molecular Biology (2024)
-
Co-transcriptional gene regulation in eukaryotes and prokaryotes
Nature Reviews Molecular Cell Biology (2024)
-
Integrated analysis of transcriptome and small RNAome reveals regulatory network of rapid and long-term response to heat stress in Rhododendron moulmainense
Planta (2024)
-
Regulation of pre-mRNA splicing: roles in physiology and disease, and therapeutic prospects
Nature Reviews Genetics (2023)
-
Alternative splicing plays key roles in response to stress across different stages of fighting in the fish Betta splendens
BMC Genomics (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.