Abstract
The movement of core-lipopolysaccharide across the inner membrane of Gram-negative bacteria is catalysed by an essential ATP-binding cassette transporter, MsbA. Recent structures of MsbA and related transporters have provided insights into the molecular basis of active lipid transport; however, structural information about their pharmacological modulation remains limited. Here we report the 2.9 Å resolution structure of MsbA in complex with G907, a selective small-molecule antagonist with bactericidal activity, revealing an unprecedented mechanism of ABC transporter inhibition. G907 traps MsbA in an inward-facing, lipopolysaccharide-bound conformation by wedging into an architecturally conserved transmembrane pocket. A second allosteric mechanism of antagonism occurs through structural and functional uncoupling of the nucleotide-binding domains. This study establishes a framework for the selective modulation of ABC transporters and provides rational avenues for the design of new antibiotics and other therapeutics targeting this protein family.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Brown, E. D. & Wright, G. D. Antibacterial drug discovery in the resistance era. Nature 529, 336–343 (2016).
Laxminarayan, R. et al. Antibiotic resistance—the need for global solutions. Lancet Infect. Dis. 13, 1057–1098 (2013).
Nikaido, H. Molecular basis of bacterial outer membrane permeability revisited. Microbiol. Mol. Biol. Rev. 67, 593–656 (2003).
Raetz, C. R. H. & Whitfield, C. Lipopolysaccharide endotoxins. Annu. Rev. Biochem. 71, 635–700 (2002).
Sperandeo, P. et al. Characterization of lptA and lptB, two essential genes implicated in lipopolysaccharide transport to the outer membrane of Escherichia coli. J. Bacteriol. 189, 244–253 (2007).
Zhou, Z., White, K. A., Polissi, A., Georgopoulos, C. & Raetz, C. R. Function of Escherichia coli MsbA, an essential ABC family transporter, in lipid A and phospholipid biosynthesis. J. Biol. Chem. 273, 12466–12475 (1998).
Doerrler, W. T., Reedy, M. C. & Raetz, C. R. An Escherichia coli mutant defective in lipid export. J. Biol. Chem. 276, 11461–11464 (2001).
Davidson, A. L., Dassa, E., Orelle, C. & Chen, J. Structure, function, and evolution of bacterial ATP-binding cassette systems. Microbiol. Mol. Biol. Rev. 72, 317–364 (2008).
Mi, W. et al. Structural basis of MsbA-mediated lipopolysaccharide transport. Nature 549, 233–237 (2017).
Ward, A., Reyes, C. L., Yu, J., Roth, C. B. & Chang, G. Flexibility in the ABC transporter MsbA: alternating access with a twist. Proc. Natl Acad. Sci. USA 104, 19005–19010 (2007).
Lee, S. C. et al. Steroid-based facial amphiphiles for stabilization and crystallization of membrane proteins. Proc. Natl Acad. Sci. USA 110, E1203–E1211 (2013).
Locher, K. P. Mechanistic diversity in ATP-binding cassette (ABC) transporters. Nat. Struct. Mol. Biol. 23, 487–493 (2016).
Jin, M. S., Oldham, M. L., Zhang, Q. & Chen, J. Crystal structure of the multidrug transporter P-glycoprotein from Caenorhabditis elegans. Nature 490, 566–569 (2012).
Zhang, Y., Rempel, D. L., Zhang, H. & Gross, M. L. An improved fast photochemical oxidation of proteins (FPOP) platform for protein therapeutics. J. Am. Soc. Mass Spectrom. 26, 526–529 (2015).
Esser, L. et al. Structures of the multidrug transporter P-glycoprotein reveal asymmetric ATP binding and the mechanism of polyspecificity. J. Biol. Chem. 292, 446–461 (2017).
Gao, M. et al. Comparison of the functional characteristics of the nucleotide binding domains of multidrug resistance protein 1. J. Biol. Chem. 275, 13098–13108 (2000).
Procko, E., Ferrin-O’Connell, I., Ng, S. L. & Gaudet, R. Distinct structural and functional properties of the ATPase sites in an asymmetric ABC transporter. Mol. Cell 24, 51–62 (2006).
Sorum, B., Töröcsik, B. & Csanády, L. Asymmetry of movements in CFTR’s two ATP sites during pore opening serves their distinct functions. eLife 6, e29013 (2017).
Nöll, A. et al. Crystal structure and mechanistic basis of a functional homolog of the antigen transporter TAP. Proc. Natl Acad. Sci. USA 114, E438–E447 (2017).
Martin, G. M., Kandasamy, B., DiMaio, F., Yoshioka, C. & Shyng, S. L. Anti-diabetic drug binding site in a mammalian KATP channel revealed by cryo-EM. eLife 6, e31054 (2017).
Perez, C. et al. Structural basis of inhibition of lipid-linked oligosaccharide flippase PglK by a conformational nanobody. Sci. Rep. 7, 46641 (2017).
Ray, B. L. & Raetz, C. R. The biosynthesis of gram-negative endotoxin. A novel kinase in Escherichia coli membranes that incorporates the 4′-phosphate of lipid A. J. Biol. Chem. 262, 1122–1128 (1987).
Garrett, T. A., Kadrmas, J. L. & Raetz, C. R. Identification of the gene encoding the Escherichia coli lipid A 4′-kinase. Facile phosphorylation of endotoxin analogs with recombinant LpxK. J. Biol. Chem. 272, 21855–21864 (1997).
Baldridge, R. D. & Graham, T. R. Identification of residues defining phospholipid flippase substrate specificity of type IV P-type ATPases. Proc. Natl Acad. Sci. USA 109, E290–E298 (2012).
Brunner, J. D., Lim, N. K., Schenck, S., Duerst, A. & Dutzler, R. X-ray structure of a calcium-activated TMEM16 lipid scramblase. Nature 516, 207–212 (2014).
Zoonens, M. & Popot, J.-L. Amphipols for each season. J. Membr. Biol. 247, 759–796 (2014).
Bayburt, T. H. & Sligar, S. G. Membrane protein assembly into nanodiscs. FEBS Lett. 584, 1721–1727 (2010).
Diederich, L., Rasmussen, L. J. & Messer, W. New cloning vectors for integration in the lambda attachment site attB of the Escherichia coli chromosome. Plasmid 28, 14–24 (1992).
Datsenko, K. A. & Wanner, B. L. One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc. Natl Acad. Sci. USA 97, 6640–6645 (2000).
Sampson, B. A., Misra, R. & Benson, S. A. Identification and characterization of a new gene of Escherichia coli K-12 involved in outer membrane permeability. Genetics 122, 491–501 (1989).
Baba, T. et al. Construction of Escherichia coli K-12 in-frame, single-gene knockout mutants: the Keio collection. Mol. Syst. Biol. 2, 2006.0008 (2006).
Wayne, P. A. Performance Standards for Antimicrobial Susceptibility Testing; Sixteenth Informational Supplement. CLSI document M100-S16CLSI. (Clinical and Laboratory Standards Institute, Wayne, 2007).
Wu, T. D. & Nacu, S. Fast and SNP-tolerant detection of complex variants and splicing in short reads. Bioinformatics 26, 873–881 (2010).
Lawrence, M., Degenhardt, J. & Gentleman, R. VariantTools: Tools for Working with Genetic Variants. R package v.1.20.0 (2016).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
Collaborative Computational Project, Number 4. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D 50, 760–763 (1994).
Chaptal, V. et al. Fluorescence detection of heavy atom labeling (FD–HAL): a rapid method for identifying covalently modified cysteine residues by phasing atoms. J. Struct. Biol. 171, 82–87 (2010).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, (213–221 (2010).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010).
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D 66, 12–21 (2010).
Kleywegt, G. J. Use of non-crystallographic symmetry in protein structure refinement. Acta Crystallogr. D 52, 842–857 (1996).
The PyMOL Molecular Graphics System v.1.8 (Schrödinger, LLC., 2015).
Zhang, Y., Wecksler, A. T., Molina, P., Deperalta, G. & Gross, M. L. Mapping the binding interface of VEGF and a monoclonal antibody Fab-1 fragment with fast photochemical oxidation of proteins (FPOP) and mass spectrometry. J. Am. Soc. Mass Spectrom. 28, 850–858 (2017).
Li, J. et al. Mapping the energetic epitope of an antibody/interleukin-23 interaction with hydrogen/deuterium exchange, fast photochemical oxidation of proteins mass spectrometry, and alanine shave mutagenesis. Anal. Chem. 89, 2250–2258 (2017).
Madhavi Sastry, G., Adzhigirey, M., Day, T., Annabhimoju, R. & Sherman, W. Protein and ligand preparation: parameters, protocols, and influence on virtual screening enrichments. J. Comput. Aided Mol. Des. 27, 221–234 (2013).
Schrödinger Release 2017-1: Schrödinger Suite 2017-2 (Schrödinger, LLC, 2017).
Schrödinger Release 2017-1: Desmond Molecular Dynamics System (Schrödinger, LLC, 2017).
Bowers, K. J. et al. Scalable Algorithms for Molecular Dynamics Simulations on Commodity Clusters. In Proc. ACM/IEEE Conference on Supercomputing (SC06) (ed. Horner-Miller, B.) (ACM, 2006).
Shivakumar, D. et al. Prediction of absolute solvation free energies using molecular dynamics free energy perturbation and the OPLS force field. J. Chem. Theory Comput. 6, 1509–1519 (2010).
Guo, Z. et al. Probing the α-helical structural stability of stapled p53 peptides: molecular dynamics simulations and analysis. Chem. Biol. Drug Des. 75, 348–359 (2010).
Acknowledgements
We thank our colleagues for support, particularly E. Brown, S. Hymowitz, B. Roth, M. Starovasnik and W. Young. We also thank V. Janakiraman, M. McCreary, E. Skippington, and K. Toy for whole-genome sequencing and analysis. Studies were carried out at the Stanford Synchrotron Radiation Lightsource 12-12 at Stanford Linear Accelerator Center National Accelerator Laboratory; use of this facility is supported by the US Department of Energy (DOE), DOE Office of Biological and Environmental Research, National Institutes of Health (NIH), and National Institute of General Medical Sciences (NIGMS). Studies were also carried out at the Berkeley Center for Structural Biology, supported by the NIH, NIGMS, and the Howard Hughes Medical Institute. The Advanced Light Source is a DOE Office of Science User Facility, contract DE-AC02-05CH11231. Reagents are available under a materials transfer agreement with Genentech.
Reviewer information
Nature thanks E. Breukink, P. Nissen and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
H.Ho. and A.O. purified and reconstituted MsbA. H.Ho. crystallized MsbA complexes. H.Ho., A.M., A.O., J.Q., K.C., M.H.B., T.M. and Y.X. performed high-throughput screening and biochemical studies. M.K.A. and M.N. performed phenotypic studies. N.K.G. and A.T.W. performed HRF–MS experiments. H.Ho., I.Z., C.T., S.S. and Y.F. performed molecular biology and protein expression. M.R. and C.D.A. performed electron micrograph studies. H.Hu., S.S.L., J.L., L.W., J.W., Y.L., H.E.P., B.D.S., M.F.T.K., and V.V. performed chemical designs and syntheses. A.M., M.K.A., N.K.G., A.O., M.R., H.E.P., M.H.B., M.-W.T., B.D.S., J.R.K., A.T.W., V.V., Y.X., M.N., J.P. and C.M.K. analysed the data; B.D.S. performed MD simulations, J.P. determined X-ray structures with input from J.R.K.; J.P. and C.M.K. wrote the manuscript with input from all authors; J.P. and C.M.K. supervised the project and are co-senior authors.
Corresponding authors
Ethics declarations
Competing interests
All authors except J.W. and Y.L. are employees of Genentech; J.W. and Y.L. are employees of WuXi.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Evaluation of the biochemical activity of MsbA and inhibitors.
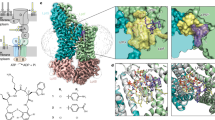
a, Quinoline inhibitors described in this study. b, Comparison of ATPase activity of increasing concentrations of E. coli MsbA in amphipol, nanodisc and FA-3 detergent matrices. c, d, Dose–response curves of compounds on purified Klebsiella pneumoniae (c) and Pseudomonas aeruginosa (d) MsbA reconstituted in amphipols. e, f, Dose–response curves of compounds on purified E. coli MsbA reconstituted in nanodiscs (e) or FA-3 detergent (f). g, h, Dose–response curves of compounds on purified E. coli A175V mutant MsbA reconstituted in amphipols (g) and nanodiscs (h). i, The LPS content of purified E. coli MsbA in the indicated matrices was measured using a chromogenic endotoxin quantitation assay (see Methods section ‘Quantitation of co-purifying LPS’). LPS from E. coli strain O111:B4 was used to generate a standard curve. Note that for both MsbA samples (0.0024 ng ml−1), the concentration of LPS (more than 0.02 ng ml−1) exceeds the concentration of MsbA, indicating a molar excess of co-purifying LPS. Data are mean ± s.e.m. from five independent experiments (b), three independent experiments (c–h) or two independent experiments (i). IC50 values (b–h, in parentheses) were determined by fitting the inhibition dose–response curve with a nonlinear four-parameter inhibition model (see Methods section ‘MsbA ATPase assay’).
Extended Data Fig. 2 Evaluation of phenotypic activity of MsbA inhibitors.
a, α-MsbA western blot of solubilized extracts from an E. coli CFT073 UPEC imp strain overexpressing MsbA from a pBAD24 vector in the presence of 2% arabinose (lane 1), an E. coli CFT073 imp strain (lane 2), an E. coli CFT073 imp strain with the arabinose-titratable msbA expression system in the presence of 2% arabinose (lane 3), an E. coli CFT073 imp strain with the arabinose-titratable msbA expression system in the presence of 0.01% arabinose (lane 4) and an E. coli CFT073 imp strain with the arabinose-titratable msbA expression system in the presence of 0.002% arabinose (lane 5). Bacteria were grown for 3 h at 37 °C before collection. The blot is representative of n = 2 independent experiments. For gel source images, see Supplementary Fig. 1. b, Dose–response curves of compounds on E. coli CFT073 imp strains expressing endogenous (WT) and overexpressed (HIGH, in the presence of 0.2% arabinose) levels of MsbA. Data are mean ± s.e.m. from two independent experiments except for G247 (n = 4). IC50 values (in parentheses) were determined by fitting the inhibition dose–response curve with a nonlinear four-parameter inhibition model (see Methods section ‘MsbA ATPase assay’). c, Representative (20 isolated cells from two independent experiments) thin section electron micrographs of CFT073 lptD(imp4213) and CFT073 wild-type strains. Treatment of the lptD(imp4213) strain with G592 (middle left) or G907 (middle right) and wild-type UPEC with G247 (bottom) leads to membrane stacking (red arrow) and enlargement of cells, similar to cells depleted of MsbA (top right). Scale bars, 0.1 μm. d, e, Nucleotide and amino acid sequence changes (indicated in red) in two G592-resistant MG1655 ΔtolC clones (mut1 and mut2, d) and a G247-resistant wild-type E. coli clone (mut3, e, data not shown). For both panels, positions are relative to the start of msbA ORF. The frequency of resistance for G247 on wild-type E. coli was 3.9 × 10−9 (data not shown).
Extended Data Fig. 3 Comparison of the G907–LPS–EcMsbA and G092–LPS–EcMsbA complexes.
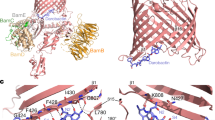
a, b, Views of the crystal packing environments in the G907–LPS–EcMsbA and G092–LPS–EcMsbA complexes show distinct crystal lattice contacts. c, The superposition of G907–LPS–EcMsbA and G092–LPS–EcMsbA complexes indicates a high degree of structural similarity, reflected in a Cα r.m.s.d. of 0.8 Å over all modelled residues. d, Chemical structure of G092. e, Dose–response curves of G092 on purified E. coli MsbA in FA-3 detergent or reconstituted into amphipols and on purified P. aeruginosa MsbA reconstituted into amphipols. Data are mean ± s.e.m. from three independent experiments. IC50 values (indicated) were determined by fitting the inhibition dose–response curve with a nonlinear four-parameter inhibition model (see Methods). f, Overall 2.92-Å crystal structure of the G092–LPS–EcMsbA complex shown from a side-view. LPS bound within the inner vestibule is shown in green sphere representation and G092 is shown in thin yellow stick representation. Fo − Fc difference electron density maps (calculated before the addition of G092 into the model and refinement) are contoured at 2σ (cyan) and 3σ (pink). g, Fo − Fc difference electron density maps of G092–LPS–EcMsbA (calculated before the addition of G092 into the model and refinement) are contoured at 2σ (cyan) and 3σ (pink). The final refined coordinates of G092 are shown for reference. h, As in g, but a side-view of the G092 binding site.
Extended Data Fig. 4 Molecular interactions of G907 in complex with EcMsbA.
a, b, The MOE (Molecular Operating Environment; Chemical Computing Group ULC) software was used to highlight the molecular interactions observed between G907 and MsbA, with two similar illustrations of this representation provided. c, A legend describing the characteristics of the interacting groups as well as the types of interactions shown in a and b.
Extended Data Fig. 5 A conserved small molecule or peptide transmembrane binding pocket in the MsbA and Pgp B-family ABC transporters.
a, Top, surface representation of the G907–LPS–EcMsbA crystal structure (subunit A, blue; subunit B, grey) with G907 in red sphere representation. LPS is omitted for clarity. Bottom, expanded view of the G907-binding site. b, Top, CePgp crystal structure in surface representation (N-terminal half, blue; C-terminal half, grey) with the extended N-terminal peptide (NTP) region of the transporter shown in red surface representation. Bottom, expanded view of the NTP-binding site. c, Similar to a, G907–LPS–EcMsbA crystal structure in surface representation (subunit A, blue; subunit B, grey) with G907 in red sphere representation, viewed approximately 180° relative to a to focus in on subunit B. LPS is omitted for clarity. Bottom, expanded view of the G907-binding site. d, Similar to a, CePgp crystal structure in surface representation (N-terminal half, blue; C-terminal half, grey) in which two modelled detergent molecules of n-undecyl-β-d-thiomaltopyranoside (UDTM) are shown in red sphere representation. Bottom, expanded view of the UDTM-binding site.
Extended Data Fig. 6 Unique relative displacement of NBDs in both the G907–LPS–EcMsbA and G092–LPS–EcMsbA complex crystal structures.
a–e, Structural superpositions of related B-family ABC transporters are provided in two orthogonal views, in which all structural superpositions were performed by aligning only the transmembrane portion of each subunit B protomer onto that of subunit A. In the case of CePgp, the C-terminal transmembrane region was superimposed onto the N-terminal transmembrane region. The structures are as follows: a, G907–LPS–EcMsbA; b, G092–LPS–EcMsbA; c, LPS–EcMsbA–cryo (PDB accession 5TV4); d, CePgp (PDB accession 4F4C); e, Thermus thermophilus TmrAB (PDB accession 5MKK). In comparing these structures, it is noteworthy that EcMsbA is a homodimeric transporter, whereas CePgp is a concatenated transporter, and TmrAB is a heterodimeric transporter; yet the structural similarity within all ‘self-pairs’ of transmembrane regions is high. Specifically, when excluding the NBDs from this analysis: a, Cα r.m.s.d. 1.26 Å; b, Cα r.m.s.d. 0.82 Å; c, Cα r.m.s.d. 1.09 Å; d, Cα r.m.s.d. 3.18 Å; e, Cα r.m.s.d. 1.66 Å. In contrast to the high structural correspondence observed between their superimposed or ‘self-paired’ transmembrane regions, the G907–LPS–EcMsbA and G092–LPS–EcMsbA complex structures both show a marked translational displacement of one NBD relative to the other, measuring up to 15 Å between equivalent Cα positions. It is notable that the extent and apparent vector of the structural translation observed at the level of the NBDs are unique to the inhibitor-bound EcMsbA complexes studied here. For reference, the three residues that constitute the highly conserved CH1–CH2–NBD coupling interaction network are shown in yellow sphere representation in all structures.
Extended Data Fig. 7 HRF–MS of LPS–EcMsbA in the presence and absence of G907 and G092.
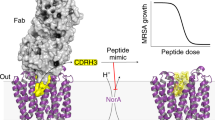
a, A schematic overview (with the target protein illustrated as a cartoon IgG shown in green) of the HRF–MS experiments performed in this study on LPS–EcMsbA in FA-3 detergent conditions in the presence or absence of G907 and G092. LC–MS/MS, liquid chromatography with tandem mass spectrometry; UPLC, ultra-performance liquid chromatography. b–c, Data are displayed as the differences measured in tryptic peptide oxidation of the ligand-bound condition with G907 present (b) or G092 present (c) relative to the inhibitor-free LPS–EcMsbA solution sample. The median residue number of the individual tryptic peptides (summarized in Extended Data Table 2 for the G907 experiment) is indicated on the x-axis. For both HRF–MS analyses, data are mean ± s.e.m. from three replicate experiments. d, A cartoon rendering of the G907–LPS–EcMsbA crystal structure coloured according to the measured differences in tryptic peptide oxidation between the inhibitor-free and G907-bound condition presented in b. Regions without significant measured changes are coloured in dark grey and are found mainly within the transmembrane region; regions lacking robust peptide coverage are coloured in light grey. The phospho-N-acetylglucosamine moieties of the bound LPS molecule are shown only for reference, the remainder of LPS and G907 are omitted for clarity.
Extended Data Fig. 8 Electron densities and electrostatic features of the LPS coordination site in the inner vestibule of the G092–LPS–EcMsbA crystal structure.
a, Fo − Fc difference electron density maps (calculated before the addition of LPS into the model and refinement) are contoured at 2.5σ (purple) and 5σ (pink). b, Fo − Fc difference electron density maps (calculated with LPS omitted from the model) contoured at 3σ (purple) and 5σ (pink). c, Fo − Fc difference electron density maps (calculated with LPS omitted from the model) contoured at 1σ (cyan) and 5σ (pink). a–c, The final refined coordinates of LPS are also shown for reference and the arginine side chains are omitted for clarity. The observed electron density positions the phospho-N-acetylglucosamine moieties of bound LPS within the conserved arginine coordination ring. d, e, Overall electrostatic surface representation of MsbA in the G092–LPS–EcMsbA complex with LPS shown in yellow stick representation and G092 omitted for clarity. For reference only, the position of the bulk membrane bilayer is depicted on the basis of a 10-ns molecular dynamics simulation (see Methods section ‘Molecular dynamics’). f, Similar to d, e, expanded view with the closest MsbA subunit removed for clarity and LPS shown in yellow stick representation. g, Similar to f, expanded view from the cytoplasmic perspective, where both MsbA subunits are displayed.
Supplementary information
Supplementary Information
This file contains: Supplementary Data on the discovery and biochemical and phenotypic characterization of selective MsbA inhibitors and the crystallization of MsbA in complex with G907 and G092; Supplementary Methods on the synthesis, structures and characterization of chemical materials; Supplementary Tables 1-2 and Supplementary Figure 1, original scans (white light and luminescence) of the western blot used to generate Extended Data Fig. 2a.
Rights and permissions
About this article
Cite this article
Ho, H., Miu, A., Alexander, M.K. et al. Structural basis for dual-mode inhibition of the ABC transporter MsbA. Nature 557, 196–201 (2018). https://doi.org/10.1038/s41586-018-0083-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-018-0083-5
This article is cited by
-
Probing the allosteric NBD-TMD crosstalk in the ABC transporter MsbA by solid-state NMR
Communications Biology (2024)
-
A new antibiotic traps lipopolysaccharide in its intermembrane transporter
Nature (2024)
-
Cerastecins inhibit membrane lipooligosaccharide transport in drug-resistant Acinetobacter baumannii
Nature Microbiology (2024)
-
Structural basis of peptide secretion for Quorum sensing by ComA
Nature Communications (2023)
-
Tackling the outer membrane: facilitating compound entry into Gram-negative bacterial pathogens
npj Antimicrobials and Resistance (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.