Abstract
Nutrient avidity is one of the most distinctive features of tumours. However, nutrient deprivation has yielded limited clinical benefits. In Gaucher disease, an inherited metabolic disorder, cells produce cholesteryl-glucoside which accumulates in lysosomes and causes cell damage. Here we develop a nanoparticle (AbCholB) to emulate natural-lipoprotein-carried cholesterol and initiate Gaucher disease-like damage in cancer cells. AbCholB is composed of a phenylboronic-acid-modified cholesterol (CholB) and albumin. Cancer cells uptake the nanoparticles into lysosomes, where CholB reacts with glucose and generates a cholesteryl-glucoside-like structure that resists degradation and aggregates into microscale crystals, causing Gaucher disease-like damage in a glucose-dependent manner. In addition, the nutrient-sensing function of mTOR is suppressed. It is observed that normal cells escape severe damage due to their inferior ability to compete for nutrients compared with cancer cells. This work provides a bioinspired strategy to selectively impede the metabolic action of cancer cells by taking advantage of their nutrient avidity.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The data supporting the findings of this study are provided in the article or Supplementary Information. The unprocessed raw data of RNA-seq are available from the corresponding author on reasonable request. Source data are provided with this paper.
References
Bartman, C. R. et al. Slow TCA flux and ATP production in primary solid tumours but not metastases. Nature 614, 349–357 (2023).
Ringel, A. E. et al. Obesity shapes metabolism in the tumor microenvironment to suppress anti-tumor immunity. Cell 183, 1848–1866 (2020).
Faubert, B., Solmonson, A. & DeBerardinis, R. J. Metabolic reprogramming and cancer progression. Science 368, eaaw5473 (2020).
Kanarek, N., Petrova, B. & Sabatini, D. M. Dietary modifications for enhanced cancer therapy. Nature 579, 507–517 (2020).
Kim, J. & Guan, K.-L. mTOR as a central hub of nutrient signalling and cell growth. Nat. Cell Biol. 21, 63–71 (2019).
Vernieri, C. et al. Fasting-mimicking diet is safe and reshapes metabolism and antitumor immunity in patients with cancer. Cancer Discov. 12, 90–107 (2022).
Vaziri-Gohar, A. et al. Limited nutrient availability in the tumor microenvironment renders pancreatic tumors sensitive to allosteric IDH1 inhibitors. Nat. Cancer 3, 852–865 (2022).
Yao, J. C. et al. Everolimus for advanced pancreatic neuroendocrine tumors. N. Engl. J. Med. 364, 514–523 (2011).
Chang, C.-H. et al. Metabolic competition in the tumor microenvironment is a driver of cancer progression. Cell 162, 1229–1241 (2015).
Nencioni, A., Caffa, I., Cortellino, S. & Longo, V. D. Fasting and cancer: molecular mechanisms and clinical application. Nat. Rev. Cancer 18, 707–719 (2018).
Spotten, L. E. et al. Subjective and objective taste and smell changes in cancer. Ann. Oncol. 28, 969–984 (2017).
Kir, S. et al. Tumour-derived PTH-related protein triggers adipose tissue browning and cancer cachexia. Nature 513, 100–104 (2014).
Biswas, A. K. & Acharyya, S. Understanding cachexia in the context of metastatic progression. Nat. Rev. Cancer 20, 274–284 (2020).
Marine, J. C., Dawson, S. J. & Dawson, M. A. Non-genetic mechanisms of therapeutic resistance in cancer. Nat. Rev. Cancer 20, 743–756 (2020).
McIntyre, A. & Harris, A. L. Metabolic and hypoxic adaptation to anti-angiogenic therapy: a target for induced essentiality. EMBO Mol. Med. 7, 368–379 (2015).
García-Jiménez, C. & Goding, C. R. Starvation and pseudo-starvation as drivers of cancer metastasis through translation reprogramming. Cell Metab. 29, 254–267 (2019).
Barton, M. K. Cancer cachexia awareness, diagnosis, and treatment are lacking among oncology providers. CA Cancer J. Clin. 67, 91–92 (2017).
Pandey, M. K. et al. Complement drives glucosylceramide accumulation and tissue inflammation in Gaucher disease. Nature 543, 108–112 (2017).
Mistry, P. K. et al. Effect of oral eliglustat on splenomegaly in patients with Gaucher disease type 1: the ENGAGE randomized clinical trial. J. Am. Med. Assoc. 313, 695–706, (2015).
Akiyama, H., Kobayashi, S., Hirabayashi, Y. & Murakami-Murofushi, K. Cholesterol glucosylation is catalyzed by transglucosylation reaction of β-glucosidase 1. Biochem. Biophys. Res. Commun. 441, 838–843 (2013).
Aerts, J. M. F. G. et al. Glycosphingolipids and lysosomal storage disorders as illustrated by Gaucher disease. Curr. Opin. Chem. Biol. 53, 204–215 (2019).
Surface, M. et al. Plasma glucosylsphingosine in GBA1 mutation carriers with and without Parkinson’s disease. Mov. Disord. 37, 416–421 (2022).
Akiyama, H. et al. Glucocerebrosidases catalyze a transgalactosylation reaction that yields a newly-identified brain sterol metabolite, galactosylated cholesterol. J. Biol. Chem. 295, 5257–5277 (2020).
Franco, R., Sánchez-Arias, J. A., Navarro, G. & Lanciego, J. L. Glucocerebrosidase mutations and synucleinopathies. Potential role of sterylglucosides and relevance of studying both GBA1 and GBA2 genes. Front. Neuroanat. 12, 52 (2018).
Shimamura, M. Structure, metabolism and biological functions of steryl glycosides in mammals. Biochem. J. 477, 4243–4261 (2020).
Marques, A. R. A. et al. Glucosylated cholesterol in mammalian cells and tissues: formation and degradation by multiple cellular β-glucosidases. J. Lipid Res. 57, 451–463 (2016).
Kałużna, M., Trzeciak, I., Ziemnicka, K., Machaczka, M. & Ruchała, M. Endocrine and metabolic disorders in patients with Gaucher disease type 1: a review. Orphanet J. Rare Dis. 14, 275 (2019).
Karpa, M. J., Duggan, P. J., Griffin, G. J. & Freudigmann, S. J. Competitive transport of reducing sugars through a lipophilic membrane facilitated by aryl boron acids. Tetrahedron 53, 3669–3678 (1997).
Zhu, J.-Y. et al. Acidity-responsive gene delivery for “superfast” nuclear translocation and transfection with high efficiency. Biomaterials 83, 79–92 (2016).
Xiao, H., Chen, W., Smeekens, J. M. & Wu, R. An enrichment method based on synergistic and reversible covalent interactions for large-scale analysis of glycoproteins. Nat. Commun. 9, 1692 (2018).
Bakh, N. A. et al. Glucose-responsive insulin by molecular and physical design. Nat. Chem. 9, 937–944 (2017).
Riscal, R., Skuli, N. & Simon, M. C. Even cancer cells watch their cholesterol! Mol. Cell 76, 220–231 (2019).
Curry, S., Mandelkow, H., Brick, P. & Franks, N. Crystal structure of human serum albumin complexed with fatty acid reveals an asymmetric distribution of binding sites. Nat. Struct. Biol. 5, 827–835 (1998).
Huang, P. et al. Cellular cholesterol directly activates Smoothened in Hedgehog signaling. Cell 166, 1176–1187 (2016).
Castellano, B. M. et al. Lysosomal cholesterol activates mTORC1 via an SLC38A9–Niemann–Pick C1 signaling complex. Science 355, 1306–1311 (2017).
Cortes, J. et al. Pembrolizumab plus chemotherapy in advanced triple-negative breast cancer. N. Engl. J. Med. 387, 217–226 (2022).
Birn, H. et al. Cubilin is an albumin binding protein important for renal tubular albumin reabsorption. J. Clin. Investig. 105, 1353–1361 (2000).
Wu, Q., Wang, L., Yu, H., Wang, J. & Chen, Z. Organization of glucose-responsive systems and their properties. Chem. Rev. 111, 7855–7875 (2011).
McDonald, O. G. et al. Epigenomic reprogramming during pancreatic cancer progression links anabolic glucose metabolism to distant metastasis. Nat. Genet. 49, 367–376 (2017).
Han, H.-J., Russo, J., Kohwi, Y. & Kohwi-Shigematsu, T. SATB1 reprogrammes gene expression to promote breast tumour growth and metastasis. Nature 452, 187–193 (2008).
Luo, J., Yang, H. & Song, B.-L. Mechanisms and regulation of cholesterol homeostasis. Nat. Rev. Mol. Cell Biol. 21, 225–245 (2020).
Baixauli, F. et al. Mitochondrial respiration controls lysosomal function during inflammatory T cell responses. Cell Metab. 22, 485–498 (2015).
Abu-Asab, M. S. et. al. in Advances in Vision Research Vol. I Essentials in Ophthalmology (eds Prakash, G. & Iwata, T.) 413–423 (Springer, 2017).
Zigdon, H. et al. Altered lysosome distribution is an early neuropathological event in neurological forms of Gaucher disease. FEBS Lett. 591, 774–783 (2017).
Uemura, N. et al. Viable neuronopathic Gaucher disease model in medaka (Oryzias latipes) displays axonal accumulation of alpha-synuclein. PLoS Genet. 11, e1005065 (2015).
Platt, F. M. Sphingolipid lysosomal storage disorders. Nature 510, 68–75 (2014).
Cantuti-Castelvetri, L. et al. Defective cholesterol clearance limits remyelination in the aged central nervous system. Science 359, 684–688 (2018).
Amaravadi, R. K., Kimmelman, A. C. & Debnath, J. Targeting autophagy in cancer: recent advances and future directions. Cancer Discov. 9, 1167–1181 (2019).
Galluzzi, L. & Green, D. R. Autophagy-independent functions of the autophagy machinery. Cell 177, 1682–1699 (2019).
Kimmey, J. M. et al. Unique role for ATG5 in neutrophil-mediated immunopathology during M. tuberculosis infection. Nature 528, 565–569 (2015).
Martina, J. A., Raben, N. & Puertollano, R. SnapShot: lysosomal storage diseases. Cell 180, 602–602 (2020).
De Leo, M. G. et al. Autophagosome–lysosome fusion triggers a lysosomal response mediated by TLR9 and controlled by OCRL. Nat. Cell Biol. 18, 839–850 (2016).
Abu-Remaileh, M. et al. Lysosomal metabolomics reveals V-ATPase- and mTOR-dependent regulation of amino acid efflux from lysosomes. Science 358, 807–813 (2017).
Liu, G. Y. & Sabatini, D. M. mTOR at the nexus of nutrition, growth, ageing and disease. Nat. Rev. Mol. Cell Biol. 21, 183–203 (2020).
Jouandin, P. et al. Lysosomal cystine mobilization shapes the response of TORC1 and tissue growth to fasting. Science 375, eabc4203 (2022).
Shin, H. R. et al. Lysosomal GPCR-like protein LYCHOS signals cholesterol sufficiency to mTORC1. Science 0, eabg6621 (2022).
Rogala, K. B. et al. Structural basis for the docking of mTORC1 on the lysosomal surface. Science 366, 468–475 (2019).
de Araujo, M. E. G. et al. Crystal structure of the human lysosomal mTORC1 scaffold complex and its impact on signaling. Science 358, 377–381 (2017).
Pineda, CarlosT. et al. Degradation of AMPK by a cancer-specific ubiquitin ligase. Cell 160, 715–728 (2015).
Wyant, G. A. et al. mTORC1 activator SLC38A9 is required to efflux essential amino acids from lysosomes and use protein as a nutrient. Cell 171, 642–654 (2017).
Bockaert, J. & Marin, P. mTOR in Brain Physiology and Pathologies. Physiol. Rev. 95, 1157–1187 (2015).
Yu, L. et al. Termination of autophagy and reformation of lysosomes regulated by mTOR. Nature 465, 942–946 (2010).
van Gool, R. et al. Targeting neurological abnormalities in lysosomal storage diseases. Trends Pharmacol. Sci. 43, 495–509 (2022).
Cang, C. et al. mTOR regulates lysosomal ATP-sensitive two-pore Na(+) channels to adapt to metabolic state. Cell 152, 778–790 (2013).
Schwörer, S., Vardhana, S. A. & Thompson, C. B. Cancer Metabolism Drives a Stromal Regenerative Response. Cell Metab. 29, 576–591 (2019).
Kannauje, P. K., Pandit, V., Wasnik, P. N., Gupta, A. K. & Venkatesan, N. Gaucher’s Disease in an Adult Female: A Rare Entity. Cureus 13, e17318 (2021).
Hu, Y. et al. Methods for drug delivery comprising unfolding and refolding proteins and peptide nanoparticles. World Intellectual Property Organization. International application published with the international search report, patent WO2011019585A1 (2010).
Yue, C. et al. Long-term and liver-selected ginsenoside C-K nanoparticles retard NAFLD progression by restoring lipid homeostasis. Biomaterials 301, 122291 (2023).
Spandidos, A., Wang, X., Wang, H. & Seed, B. PrimerBank: a resource of human and mouse PCR primer pairs for gene expression detection and quantification. Nucleic Acids Res. 38, D792–D799 (2009).
Afgan, E. et al. The Galaxy platform for accessible, reproducible and collaborative biomedical analyses: 2018 update. Nucleic Acids Res. 46, W537–W544 (2018).
Xia, J., Gill, E. E. & Hancock, R. E. W. NetworkAnalyst for statistical, visual and network-based meta-analysis of gene expression data. Nat. Protoc. 10, 823–844 (2015).
Yuan, M., Breitkopf, S. B., Yang, X. & Asara, J. M. A positive/negative ion-switching, targeted mass spectrometry–based metabolomics platform for bodily fluids, cells, and fresh and fixed tissue. Nat. Protoc. 7, 872–881 (2012).
Giacomoni, F. et al. Workflow4Metabolomics: a collaborative research infrastructure for computational metabolomics. Bioinformatics 31, 1493–1495 (2014).
Gonzalez, P. S. et al. Mannose impairs tumour growth and enhances chemotherapy. Nature 563, 719–723 (2018).
Tang, Z., Kang, B., Li, C., Chen, T. & Zhang, Z. GEPIA2: an enhanced web server for large-scale expression profiling and interactive analysis. Nucleic Acids Res. 47, W556–W560 (2019).
Acknowledgements
This work was supported by the National Natural Science Foundation of China (82204293), the Collaborative Innovation Project of Yangtze River Delta Science and Technology Community (2023CSJZN0800) and the Logistics Research Program (BWS20J017). We thank P. Li for manuscript advice, Y. Qiu for grammar editing, H. Guo for help with TEM detection, and H. Wang, C. Wang, Y. Cheng and Y. Liu for advice on the introduction.
Author information
Authors and Affiliations
Contributions
Y.H. conceived the study. C.Y. and Y.H. designed the study. C.Y. conducted cell growth, GLUT1 and LDLR knockdown, movie making, immunocytochemistry assays, western blotting, all confocal imaging experiments, RNA-seq and metabonomics analyses and all statistical analyses. C.Y., W.L., S.F. and D.L. conducted the preparation and characterization of AbCholB and AbChol, and flow cytometry analysis. Z.H. synthesized CholB and conducted 1H NMR analysis. H.D., W.L., X.Z. and S.F. performed HPLC detection. W.L. and S.F. performed TLC analysis. T.D. and Y.S. downloaded and analysed TCGA data. W.L. conducted MALDI–TOF MS and LC–MS/MS scans. W.L., S.F., J.Y. and D.L. performed animal experiments. C.Y. and Y.H wrote the manuscript. A.Y., J.W. and L.K. revised the manuscript. All authors read and edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Nanotechnology thanks Dong-Bing Cheng, Chi (V) Dang and Ming Tan for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Characteristic features of AbCholB.
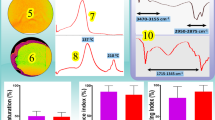
a, 1H-NMR showing the molecular structure of CholB. 1H-NMR (500 MHz, CDCl3): δ 8.02 – 7.74 (m, 2H), 7.57 – 7.28 (m, 2H), 6.69 (d, J = 99.9 Hz, 1H), 5.52 – 5.36 (m, 1H), 4.67 (tt, J = 11.1, 4.5 Hz, 1H), 2.59 – 2.35 (m, 2H), 2.10 – 1.82 (m, 6H), 1.67 – 1.47 (m, 8H), 1.41 – 1.34 (m, 3H), 1.28 (dq, J = 7.0, 3.8 Hz, 4H), 1.16 (tt, J = 19.7, 7.2 Hz, 6H), 1.07 (d, J = 6.5 Hz, 3H), 1.01 (dd, J = 11.1, 5.1 Hz, 2H), 0.94 (dd, J = 6.7, 4.3 Hz, 3H), 0.90 – 0.88 (m, 3H), 0.71 (q, J = 4.7, 3.1 Hz, 3H). b, TLC analysis for Chol and CholB. TLC analysis was developed in petroleum ether/ethyl acetate (1/1, vol/vol) and visualized with iodine vapour. c, d, In situ MALDI–TOF–MS analysis showing the binding of CholB and glucose (Glu) based on a typical ion (m/z = 568.19, M + NH4) of CholB (c) and a typical ion (m/z = 716.48, M+Na) of GlcCholB complex (d). e, Morphologies of AbChol under TEM. n = 3 independent experiments. f, g, Dynamic light scattering (DLS) revealing the size of AbCholB (f) and AbChol (g). Right top was the aqueous solution of AbCholB (f) and AbChol (g) at 5 mM. h, Diameter changes of AbCholB dispersed in saline with a concentration of 50 μM.
Extended Data Fig. 2 Uptake of AbCholB could be inhibited by LDLR knockdown.
a, A higher magnification view of Fig. 1e with bright field. Scale bar, 10 μm. b, FCM analysis showing the uptake of Dil-AbChol (left) or Dil-AbChoB (middle) in DU145 cells treated with or without LDL blockade. The histogram (right) was quantitative MFI of FCM data. n = 3 replicates. Mean ± s.e.m. Paired two-tailed Student’s t-test as indicated (right). c, WB analysis for LDLR knockdown in MDA-MB-231 cells. n = 3 independent experiments. d, Representative fluorescence confocal images of Dil-AbCholB (upper) or Dil-AbChol (lower) uptake in MDA-MB-231 cells with or without LDLR knockdown as indicated. Scale bar, 20 μm. n = 3. Cells were incubated with Dil-AbCholB or Dil-AbChol for 4 h before collection. e, FCM analysis showing the uptake of Dil-AbChol (left) or Dil-AbCholB (middle) in MDA-MB-231 cells treated with or without LDLR knockdown. The histogram (right) was quantitative MFI of FCM data. n = 3 replicates. Mean ± s.e.m. Paired two-tailed Student’s t-test as indicated (right).
Extended Data Fig. 3 AbCholB inhibits the proliferation of multiple cancer cells.
a-e, Growth curves showing the effect of AbCholB on proliferation of breast cancer MDA-MB-231 cells (a), prostate cancer DU145 cells (b), pancreas cancer Hs766T cells (c), lung cancer A549 cells (d) and normal breast MCF10A cells (e). All cells were cultured for 48 h in the presence of AbChol (grey), Ph-B+AbChol (blue) and AbCholB (magenta) with a series of gradient concentrations. MTT analysis was performed on 6 replicates. Mean ± s.e.m. Two-way ANOVA with Dunnett’s multiple comparisons test. *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001. From low concentration to high concentration, the exact P values in (a) were 0.2276, 0.9973, 0.0123, 0.0001, 4.637 x 10-9, 6.337 x 10-7, 2.364 x 10-10, 2.8 x 10-14 and 2.8 x 10-14. The exact P values in (b) were 0.0263, 0.9628, 0.7987, 0.962, 0.0007, 6.6 x 10-6, 7.791 x 10-11, 3.2 x 10-14 and 2.8 x 10-14. The exact P values in (c) were 0.9942, 0.9888, 0.4375, 0.0012, 7.634 x 10-5, 1.436 x 10-8, 3.4 x 10-14, 2.8 x 10-14 and 2.8 x 10-14. The exact P values in (d) were 0.967, 0.9371, 0.9532, 0.004, 0.0013, 8.342 x 10-5, 5.432 x 10-7, 1.01 x 10-13 and 2.8 x 10-14. The exact P values in (e) were 1, 0.9956, 1, 0.8813, 0.1854, 0.0209, 1.327 x 10-8, 2.8 x 10-14 and 2.8 x 10-14. f, Western blots showing GLUT1 levels in sh.Ctrl and sh.GLUT1 MDA-MB-231 cells. n = 3 independent experiments.
Extended Data Fig. 4 AbCholB affects cholesterol homeostasis in MDA-MB-231 cells.
a, PCA for untargeted metabolism analysis in AbCholB treated, Ph-B+AbChol treated and control MDA-MB-231 cells. Left for positive ion model and right for negative ion model. b, c, Relative levels of intermediates involved in cholesterol biosynthesis including farnesyl pyrophosphate (b) and squalene (c). n = 6 replicates. Minimum to maximum, 0–100. Centre line for the median (50th percentile). The box indicates the 25th-75th percentiles of dataset. Paired two-tailed Student’s t-test. d-l, Relative levels of representative metabolites involved in cholesterol metabolic progress including cholesteryl ester (d), steroid hormone biosynthetic intermediates (e to h), bile acid biosynthetic intermediates (i to l). n = 6 replicates. Minimum to maximum, 0–100. Centre line for the median (50th percentile). The box indicates the 25th-75th percentiles of dataset. Statistical analysis was conducted by paired two-tailed Student’s t-test. All cells were subjected to the indicated treatments for 48 h before metabolites extraction. LC-MS/MS analysis revealing the deficiency in cholesterol synthesis and accumulation of various cholesterol derivatives in the presence of AbCholB. Relative levels of metabolites expressed as normalized peak intensity. m, Model revealing the detailed position of differential intermediates in cholesterol metabolic progress.
Extended Data Fig. 5 AbCholB disturbs mRNA levels of genes enriched in glucose and cholesterol metabolisms.
a, PCA for RNA-seq in AbCholB treated, Ph-B+AbChol treated and control MDA-MB-231 cells. b-d, Volcano plot showing the significantly changed mRNAs between groups as follow: AbCholB vs Ctrl (b), AbCholB vs Ph-B + AbChol (c), Ph-B + AbChol vs Ctrl (d). All RNA-seq samples were collected after treatment for 48 h. n = 3 replicates. e-g, GSEA analysis showing depletion of glucose metabolic gene signatures, including glycolysis/gluconeogenesis (e), pentose phosphate pathway (f) and glycosaminoglycan degradation (g) in AbCholB treated MDA-MB-231 cells compared to Ph-B + AbChol treatment. h, GSEA analysis showing depletion of sterol homeostasis gene signatures in the presence of AbCholB compared to Ph-B + AbChol. i, GSEA analysis showing enrichment of sterol homeostasis gene signatures in Ph-B + AbChol treated cells compared to control cells. GO, gene ontology. j-l, RNA levels of glucose metabolic enzymes G6PD (j), PFKL (k), UGDH (l). G6PD: glucose-6-phosphate dehydrogenase, participates in pentose phosphate pathway; PFKL: phosphofructokinase, key enzyme in glycolysis; UGDH: UDP-glucose-6-dehydrogenase, participates in the biosynthesis of glycosaminoglycans. n = 3 replicates. Floating bar, minimum–maximum; the centre line indicates the mean. Statistical analysis was conducted by paired two-tailed Student’s t-test. m-r, RNA levels of cholesterol homeostasis regulatory factors SREBF2 (m), FDFT1 (n), SREBF1 (o), ABCA1 (p), ABCG1 (q) and SCD (r). n = 3 replicates. Floating bar, minimum–maximum; the centre line indicates the mean. Statistical analysis was conducted by paired two-tailed Student’s t-test. SREBF1/SREBF2: sterol regulatory element binding transcription factor 1/2, necessary for cholesterol and lipid homeostasis. FDFT1: farnesyl-diphosphate farnesyltransferase 1, key enzyme catalysing the sterol biosynthesis. ABCA1: ATP Binding Cassette Subfamily A Member 1, ABCG1: ATP Binding Cassette Subfamily G Member 1, ABCA1 and ABCG1 mediate cholesterol and lipid efflux. SCD: stearoyl-CoA desaturase, key enzyme involved in fatty acid biosynthesis. mRNA levels in (j-r) are from RNA-seq data.
Extended Data Fig. 6 GlcCholB formation and accumulation causes GD-like storage features in MDA-MB-231 cells.
a, FCM analysis for the uptake of Dil-AbChol (left) or Dil-AbCholB (middle) and the quantitative MFI histogram (right) in DU145 cells at 24 h, 48 h and 72 h. MFI was presented as mean ± s.e.m. n = 3 replicates. Statistical analysis was conducted by paired two-tailed Student’s t-test. b, GSEA showing depletion of lysosomal lumen gene signatures in AbCholB treated MDA-MB-231 cells compared to Ph-B+AbChol treated cells. c, Heatmap of lysosome lumen gene expressions based on GSEA analysis. n = 3 replicates. d, Representative TEM images of autolysosomes (black arrows) and MLB (orange arrows) in DU145 cells subjected to the indicated treatments for 48 h. e, Histogram of MLB counts from (d). MLBs were counted under at least 5 visual fields. Minimum to maximum, 0–100. Centre line for the median (50th percentile). 25th-75th percentiles of dataset in the box. Statistical analysis was conducted by one-way ANOVA. f, TLC analysis for the GlcCholB treated with GBA. GlcCholB was incubated with GBA for 16 h. TLC was developed with petroleum ether/ethyl acetate (1:1, vol/vol). g, GBA activity in shGLUT1 MDA-MB-231 cells subjected to the indicated treatments for 48 h. n = 3 replicates. Statistical analysis was conducted by paired Student’s two-tailed t-test. h, Western blots showing the protein level of caspase 3 (including cleaved-caspase3). n = 3 independent experiments. i, Representative TEM images with lower magnification for Fig. 4e. Autolysosomes (black arrows), MLB (orange arrows), mitochondria (yellow arrows).
Extended Data Fig. 7 AbCholB specifically accumulates in tumour tissues without significant systematic toxicity.
a, Tumour growth kinetics in 4T1-Luci-sh.GLUT1 implanted mice treated with saline and AbCholB. n = 9. Mean ± s.e.m. Statistical analysis was conducted by two-way ANOVA. b, Body weights during the treatment period. n = 9. Mean ± s.e.m. Statistical analysis was conducted by two-way ANOVA. ns: no significance. *P < 0.05. The exact P values were 0.0344 (AbChol vs Saline), 0.0748 (Ph-B+AbChol vs Saline) and 0.0603 (AbCholB vs Saline), respectively. c, Representative images of time-dependent in vivo fluorescence imaging in 4T1-Luci tumour bearing mice. Imaging was performed after tail-vein injection of DiD-AbCholB or DiD-AbChol for 2 h, 6 h, 12 h, 24 h and 48 h. d, Fluorescence images showing DiD-AbCholB distribution in tissues of heart, liver, spleen, lung, kidney, brain and tumour. n = 3. Scale bar, 200 μm. Tumour-bearing mice were sectioned and stained for DAPI after i.v. of DiD-AbCholB for 72 h. e, LC-MS analysis revealing the formation of GlcCholB in tumour treated with AbCholB. Specific peak of GlcCholB (m/z = 694.44, M + H) was only found in AbCholB treated tumour. f, PAS and H&E staining for liver tissues from mice subjected to indicated treatments. Black arrow in H&E images was for macrophages. Scale bar, 30 μm. g, h, Serum biochemical measurement for glucose (g) and glycated albumin (GA) (h). GA was presented as ratio of glycated albumin to total albumin. n = 9 for glucose and n = 8 for GA. Minimum to maximum, 0–100. The centre line indicates the median (50th percentile); the box indicates the 25th–75th percentiles of the dataset. One-way ANOVA. i-l, Serum biochemical measurement for triglyceride (TG) (i), total cholesterol (j), HDL-C (k), and LDL-C (l). n = 8. Minimum to maximum, 0–100. The centre line indicates the median (50th percentile); the box indicates the 25th–75th percentiles of the dataset. Statistical analysis was conducted by one-way ANOVA.
Extended Data Fig. 8 High GLUT1 or LDLR level is correlated to poor prognosis in multiple cancers.
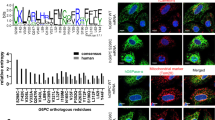
a, Analysis of GLUT1 expression across multiple tumours and normal tissues from TCGA RNA-seq data. Minimum to maximum, 0–100. The centre line indicates the median (50th percentile); the box indicates the 25th–75th percentiles of the dataset. Paired two-tailed Student’s t-test. b-k, Kaplan–Meier curves revealing the overall survival of patients with bladder cancer (BLCA, n = 403), breast cancer (BRCA, n = 1096), brain cancer (n = 667), cervical squamous cell carcinoma and endocervical adenocarcinoma (CESC, n = 304), head and neck squamous carcinoma (HNSC, n = 518), kidney renal clear cell carcinoma (KIRC, n = 532), liver hepatocellular carcinoma (LIHC, n = 369), lung adenocarcinoma (LUAD, n = 476), pancreatic adenocarcinoma (PAAD, n = 178) and prostate adenocarcinoma (PRAD, n = 597) based on GLUT1 mRNA levels in TCGA online database. Log-rank (Mantel-Cox) test. Error bar: 95% CI. l, Analysis of LDLR expression across multiple tumours and normal tissues from TCGA RNA-seq data. Minimum to maximum, 0–100. The centre line indicates the median (50th percentile); the box indicates the 25th–75th percentiles of the dataset. Paired two-tailed Student’s t-test. m-s, Kaplan–Meier curves revealing the overall survival of patients with bladder cancer (BLCA, n = 407), brain cancer (n = 653), kidney renal clear cell carcinoma (KIRC, n = 532), liver hepatocellular carcinoma (LIHC, n = 369), lung adenocarcinoma (LUAD, n = 496), pancreatic adenocarcinoma (PAAD, n = 178) and prostate adenocarcinoma (PRAD, n = 497) based on LDLR mRNA levels in TCGA online database. Log-rank (Mantel-Cox) test. Error bar: 95% CI.
Supplementary information
Supplementary Information
Supplementary Figs. 1–4 and Discussion.
Supplementary Table 1
Differential metabolites in AbCholB treated MDA-MB-231 cells compared to Ctrl. Including statistical source data for Figs. 3c–m and 4g and Extended Data Fig. 4.
Supplementary Video 1
Uptake of Dil-AbCholB by MDA-MB-231 cells in 12 h. MDA-MB-231 cells were seeded into a confocal dish. 50 μM Dil-AbCholB was added and the uptake process was videoed for 12 h. Scale bar, 20 μm.
Supplementary Video 2
LDL inhibits the uptake of Dil-AbCholB. LDL was added into MDA-MB-231 cells to block LDLR. 1 h later, 50 μM Dil-AbCholB was added for 12 h video. Scale bar, 20 μm.
Supplementary Video 3
Uptake of Dil-AbChol by MDA-MB-231 cells in 12 h. MDA-MB-231 cells were seeded into a confocal dish. 50 μM Dil-AbChol was added and the uptake process was videoed for 12 h.
Supplementary Video 4
LDL inhibits the uptake of Dil-AbChol. LDL was added into MDA-MB-231 cells to block LDLR. 1 h later, 50 μM Dil-AbChol was added for 12 h video. Scale bar, 20 μm.
Supplementary Video 5
Access of Dil-AbCholB into lysosomes in 12 h. MDA-MB-231 cells were seeded into a confocal dish. LAMP1-GFP probe was added to trace lysosomes prior to Dil-AbCholB addition. The medium was replenished with fresh medium contained 50 μM Dil-AbCholB and cells were videoed for 12 h. Scale bar, 20 μm.
Supplementary Video 6
Changes of co-localized positions of Dil-AbCholB and lysosomes from (Supplementary Video 5). The co-localized progress of Dil-AbCholB and LAMP1-positive lysosomes was analysed using cellSens Dimension software.
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Unprocessed western blots.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 5
Statistical source data.
Source Data Fig. 5
Unprocessed western blots.
Source Data Fig. 6
Statistical source data.
Source Data Extended Data Table 1
Statistical source data.
Source Data Extended Data Fig.1
Unprocessed western blots.
Source Data Extended Data Table 2
Statistical source data.
Source Data Extended Data Fig.2
Unprocessed western blots.
Source Data Extended Data Table 3
Statistical source data.
Source Data Extended Data Table 4
Statistical source data.
Source Data Extended Data Fig.3
Unprocessed western blots.
Source Data Extended Data Table 5
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Yue, C., Lu, W., Fan, S. et al. Nanoparticles for inducing Gaucher disease-like damage in cancer cells. Nat. Nanotechnol. (2024). https://doi.org/10.1038/s41565-024-01668-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41565-024-01668-4