Abstract
Oncogenic alterations in the RAS/RAF/MEK/ERK pathway drive the growth of a wide spectrum of cancers. While BRAF and MEK inhibitors are efficacious against BRAFV600E-driven cancers, effective targeted therapies are lacking for most cancers driven by other pathway alterations, including non-V600E oncogenic BRAF, RAS GTPase-activating protein (GAP) NF1 (neurofibromin 1) loss and oncogenic KRAS. Here, we show that targeting the SHP2 phosphatase (encoded by PTPN11) with RMC-4550, a small-molecule allosteric inhibitor, is effective in human cancer models bearing RAS–GTP-dependent oncogenic BRAF (for example, class 3 BRAF mutants), NF1 loss or nucleotide-cycling oncogenic RAS (for example, KRASG12C). SHP2 inhibitor treatment decreases oncogenic RAS/RAF/MEK/ERK signalling and cancer growth by disrupting SOS1-mediated RAS–GTP loading. Our findings illuminate a critical function for SHP2 in promoting oncogenic RAS/MAPK pathway activation in cancers with RAS–GTP-dependent oncogenic BRAF, NF1 loss and nucleotide-cycling oncogenic KRAS. SHP2 inhibition is a promising molecular therapeutic strategy for patients with cancers bearing these oncogenic drivers.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Dhillon, A. S., Hagan, S., Rath, O. & Kolch, W. MAP kinase signalling pathways in cancer. Oncogene 26, 3279–3290 (2007).
Bos, J. L., Rehmann, H. & Wittinghofer, A. GEFs and GAPs: critical elements in the control of small G proteins. Cell 130, 385 (2007).
The Cancer Genome Atlas Research Network Comprehensive molecular profiling of lung adenocarcinoma. Nature 511, 543–550 (2014).
Krauthammer, M. et al. Exome sequencing identifies recurrent mutations in NF1 and RASopathy genes in sun-exposed melanomas. Nat. Genet. 47, 996–1002 (2015).
Redig, A. J. et al. Clinical and molecular characteristics of NF1-mutant lung cancer. Clin. Cancer Res. 22, 3148–3156 (2016).
Caunt, C. J., Sale, M. J., Smith, P. D. & Cook, S. J. MEK1 and MEK2 inhibitors and cancer therapy: the long and winding road. Nat. Rev. Cancer 15, 577–592 (2015).
Bollag, G. et al. Vemurafenib: the first drug approved for BRAF-mutant cancer. Nat. Rev. Drug Discov. 11, 873–886 (2012).
Nieto, P. et al. A Braf kinase-inactive mutant induces lung adenocarcinoma. Nature 548, 239–243 (2017).
Yao, Z. et al. Tumours with class 3 BRAF mutants are sensitive to the inhibition of activated RAS. Nature 548, 234–238 (2017).
Lito, P., Solomon, M., Li, L.-S., Hansen, R. & Rosen, N. Allele-specific inhibitors inactivate mutant KRAS G12C by a trapping mechanism. Science 351, 604–608 (2016).
Ostrem, J. M., Peters, U., Sos, M. L., Wells, J. A. & Shokat, K. M. K-Ras(G12C) inhibitors allosterically control GTP affinity and effector interactions. Nature 503, 548–551 (2013).
Patricelli, M. P. et al. Selective inhibition of oncogenic KRAS output with small molecules targeting the inactive state. Cancer Discov. 6, 316–329 (2016).
Prahallad, A. et al. PTPN11 is a central node in intrinsic and acquired resistance to targeted cancer drugs. Cell Rep. 12, 1978–1985 (2015).
Niihori, T. et al. Functional analysis of PTPN11/SHP-2 mutants identified in Noonan syndrome and childhood leukemia. J. Hum. Genet. 50, 192–202 (2005).
Tartaglia, M. & Gelb, B. D. Germ-line and somatic PTPN11 mutations in human disease. Eur. J. Med. Genet. 48, 81–96 (2005).
Chen, Y.-N. P. et al. Allosteric inhibition of SHP2 phosphatase inhibits cancers driven by receptor tyrosine kinases. Nature 535, 148–152 (2016).
Wildes, D. et al. Allosteric inhibition of SHP2 variants containing cancer-associated activating mutations. Proc. Ann. Am. Ass. Cancer Res. 109, 4877 (2018).
Garcia-Fortanet, J. et al. Allosteric inhibition of SHP2: identification of a potent, selective, and orally efficacious phosphatase inhibitor. J. Med. Chem. 59, 7773–7782 (2016).
Poulikakos, P. I. et al. RAF inhibitor resistance is mediated by dimerization of aberrantly spliced BRAF(V600E). Nature 480, 387–390 (2011).
Yao, Z. et al. BRAF mutants evade ERK-dependent feedback by different mechanisms that determine their sensitivity to pharmacologic inhibition. Cancer Cell 28, 370–383 (2015).
Lito, P. et al. Relief of profound feedback inhibition of mitogenic signaling by RAF inhibitors attenuates their activity in BRAFV600E melanomas. Cancer Cell 22, 668–682 (2012).
Corbalan-Garcia, S., Yang, S. S., Degenhardt, K. R. & Bar-Sagi, D. Identification of the mitogen-activated protein kinase phosphorylation sites on human Sos1 that regulate interaction with Grb2. Mol. Cell. Biol. 16, 5674–5682 (1996).
Kamioka, Y., Yasuda, S., Fujita, Y., Aoki, K. & Matsuda, M. Multiple decisive phosphorylation sites for the negative feedback regulation of SOS1 via ERK. J. Biol. Chem. 285, 33540–33548 (2010).
Nissan, M. H. et al. Loss of NF1 in cutaneous melanoma is associated with RAS activation and MEK dependence. Cancer Res. 74, 2340–2350 (2014).
Shapira, S., Barkan, B., Friedman, E., Kloog, Y. & Stein, R. The tumor suppressor neurofibromin confers sensitivity to apoptosis by Ras-dependent and Ras-independent pathways. Cell Death Differ. 14, 895–906 (2007).
Hunter, J. C. et al. Biochemical and structural analysis of common cancer-associated KRAS mutations. Mol. Cancer Res. 13, 1325–1335 (2015).
Watson, I. R. et al. The RAC1 P29S hotspot mutation in melanoma confers resistance to pharmacological inhibition of RAF. Cancer Res. 74, 4845–4852 (2014).
Kobayashi, T. et al. Molecular and clinical analysis of RAF1 in Noonan syndrome and related disorders: dephosphorylation of serine 259 as the essential mechanism for mutant activation. Hum. Mutat. 31, 284–294 (2010).
Fujita-Sato, S. et al. Enhanced MET translation and signaling sustains K-Ras-driven proliferation under anchorage-independent growth conditions. Cancer Res. 75, 2851–2862 (2015).
Foster, R., Mudd, C., Wiggins, C. & Torrance, C. 3-Dimensional growth reveals KRAS dependency. Cancer Res. 73, 4289 (2013).
Vartanian, S. et al. Identification of mutant K-Ras-dependent phenotypes using a panel of isogenic cell lines. J. Biol. Chem. 288, 2403–2413 (2013).
Huang, H. et al. Oncogenic K-Ras requires activation for enhanced activity. Oncogene 33, 532–535 (2014).
Tashiro, T. et al. In vivo and ex vivo cetuximab sensitivity assay using three-dimensional primary culture system to stratify KRAS mutant colorectal cancer. PLoS One 12, e0174151 (2017).
McDonald, E. R.III et al. Project DRIVE: a compendium of cancer dependencies and synthetic lethal relationships uncovered by large-scale, deep RNAi screening. Cell 170, 577–592 (2017).
McDonald, E. R.III et al. Project DRIVE: a compendium of cancer dependencies and synthetic lethal relationships uncovered by large-scale, deep RNAi screening. Cell 170, 577–592.e10 (2017).
Drosten, M. et al. Genetic analysis of Ras signalling pathways in cell proliferation, migration and survival. EMBO J. 29, 1091–1104 (2010).
Dance, M., Montagner, A., Salles, J. P., Yart, A. & Raynal, P. The molecular functions of Shp2 in the Ras/Mitogen-activated protein kinase (ERK1/2) pathway. Cell Signal. 20, 453–459 (2008).
Hadari, Y. R., Kouhara, H., Lax, I. & Schlessinger, J. Binding of Shp2 tyrosine phosphatase to FRS2 is essential for fibroblast growth factor-induced PC12 cell differentiation. Mol. Cell. Biol. 18, 3966–3973 (1998).
Aronheim, A. et al. Membrane targeting of the nucleotide exchange factor Sos is sufficient for activating the Ras signaling pathway. Cell 78, 949–961 (1994).
Cunnick, J. M., Dorsey, J. F., Munoz-Antonia, T., Mei, L. & Wu, J. Requirement of SHP2 binding to Grb2-associated binder-1 for mitogen-activated protein kinase activation in response to lysophosphatidic acid and epidermal growth factor. J. Biol. Chem. 275, 13842–13848 (2000).
Cunnick, J. M. et al. Regulation of the mitogen-activated protein kinase signaling pathway by SHP2. J. Biol. Chem. 277, 9498–9504 (2002).
Batth, T. S. et al. Large-scale phosphoproteomics reveals Shp-2 phosphatase-dependent regulators of Pdgf receptor signaling. Cell Rep. 22, 2784–2796 (2018).
Niihori, T. et al. Germline KRAS and BRAF mutations in cardio-facio-cutaneous syndrome. Nat. Genet. 38, 294–296 (2006).
Janes, M. R. et al. Targeting KRAS mutant cancers with a covalent G12C-specific inhibitor. Cell 172, 578–589e.17 (2018).
Zehir, A. et al. Mutational landscape of metastatic cancer revealed from prospective clinical sequencing of 10,000 patients. Nat. Med. 23, 703–713 (2017).
Koltun, E. & Revolution Medicines, Inc. (WO2018013597) 2,5-Disubstituted 3-methyl pyrazines and 2,5,6-trisubstituted 3-methyl pyrazines as allosteric SHP2 inhibitors. Patent no. PCT/US2017/041577 (2017); https://go.nature.com/2mv9pBw
Lee, S. J. et al. Regulation of hypoxia-inducible factor 1alpha (HIF-1alpha) by lysophosphatidic acid is dependent on interplay between p53 and Kruppel-like factor 5. J. Biol. Chem. 288, 25244–25253 (2013).
National Cancer Institute. The NCI RAS initiative. National Institutes of Health https://www.cancer.gov/research/key-initiatives/ras (2018).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Acknowledgements
Revolution Medicines thanks S. Kelsey for scientific guidance and critical review of the manuscript, K. Shokat for scientific guidance, N. Tibrewal for development of the phosphatase selectivity panel and J. Jasper for scientific input. F. Haderk was supported by the Mildred Scheel postdoctoral fellowship from the German Cancer Aid. Funding support from NIH/NCI U54CA224081, NIH/NCI R01CA204302, NIH/NCI R01CA211052, NIH/NCI R01CA169338 and Pew Trust and Stewart Foundation (T.G.B.) is acknowledged. We are grateful to researchers at MI Bioresearch, WuXi Apptec and Genendesign who support our in vivo pharmacology efforts.
Author information
Authors and Affiliations
Contributions
R.J.N. contributed to the design, conduct and interpretation of all experiments. F.H. contributed in vitro and in vivo experimental design, conduct and interpretation. C.S., C.J.S, G.H., D.W., C.T., K.M., A.M., J.R., T.H. and A.Z. contributed in vitro experimental design, conduct and interpretation. V.O., C.M. and Z.W. contributed design, conduct and interpretation of in vivo experiments. C.M.B. established and maintains the TH77 model. G.K., E.S.K. and A.L.G. directed the design and synthesis of RMC-4550. M.S. contributed design, conduct and interpretation of in vivo experiments. M.A.G. contributed to the interpretation of all experiments. J.A.M.S. and T.G.B. supervised the project and contributed to the design and interpretation of all experiments. R.J.N., J.A.M.S. and T.G.B. wrote the manuscript with input from all co-authors.
Corresponding authors
Ethics declarations
Competing interests
T.G.B. is a consultant to Novartis, AstraZeneca, Array Biopharma (Scientific Advisory Board) and Inventiva, and is the recipient of research grants from Ignyta and Revolution Medicines. T.G.B. is a member of the Revolution Medicines Scientific Advisory Board. R.J.N., C.S., C.J.S., D.W., C.T., K.M., A.M., J.R., T.H., Z.W., G.K., E.S.K., A.L.G., M.S., M.A.G. and J.A.M.S. are employees of Revolution Medicines, Inc. Revolution Medicines holds the patent mentioned in the Methods section. The other authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Western blot validation of a RAS-GTP ELISA.
NCI-H358 cells were grown in 2D culture and incubated with increasing concentrations of RMC-4450 for one hour. a) Cellular lysates were prepared and levels of RAS-GTP were measured by ELISA. Shown are three independent biological replicates performed in technical duplicate. b) Assessment of RAS-GTP in samples from replicate 2 by western blot. RAS-GTP was affinity purified from the cellular lysates using the GST-RAF-RBD bound to glutathione resin. Source data is provided in Supplementary Table 9.
Supplementary Figure 2 Expression of SHP2E76K rescues Class 3 BRAF, NF1LOF and KRASG12C mutant cell lines from RMC-4550.
Stable expression of SHP2-WT and SHP2-E76K was induced upon lentiviral transfection of NCI-H508 (Class 3 BRAF), NCI-H1838 (NF1LOF), and NCI-H358 (KRASG12C) cells (n = 3 biologically independent experiments; figures show mean +/− S.D.) a, c, e) Effect of RMC-4550 on proliferation of NCI-H508 (a), NCI-H1838 cells (c), and NCI-H358 cells (e) in 3D culture. Three days after seeding, cells were treated with RMC-4550 and cell viability measured on Day 8 using CTG (data are representative of n = 2 independent experiments for NCI-H508 and NCI-H1838, and n = 3 observations for H358, in technical duplicate; figures show mean +/− S.D.). RMC-4550 exhibited concentration-dependent inhibition of growth in parental and SHP2WT-expressing cell lines representing all three genotypes but failed to elicit a reduction in proliferation ≥50% of control signal in SHP2E76K expressing cells up to the maximal test concentration of 10 μM. b, d, f) Effect of RMC-4550 on pERK levels in NCI-H508 (b), NCI-H1838 cells (d), and NCI-H358 cells (f). Cells were grown in 2D culture and incubated with 1 µM RMC-4550 for 90 minutes. Cellular lysates were prepared and levels of pERK determined by immunoblot. RMC-4550 treatment resulted in reduction of pERK levels in parental and SHP2WT expressing cells across all genotypes, but failed to suppress pERK in SHP2E76K expressing cells. Shown is one representative example of three independent experiments. Source data is provided in Supplementary Table 9.
Supplementary Figure 3 Effects of SHP2 inhibition in RAS pathway-driven cancer cells.
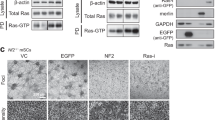
(a) Effect of RMC-4550 on RAS-GTP levels in BEAS-2B cells stably transfected with scrambled (shScramble) or NF1 targeting (shNF1) shRNA. Data shows a representative of three independent experiments. (b-d) (b) Cell sensitivities to the knockdowns of the indicated genes were determined using crystal violet staining assays (n = 2). The spectrophotometric intensities of the crystal violet stains were quantified and represented as a bar plot. (c) RAS-GTP levels in cellular lysates are shown by GST-RAF-RBD affinity capture. (d) Individual western blot band intensities were quantified and shown for pERK/ERK, RAS-GTP/RAS (total) level and knock down efficiency, expressed relative to a control scrambled shRNA intensity of 1.0 and the effect of knockdown on pERK. (e-h) (e), (f) Effect of RMC-4550 and the KRASG12C-specific inhibitor ARS-853 on pERK levels in KRASG12C cancer cell lines. Cells were incubated with increasing concentrations of RMC-4550 or ARS-853 for one hour. Cellular lysates were prepared and levels of pERK determined. RMC-4550 produced a concentration-dependent reduction in pERK levels. Geometric mean IC50 values for reduction in pERK by RMC-4550 in NCI-H358 and CALU-1 were 36 nM and 7 nM, respectively. A concentration-dependent reduction in pERK levels was observed for ARS-853 (geometric mean IC50 values of 4700 nM and 430 nM for NCI-H358 and CALU-1 cells, respectively); data represent n = 4 independent experiments, performed in technical duplicate; figures show mean +/− S.D.). (g), (h) Effect of RMC-4550 and ARS-853 on proliferation of two KRASG12C cancer cell lines in 3D culture. NCI-H358 and CALU-1 cells were treated with RMC-4550 or ARS-853 and cell viability measured on Day 7 using Cell Titer Glo (data represent n = 4 independent experiments, performed in duplicate for NCI-H358, and n = 3 independent experiments, performed in duplicate for CALU-1; figures show mean +/− S.D.). RMC-4550 and ARS-853 exhibited geometric mean IC50 values of 43 nM and 1020 nM, respectively, in NCI-H358 cells. Source data is provided in Supplementary Table 9.
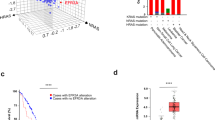
Supplementary Figure 4 Effects of SHP2 inhibition in KRAS-driven cancer cells.
(a) Comparison of the effect of SHP2 inhibition and KRAS knockdown on the proliferation of KRAS-mutant cancer cell lines bearing different oncogenic mutations in KRAS as indicated by color coding. The negative logarithm of RMC-4550 GI50 values for cancer cells bearing activating KRAS mutations (described in Supplementary Table 5) were plotted against the negative value of their Project Drive Sensitivity Score for KRAS shRNA knockdown. Cells with high sensitivity to KRAS knockdown have larger negative Sensitivity Scores, and cells with greater sensitivity to SHP2 inhibition have larger negative log(RMC-4550 GI50) values. (b-c) Effect of RMC-4550 on RAS-GTP and pERK levels in Hras−/−, Nras−/−, Kras−/− mouse embryonic fibroblasts (MEFs) expressing solely the indicated RAS or BRAF transcript. Cells were grown in 2D culture and incubated with 1 µM RMC-4550 for 24 h. b) Cellular lysates were prepared and levels of RAS-GTP and pERK determined by immunoblot. RAS-GTP was affinity purified from the cellular lysates using GST-tagged RAF-RBD coupled to glutathione agarose beads. (c) RAS-GTP levels were quantified in whole cell lysates via RAS-GTP ELISA. Significant reduction in RAS-GTP and pERK levels is observed for KRAS WT (WT-4B) and KRAS G12 expressing MEFs, while cells expressing KRASQ61H lack response. Values above bars represent fraction of RAS-GTP in RMC-4550 vs. DMSO treated cells. Data in (b) shows a representative example of three independent experiments; data in (c) shows a representative example of two independent experiments, and bars indicates mean of two technical replicates. (d-e) (d) Expression of the wild type SOS1 and SOS-F proteins was induced by doxycycline treatment for 24 h. Cells were treated with 50 ng/mL EGF at various time intervals, lysed, and total pERK levels were measured by AlphaLisa. The SOS-F construct showed increased and sustained levels of pERK signal in the absence of EGF stimulation. Shown is one representative example performed in technical duplicate of three independent experiments. Figure indicates mean +/− S.D. (e) NCI-H358 cells transiently expressing SOS1 and SOS-F encoding cDNA for 48 h were treated with vehicle or 1 µM RMC-4550 for 1 and 6 h. Harvested lysates from the cells were probed for the indicated proteins using western blot. Data shown represent at least 3 independent biological replicates. Source data is provided in Supplementary Table 9.
Supplementary Figure 5 Xenograft models of KRASG13D and KRASQ61K are insensitive to RMC-4550.
Daily oral administration of RMC-4550 (30 mg/kg) to athymic nude mice bearing (a) HCT-116 (KRASG13D) or (b) CALU-6 (KRASQ61K) xenograft tumors did not produce significant tumor growth inhibition. Treatment with RMC-4550 was well tolerated in both models, as assessed by body weight (c,d). Source data is provided in Supplementary Table 9.
Supplementary Figure 6 Tolerability assessment in animals treated with the SHP2 inhibitor.
Repeated daily oral administration of RMC-4550 of all dose groups in the MeWo (a), NCI-H1838 (b), NCI-H358 (c), and MIA PaCa-2 (d), xenograft models and LUN023 (e), LUN037 (f), TH77 (g), LUN092 (h), and LUN156 (i) PDX models was well tolerated as assessed by body weight. Source data is provided in Supplementary Table 9.
Supplementary Figure 7 Durable response to RMC-4550 in NF1LOF model.
(a) Western blot demonstrates loss of NF1 protein in TH77 PDX model. A previously published EGFRL858R, NF1wt PDX model is shown as comparator1. (b) Tumor specimen of vehicle and RMC-4550 treated groups were analyzed regarding their general tissue morphology by H&E stain (representative of n = 3 independent samples per treatment arm, 10X magnification, size bar: 100 nm). Significant areas of necrosis (arrows) were present in tumors of RMC-4550 treated animals.
Supplementary Figure 8 Unprocessed scans of raw blots.
(a) SHP2 IP from Fig. 5c, (b) GAB1 IP from Fig. 5d, and (c) membrane fractionation from Fig. 5e. (d) Effect of RMC-4550 on RAS-GTP levels from Supplementary Figure 1b, (e) Effect of SHP2 variants on pERK in H508 cells from Supplementary Figure 2b, (f) Effect of SHP2 variants on pERK in H1838 cells from Supplementary Figure 2d, (g) Effect of SHP2 variants on pERK in H358 cells from Supplementary Figure 2f. (h) Effect of NF1 shRNA knockdown on RAS-GTP levels from Supplementary Figure 3a, and (i) RAS-GTP levels from Supplementary Figure 3c, (j) Effect of SHP2 and SOS1 shRNA knockdown in RAS-GTP levels from Supplementary Figure 3d, (k) Effect of RMC-4550 on RAS-GTP and pERK levels in RAS-less MEFs from Supplementary Figure 4b, (l) Effect of SOS or SOS-F expression on pERK from Supplementary Figure 4e, (m) NF1 levels in TH77 PDX model from Supplementary Figure 7a. Note that panels (d) through (e) are located in additional pages.
Supplementary information
Supplementary Information
Supplementary Figures 1–9 and Supplementary Table legends.
Supplementary Table 1
RMC-4550 activity against a panel of 15 phosphatases.
Supplementary Table 2
RMC-4550 activity against a panel of 468 kinases at a test concentration of 10 µM.
Supplementary Table 3
RMC-4550 activity against a panel of 44 safety targets at a test concentration of 10 μM.
Supplementary Table 4
RMC-4550 activity against a panel of NF1 mutant cell lines.
Supplementary Table 5
RMC-4550 activity against a panel of 33 KRAS mutant cell lines.
Supplementary Table 6
Pharmacokinetic parameters of RMC-4550 from single dose NCI-H358 study.
Supplementary Table 7
Pharmacokinetic parameters of RMC-4550 from end of MIA PaCa-2 efficacy study.
Supplementary Table 8
Antibodies.
Supplementary Table 9
Statistics source data.
Rights and permissions
About this article
Cite this article
Nichols, R.J., Haderk, F., Stahlhut, C. et al. RAS nucleotide cycling underlies the SHP2 phosphatase dependence of mutant BRAF-, NF1- and RAS-driven cancers. Nat Cell Biol 20, 1064–1073 (2018). https://doi.org/10.1038/s41556-018-0169-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-018-0169-1
This article is cited by
-
Preclinical translational platform of neuroinflammatory disease biology relevant to neurodegenerative disease
Journal of Neuroinflammation (2024)
-
Combinatorial strategies to target RAS-driven cancers
Nature Reviews Cancer (2024)
-
Targeting KRAS in cancer
Nature Medicine (2024)
-
New clinical trial design in precision medicine: discovery, development and direction
Signal Transduction and Targeted Therapy (2024)
-
BRAF — a tumour-agnostic drug target with lineage-specific dependencies
Nature Reviews Clinical Oncology (2024)