Abstract
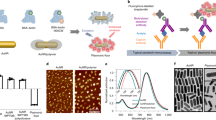
The ability to detect biomarkers with ultrahigh sensitivity radically transformed biology and disease diagnosis. However, owing to incompatibilities with infrastructure in current biological and medical laboratories, recent innovations in analytical technology have not yet been adopted broadly. Here, we report a simple, universal ‘add-on’ technology (dubbed EASE) that converts the ordinary sensitivities of common bioassays to extraordinary ones, and that can be directly plugged into the routine practices of current research and clinical laboratories. The assay relies on the bioconjugation capabilities and ultrafast and localized deposition of polydopamine at the target site, which permit a large number of reporter molecules to be captured and lead to detection-sensitivity enhancements exceeding three orders of magnitude. The application of EASE in the ELISA-based detection of the HIV antigen in blood from patients leads to a sensitivity lower than 3 fg ml−1. We also show that EASE allows for the direct visualization, in tissues, of the Zika virus and of low-abundance biomarkers related to neurological diseases and cancer immunotherapy.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$99.00 per year
only $8.25 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Howes, P. D., Chandrawati, R. & Stevens, M. M. Colloidal nanoparticles as advanced biological sensors. Science 346, 1247390 (2014).
Chan, W. C. & Nie, S. Quantum dot bioconjugates for ultrasensitive nonisotopic detection. Science 281, 2016–2018 (1998).
Kelley, S. O. et al. Advancing the speed, sensitivity and accuracy of biomolecular detection using multi-length-scale engineering. Nat. Nanotech. 9, 969–980 (2014).
Nam, J.-M., Thaxton, C. S. & Mirkin, C. A. Nanoparticle-based bio-bar codes for the ultrasensitive detection of proteins. Science 301, 1884–1886 (2003).
Kosaka, P. M. et al. Detection of cancer biomarkers in serum using a hybrid mechanical and optoplasmonic nanosensor. Nat. Nanotech. 9, 1047–1053 (2014).
Rodriguez-Lorenzo, L., de La Rica, R., Alvarez-Puebla, R. A., Liz-Marzan, L. M. & Stevens, M. M. Plasmonic nanosensors with inverse sensitivity by means of enzyme-guided crystal growth. Nat. Mater. 11, 604–607 (2012).
He, L., Ozdemir, S. K., Zhu, J., Kim, W. & Yang, L. Detecting single viruses and nanoparticles using whispering gallery microlasers. Nat. Nanotech. 6, 428–432 (2011).
Thomas, R. K. et al. Sensitive mutation detection in heterogeneous cancer specimens by massively parallel picoliter reactor sequencing. Nat. Med. 12, 852–855 (2006).
Zheng, G., Patolsky, F., Cui, Y., Wang, W. U. & Lieber, C. M. Multiplexed electrical detection of cancer markers with nanowire sensor arrays. Nat. Biotechnol. 23, 1294–1301 (2005).
Wu, G. et al. Bioassay of prostate-specific antigen (PSA) using microcantilevers. Nat. Biotechnol. 19, 856–860 (2001).
Schallmeiner, E. et al. Sensitive protein detection via triple-binder proximity ligation assays. Nat. Methods 4, 135–137 (2007).
Watanabe, R. et al. Arrayed lipid bilayer chambers allow single-molecule analysis of membrane transporter activity. Nat. Commun. 5, 4519 (2014).
Ma, W. et al. Attomolar DNA detection with chiral nanorod assemblies. Nat. Commun. 4, 2689 (2013).
Haun, J. B., Devaraj, N. K., Hilderbrand, S. A., Lee, H. & Weissleder, R. Bioorthogonal chemistry amplifies nanoparticle binding and enhances the sensitivity of cell detection. Nat. Nanotech. 5, 660–665 (2010).
Vollmer, F. & Arnold, S. Whispering-gallery-mode biosensing: label-free detection down to single molecules. Nat. Methods 5, 591–596 (2008).
Li, M., Tang, H. X. & Roukes, M. L. Ultra-sensitive NEMS-based cantilevers for sensing, scanned probe and very high-frequency applications. Nat. Nanotech. 2, 114–120 (2007).
Cooper, M. A. et al. Direct and sensitive detection of a human virus by rupture event scanning. Nat. Biotechnol. 19, 833–837 (2001).
Rissin, D. M. et al. Single-molecule enzyme-linked immunosorbent assay detects serum proteins at subfemtomolar concentrations. Nat. Biotechnol. 28, 595–599 (2010).
Burst bubbles. Nature 526, 609–610 (2015).
Lee, H., Dellatore, S. M., Miller, W. M. & Messersmith, P. B. Mussel-inspired surface chemistry for multifunctional coatings. Science 318, 426–430 (2007).
Lee, H., Rho, J. & Messersmith, P. B. Facile conjugation of biomolecules onto surfaces via mussel adhesive protein inspired coatings. Adv. Mater. 21, 431–434 (2009).
Soderhall, K. & Cerenius, L. Role of the prophenoloxidase-activating system in invertebrate immunity. Curr. Opin. Immunol. 10, 23–28 (1998).
Cerenius, L. & Soderhall, K. The prophenoloxidase-activating system in invertebrates. Immunol. Rev. 198, 116–126 (2004).
Weber, R. et al. Threshold of detection of Cryptosporidium oocysts in human stool specimens: evidence for low sensitivity of current diagnostic methods. J. Clin. Microbiol. 29, 1323–1327 (1991).
Mahler, M., Ngo, J. T., Schulte-Pelkum, J., Luettich, T. & Fritzler, M. J. Limited reliability of the indirect immunofluorescence technique for the detection of anti-Rib-P antibodies. Arth. Res. Ther. 10, R131 (2008).
Zrazhevskiy, P., Sena, M. & Gao, X. Designing multifunctional quantum dots for bioimaging, detection, and drug delivery. Chem. Soc. Rev. 39, 4326–4354 (2010).
Medintz, I. L., Uyeda, H. T., Goldman, E. R. & Mattoussi, H. Quantum dot bioconjugates for imaging, labelling and sensing. Nat. Mater. 4, 435–446 (2005).
Zrazhevskiy, P. et al. Cross-platform DNA encoding for single-cell imaging of gene expression. Angew. Chem. Int. Ed. 55, 8975–8978 (2016).
Battersby, B. J., Lawrie, G. A. & Trau, M. Optical encoding of microbeads for gene screening: alternatives to microarrays. Drug Discov. Today 6, 19–26 (2001).
Braeckmans, K., De Smedt, S. C., Leblans, M., Pauwels, R. & Demeester, J. Encoding microcarriers: present and future technologies. Nat. Rev. Drug Discov. 1, 447–456 (2002).
Nolan, J. P. & Sklar, L. A. Suspension array technology: evolution of the flat-array paradigm. Trends Biotechnol. 20, 9–12 (2002).
Fulton, R. J., McDade, R. L., Smith, P. L., Kienker, L. J. & Kettman, J. R. Advanced multiplexed analysis with the FlowMetrixTM system. Clin. Chem. 43, 1749–1756 (1997).
Dunbar, S. A. Applications of Luminex xMAP technology for rapid, high-throughput multiplexed nucleic acid detection. Clin. Chim. Acta 363, 71–82 (2006).
Chan, W. C. et al. Luminescent quantum dots for multiplexed biological detection and imaging. Curr. Opin. Biotechnol. 13, 40–46 (2002).
Han, M., Gao, X., Su, J. Z. & Nie, S. Quantum-dot-tagged microbeads for multiplexed optical coding of biomolecules. Nat. Biotechnol. 19, 631–635 (2001).
Klein, D., Hurley, L. B., Merrill, D. & Quesenberry, C. P. Jr Review of medical encounters in the 5 years before a diagnosis of HIV-1 infection: implications for early detection. J. Acq. Immune Defic. Syndr. 32, 143–152 (2003).
Fu, E. et al. Enhanced sensitivity of lateral flow tests using a two-dimensional paper network format. Analyt. Chem. 83, 7941–7946 (2011).
Palella, F. J. et al. Survival benefit of initiating antiretroviral therapy in HIV-infected persons in different CD4+ cell strata. Ann. Intern. Med. 138, 620–626 (2003).
Holodniy, M. et al. Relationship between antiretroviral prescribing patterns and treatment guidelines in treatment-naive HIV-1-infected US veterans (1992–2004). J. Acq. Immune Defic. Syndr 44, 20–29 (2007).
Marks, G., Crepaz, N. & Janssen, R. S. Estimating sexual transmission of HIV from persons aware and unaware that they are infected with the virus in the USA. AIDs 20, 1447–1450 (2006).
Miles, S. A. et al. Rapid serologic testing with immune-complex-dissociated HIV p24 antigen for early detection of HIV infection in neonates. New Engl. J. Med. 328, 297–302 (1993).
Nishanian, P., Huskins, K. R., Stehn, S., Detels, R. & Fahey, J. L. A simple method for improved assay demonstrates that HIV p24 antigen is present as immune complexes in most sera from HIV-infected individuals. J. Infect. Dis. 162, 21–28 (1990).
Marozsan, A. J. et al. Relationships between infectious titer, capsid protein levels, and reverse transcriptase activities of diverse human immunodeficiency virus type 1 isolates. J. Virol. 78, 11130–11141 (2004).
Bale, T. L. & Vale, W. W. CRF and CRF receptors: role in stress responsivity and other behaviors. Annu. Rev. Pharmacol. Toxicol. 44, 525–557 (2004).
Zorrilla, E. P., Logrip, M. L. & Koob, G. Corticotropin releasing factor: a key role in the neurobiology of addiction. Front. Neuroendocrinol. 35, 234–244 (2014).
Van Pett, K. et al. Distribution of mRNAs encoding CRF receptors in brain and pituitary of rat and mouse. J. Comp. Neurol. 428, 191–212 (2000).
Weathington, J. M. & Cooke, B. M. Corticotropin-releasing factor receptor binding in the amygdala changes across puberty in a sex-specific manner. Endocrinol. 153, 5701–5705 (2012).
Waldorf, K. M. A. et al. Fetal brain lesions after subcutaneous inoculation of Zika virus in a pregnant nonhuman primate. Nat. Med. 22, 1256–1259 (2016).
Keir, M. E. et al. PD-1 and its ligands in tolerance and immunity. Annu. Rev. Immunol. 26, 677–704 (2008).
Brahmer, J. R. et al. Phase I study of single-agent anti–programmed death-1 (MDX-1106) in refractory solid tumors: safety, clinical activity, pharmacodynamics, and immunologic correlates. J. Clin. Oncol. 28, 3167–3175 (2010).
Hamid, O. et al. Safety and tumor responses with lambrolizumab (anti–PD-1) in melanoma. New Engl. J. Med. 369, 134–144 (2013).
Garon, E. B. et al. Pembrolizumab for the treatment of non-small-cell lung cancer. New Engl. J. Med. 372, 2018–2028 (2015).
Rizvi, N. A. et al. Activity and safety of nivolumab, an anti-PD-1 immune checkpoint inhibitor, for patients with advanced, refractory squamous non-small-cell lung cancer (CheckMate 063): a phase 2, single-arm trial. Lancet Oncol. 16, 257–265 (2015).
Nghiem, P. T. et al. PD-1 blockade with pembrolizumab in advanced Merkel-cell carcinoma. New Engl. J. Med. 374, 2542–2552 (2016).
Wang, X. et al. PD-L1 expression in human cancers and its association with clinical outcomes. OncoTarg. Ther. 9, 5023–5039 (2016).
Liu, Y., Ai, K. & Lu, L. Polydopamine and its derivative materials: synthesis and promising applications in energy, environmental, and biomedical fields. Chem. Rev. 114, 5057–5115 (2014).
Li, J. et al. Stably doped conducting polymer nanoshells by surface initiated polymerization. Nano Lett. 15, 8217–8222 (2015).
Sanford, C. A. et al. A central amygdala CRF circuit facilitates learning about weak threats. Neuron 93, 164–178 (2017).
Zhao, H. et al. Structural basis of Zika virus-specific antibody protection. Cell 166, 1016–1027 (2016).
Acknowledgements
This work was supported in part by the NIH (R21CA192985, R01AI100989, AI083019, AI104002, and AI060389) and the Department of Bioengineering at the University of Washington. J.L. thanks the Howard Hughes Medical Institute for a student fellowship. We are also grateful to B. Lutz and D. Leon for help with the lateral flow test, and P. Zrazhevskiy for discussions on immunostaining.
Author information
Authors and Affiliations
Contributions
J.L. and X.G. conceived the idea and designed the project. J.L. and W.T. performed the majority of the experiments, with help from M.A.B. and L.S.Z. for CRF imaging in the brain, from M.A.D., K.M.A.W., M.G. and L.R. for ZIKA imaging, and R.H.P. for PD-L1 imaging. All authors were involved in data analysis. J.L., L.S.Z., K.M.A.W., M.G., L.R., R.H.P. and X.G. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary methods, figures and tables. (PDF 7035 kb)
Rights and permissions
About this article
Cite this article
Li, J., Baird, M., Davis, M. et al. Dramatic enhancement of the detection limits of bioassays via ultrafast deposition of polydopamine. Nat Biomed Eng 1, 0082 (2017). https://doi.org/10.1038/s41551-017-0082
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41551-017-0082
This article is cited by
-
Renal clearable polyfluorophore nanosensors for early diagnosis of cancer and allograft rejection
Nature Materials (2022)
-
Biosensor-based assay of exosome biomarker for early diagnosis of cancer
Frontiers of Medicine (2022)
-
Recent advances in sensitivity enhancement for lateral flow assay
Microchimica Acta (2021)
-
An electrochemiluminescence immunosensor for the N-terminal brain natriuretic peptide based on the high quenching ability of polydopamine
Microchimica Acta (2019)
-
Melanin/polydopamine-based nanomaterials for biomedical applications
Science China Chemistry (2019)