Abstract
Common and rare variants in the LRRK2 locus are associated with Parkinson’s disease (PD) risk, but the downstream effects of these variants on protein levels remain unknown. We performed comprehensive proteogenomic analyses using the largest aptamer-based CSF proteomics study to date (7006 aptamers (6138 unique proteins) in 3107 individuals). The dataset comprised six different and independent cohorts (five using the SomaScan7K (ADNI, DIAN, MAP, Barcelona-1 (Pau), and Fundació ACE (Ruiz)) and the PPMI cohort using the SomaScan5K panel). We identified eleven independent SNPs in the LRRK2 locus associated with the levels of 25 proteins as well as PD risk. Of these, only eleven proteins have been previously associated with PD risk (e.g., GRN or GPNMB). Proteome-wide association study (PWAS) analyses suggested that the levels of ten of those proteins were genetically correlated with PD risk, and seven were validated in the PPMI cohort. Mendelian randomization analyses identified GPNMB, LCT, and CD68 causal for PD and nominate one more (ITGB2). These 25 proteins were enriched for microglia-specific proteins and trafficking pathways (both lysosome and intracellular). This study not only demonstrates that protein phenome-wide association studies (PheWAS) and trans-protein quantitative trail loci (pQTL) analyses are powerful for identifying novel protein interactions in an unbiased manner, but also that LRRK2 is linked with the regulation of PD-associated proteins that are enriched in microglial cells and specific lysosomal pathways.
Similar content being viewed by others
Introduction
Leucine-rich repeat kinase 2 (LRRK2) gene mutations can cause Mendelian Parkinson’s disease (PD). At the same time, common variant in this locus are some of the most significant PD risk variants. One of the largest PD risk genome-wide association studies (GWAS) to date1 found multiple independent signals in the LRRK2 gene. Subsequent studies2 examined the association between the common variant (SNP) rs76904798 (chr12:40220632:C:T) with LRRK2 RNA expression and found that the minor allele (T) was associated with increased expression levels of LRRK2. Like most of neurodegenerative diseases, PD is a complex disorder with several, and sometimes independently interconnected, pathways encompassing multiple proteins contributing to disease pathogenesis. Identifying those pathways, as well as the included proteins and protein interactions, is instrumental to decipher the pathobiology of the disease.
A recent study3 leveraged the cerebrospinal fluid (CSF) proteome to identify the causal proteins under the PD loci by performing cis-protein quantitative trait loci (pQTL) and Mendelian randomization (MR). A limitation of using cis-signals is the non-assessment of potential protein–protein interaction. We recently demonstrated that common variants in MS4A4A are a trans-pQTL for CSF soluble TREM2 (sTREM2) levels and are associated with Alzheimer’s disease (AD) risk4, indicating that MS4A4A modifies risk for AD by modifying TREM2 biology, and that MS4A4A is a potential therapeutic target. This led to several active clinical trials targeting MS4A4A to treat AD. Along the same lines, we performed unbiased multi-tissue pQTL mapping and identified hundreds of cis and trans pQTLs. These pQTLs were later used to identify causal and druggable targets for AD, PD, frontotemporal dementia (FTD), amyotrophic lateral sclerosis (ALS), and stroke5. Together, all these findings strongly support the use of pQTL analyses for the identification of new causal proteins, causal pathways, and potential therapeutic targets.
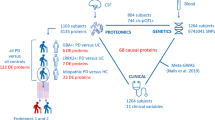
Here, we performed a trans-pQTL study to identify proteins associated with LRRK2 variants using the largest aptamer-based CSF proteomics study to date (7006 aptamers (6138 unique proteins) in 3107 individuals), and additional mediation and pathway analyses to understand the downstream effects of LRRK2 variants (Fig. 1). We discovered that a majority of the identified proteins were enriched in the endolysosomal pathway and immune function.
Results
Proteins associated with LRRK2 Parkinson’s disease risk variant
To identify proteins associated with LRRK2 variants, we examined the largest CSF pQTL dataset to identify proteins associated with the sentinel variant in this locus: rs76904798. PheWAS analyses identified 14 proteins in the pQTL atlas with protein levels associated to this variant (C1QTNF1, CD63, CD68, ENTPD1, GPNMB, GREM2, GRN, HLA-DQA2, ITGB2, NIPAL4, OLR1, SDCBP2, TLR3, and TMEM106A). Then, we performed conditional analyses to identify independent signals in this region for these proteins. We identified a total of 15 SNPs associated with these proteins, and 12 SNPs were not in LD (r2 < 0.85). Next, we performed a PheWAS of the 12 independent SNPs and identified 11 additional proteins (AGFG2, CA1, CHIT1, DNAJC15, EID3, FCGR1A, FTL, GAA, LCT, LGALS9, and SRI) (Fig. 2a and Supplementary Fig. 1). Overall, 25 unique proteins were found to be associated with 12 independent SNPs in the LRRK2 locus (Fig. 2b; plot made using ref. 6 Circlize).
a LRRK2 PheWAS plot of 24 CSF proteins over four SNPs. The PD risk SNP is highlighted red. Genes previously associated with PD is shown as a color in a circle. x axis is proteins sorted by chr12:40220632:C:T P values and y axis is FDR corrected −log10(P value). For proteins with multiple aptamers, the most significant P value is represented. b Circos plot of proteins in chr12:40220632:C:T, chr12:40204350:T:G, and chr12:40035894:C:T. Width of connection represents the SNP effect size. All 14 proteins in chr12:40220632:C:T had positive effect size and the 16 proteins in chr12:40035894:C:T had a negative effect size. c LRRK2 PheWAS plot of 14 CSF proteins (18 aptamers). All FDR passing aptamers had positive chr12:40220632:C:T direction (light red). FDR corrected −log10(P value) y axis and x axis is entrez gene ID.
Multiple proteins (such as GPNMB, CD68, OLR1, and FTL) were measured with more than one aptamer and showed high correlation (R > 0.9). We also analyzed the correlation accross different proteins and found that GRN and C1QTNF1 (R = 0.88) showed high correlation, while the remaining 20 proteins showed acorrelation of R < 0.7 (Supplementary Fig. 2).
Eleven (GRN, GPNMB, HLA-DQA2, CD63, CD68, EID3, ENTPD1, GAA, LCT, SRI, and TLR3) of the 25 proteins have been reported to be implicated in PD pathogenesis in previous studies (Supplementary Table 3). Six proteins (AGFG2, C1QTNF1, CHIT1, DNAJC15, FTL, and OLR1) have been reported to be implicated in other neurodegenerative diseases such as AD, ALS, or stroke, but not in PD. The rest (eight) are proteins not implicated with PD or neurodegeneration, such as GREM2, ITGB2, SDCBP2, and TMEM106A.
In order to determine if these results are robust and replicate across cohorts, we performed analyses for each of the top aptamer-SNP pairs in each of the six independent cohorts (Supplementary Fig. 3). For example, the pair GRN and chr12:40220632:C:T (MAF = 0.134) had the effect sizes of all the cohorts between 0.238 and 0.389 and were all consistently in the positive direction. The pair HLA-DQA2 and chr12:39994307:C:T (MAF = 0.049) had similar and large effect sizes between the meta pQTL analysis and the PPMI cohort with 0.725 and 0.739, respectively. We did not see any heterogeneity in any of the analyses and all the cohorts contributed to the associations. To further verify result replicability, the associations were examined in a previous study5. In ref. 5, SomaScan1.3 K (1305 proteins) was generated in 971 CSF samples, and the association of LRRK2 variants with GPNMB, GRN, OLR1, and CD63 was reported previously. These analyses indicate that the analyses are robust as all cohorts contribute to the associations and can be replicated in additional studies.
Genetically differentially expressed proteins
Then, we integrated the protein QTL with the latest GWAS for PD risk, under the TWAS/FUSION7 framework to identify genetically driven differential expressed proteins in PD (Fig. 2c). This analysis identified eight proteins (HLA-DQA2, GRN, GPNMB, CD68, C1QTNF1, ENTPD1, SDCBP2, and TLR3) with differentially abundant protein levels in PD cases compared to controls, and two (ITBG2 and TMEM106A) with suggestive association (Supplementary Table 4). Higher levels in PD cases compared to controls were predicted for all proteins.
To validate these results, we used the PPMI data to determine if the levels of the PWAS-significant proteins were significantly different between controls, sporadic, PD cases, or individuals with autosomal dominant PD mutations (LRRK2 + , GBA + , SNCA + ; prodromal cases). Of the 25 proteins (30 aptamers) identified in the initial analyses, 24 aptamers had PPMI protein data measure in 741 PPMI samples and diagnosis. We found 15 proteins had significantly different protein levels between controls and cases, controls and prodromal samples, and/or control and mutation carriers (Supplementary Fig. 4 and Supplementary Table 5), which included the ones predicted by the FUSION approach: HLA-DQA2, GRN, GPNMB, ENTPD1, ITBG2, and TMEM106A were dysregulated in PD cases, as well as even at the prodromal stages.
The PPMI data was also used to determine whether these proteins differed in expression based on sex, and 12 proteins (13 aptamers) did not have sex differences in protein expression in the PPMI dataset or when selecting only case, control, prodromal, LRRK2+, GBA+, or SNCA+ samples (Supplementary Table 6). The remaining 10 proteins (11 aptamers) had significantly different protein levels between male and female samples in PPMI. CD63, FCGR1A, a GPNMB aptamer, and a OLR1 aptamer were significant in only the case samples. GAA, LCT, and LGALS9 had significant sex differences in expression in only the control samples.
Identifying causal proteins for PD risk
Mendelian randomization was performed to estimate the causal effect of PD exposure through links between the PD GWAS8 and pQTL summary statistics. Our results identified seven proteins, out of the 25, to be causally associated to PD risk (SDCBP2, GPNMB, CD68, LCT, ENTPD1, ITGB2, and C1QTNF1) (Table 1, Supplementary Table 7, and Supplementary Fig. 5). To address the horizontal pleiotropy of the LRRK2 locus, we performed MR analyses by removing all the variants in this locus. GPNMB, CD68, and LCT remained significant. To further confirm that pleiotropy did not cause false positive findings, MR analyses were performed including only cis-pQTL, with identical results as the previous analyses. These results indicate that GPNMB, CD68, LCT, and potentially ITGB2 are part of the causal pathway for PD (Table 1).
Biological insights of the proteogenomic analyses
Interaction and pathway analysis
Pathway analysis was performed to understand how proteins interact between them, and if they were enriched in specific pathways. STRING analysis found LRRK2 to interact with GRN and GPNMB and is part of a network that also included CD68 and ITGB2 (Fig. 3a). CD68 is known to interact with GRN, CD63, and ORL1, and predicted to co-express with ITGB2 and FCGR1A. Protein enrichment analyses indicate that the 25 proteins are enriched in proteins involved in leukocyte activation involved in inflammatory response (GO:0002269, P = 0.008) and the microglial cell activation pathway (GO:0001774 microglial cell activation, P = 0.008), along with enrichment in the lysosome cellular component (GO:0005764 lysosome, P = 0.001; Fig. 3b). Gene-disease enrichment analyses also identified an enrichment in lysosomal storage disease genes (C0085078 Lysosomal storage disease, P = 0.004; Fig. 3c). Lysosomal storage disease genes include GRN, GPNMB, GAA, and CHIT1 which, according to STRING analyses, interact with CD68, CD63, and TLR3, and are known to have lysosomal membrane function (Supplementary Fig. 6).
a STRING interactions of the 25 proteins and LRRK2 (red arrow). b EnrichGO of 6138 SomaScan7K genes with recognized unique entrez ID as background and 26 genes of interest (25 proteins and LRRK2). Biological process (BP) and cellular component (CC) are shown. c DisGeNET gene-disease enrichment of 26 genes and 6138 SomaScan7K genes. The top five disease-gene connections are shown. d Brain cell-type fold-change (FC) barplot using 5709 CSF SomaScan7K proteins with brain cell-type data and 25 proteins. *** is FC > |3.7| (P = 3.45 × 10−5) and * is FC > |3.0 | . e Cellular pathways. Pathway involvement of the 25 LRRK2-associated proteins in the endosome, Golgi apparatus, endoplasmic reticulum (ER), nucleus, mitochondrion, cytoplasm, plasma membrane, extracellular space, and lysosome. A large proportion of LRRK2-associated proteins are involved (or enriched) in the endolysosomal pathway. *Most likely involved. +Druggable protein target. Cell structure images are from Bioicons.
Brain cell-type enrichment analysis
In line with the gene enrichment analyses, cell-type enrichment analyses indicate enrichment of microglia-specific proteins (fold change: 3.92, P = 3.45 × 10−5; Fig. 3d; Supplementary Table 8). Nine of the 25 proteins were microglial/macrophage-specific (Supplementary Table 9). Of the eleven PD-associated proteins, our analyses indicate that four (CD68, GRN, HLA-DQA2, TLR3) were microglia-specific, SRI was astrocyte-specific, and GPNMB was oligodendrocyte-specific. LRRK2, GRN, and GPNMB had predominant glial (microglia and oligodendrocyte) cell-type expression and were known to interact based on STRING (Fig. 3a).
Discussion
Genome-wide association studies have identified over 90 loci associated with PD8. However, the genes driving the association for those loci and the functional mechanism by which those signals lead to PD are unknown for most of these loci. In addition, those analyses were not designed to identify complex interactions between proteins. A common approach to identify the functional and causal genes driving the association is QTL mapping combined with colocalization and Mendelian Randomization analyses. This is usually performed with transcriptomic data, but it is now understood that gene expression levels do not correlate with protein levels5. In fact, proteins are likely closer mechanistically to the disease outcome than gene expression. A recent study extended the eQTL approach to protein levels by analyzing pQTL for 1,305 proteins and 131 individuals from the PPMI study5. Kaiser et al.3 found that pQTLs for GPNMB, HLA-DAQ2, and FCGR2A not only colocalized with PD risk loci but were also significant in the MR analyses, implicating those proteins in the causal pathway of PD. The main limitation of this study is that they only included cis-pQTL, missing on additional causal proteins along with protein–protein interactions.
In recent studies by our group4,5, we used cis and trans-pQTL mapping to identify causal and druggable targets for AD and PD. We identified a highly significant trans-pQTL for TREM2 in the MS4A4A region that resolved the MS4A4A loci for AD risk4. We also generated a multi-tissue (brain, CSF, and plasma) pQTL atlas that was integrated with colocalization, MR, and drug repositioning to identify causal proteins and therapies for AD, PD, and other neurodegenerative diseases5. Yang et al.5 identified 35 (brain), 13 (CSF), and 15 (plasma) proteins that were PD causal based on MR with more than 50% colocalizing with PD risk loci. In addition, Yang et al.5 identified seven FDA compounds that target those proteins and in the same direction.
Here, we have increased the sample size of our CSF-published studies to 3107 individuals and focused on the LRRK2 locus to resolve the most significant locus for PD risk and elucidate the downstream mechanism of LRRK2 common variants. Briefly, we identified 25 proteins associated with PD risk, 14 of them not previously reported to be PD-associated. Subsequent TWAS/FUSION-like PWAS approaches determined that ten of those proteins are genetically associated with PD risk, and five were causally linked via MR. Among these proteins, GPNMB and HLA-DAQ2 were already identified in ref. 3 in their cis-only pQTL analyses. In addition, we identified other proteins (CD68 and LCT) that are causal based on all our MR analyses including stringent sensitivity analyses. Other proteins (SDCBP2, ENTPD1, ITGB2, and C1QTNF1) that our MR analyses suggest are causal were genetically associated with PD risk based on the PWAS analyses and some (ENTPD1, ITGB2, and C1QTNF1) showed significant differential protein levels in PD cases compared to controls on the PPMI dataset.
The endolysosomal pathway dysfunction is strongly implicated in the pathobiology of PD1,3,9. Endolysosomal system dysfunction cause disruptions in various essential cellular processes such as endocytosis, phagocytosis, macroautophagy, mitophagy, and/or lysosomal function. LRRK2 is a part of endolysosomal pathway and one of its known functions is the involvement in endosome sorting prior to lysosome fusion10. Interestingly, we found that LRRK2 variants modify the protein level of two well-established PD-associated proteins: GRN and GNMPB. GRN localizes in endosomes and lysosomes and GRN protein levels are known to tightly regulate lysosomal function11. In addition to the localization of GPNMB to plasma membrane, it is also present in intracellular vesicles such as endosomes and lysosomes12. Recently, GPNMB was shown to directly interact with α-synuclein, a disease-defining protein aggregating in PD13. Membrane protein HLA-DQA2, a protein of HLA family that has been previously implicated in PD pathogenesis14, also localizes in endosomes and lysosomes together with lysosome-associated proteins CD6315 and CD6816. Previously, expression of CD68 has been shown to be increased in the p.G2019S LRRK2 transgenic mice compared to control mice upon cerebral injection of α-synuclein pre-formed fibrils17. Importantly, the proteins identified here were not only enriched in the endolysosomal pathway, but other pathways are also related to the intracellular trafficking including the secretory vesicle pathway (GO:0099503 secretory vesicle; P = 0.002), or the vacuole (GO:0005773 vacuole; P = 0.002), indicating that changes in protein/vesicle tracking, and not just the lysosome, are important in PD pathogenesis (Fig. 3e).
In addition to identifying specific biological pathways, our analyses can also inform about the cell types in which these pathways are disrupted. We found that the proteins identified in our study were enriched in microglia (fold change: 3.92; P < 0.05) and immune response (P = 0.008). Microglia dysregulation is known to be implicated in AD, but few studies have identified a strong enrichment of microglia genes/proteins and immune function in PD1,18. Indeed, a large proportion of the proteins identified here play a role in the immune response. Many of these proteins are localized in plasma membrane of microglia and macrophages (such as ENTPD1 and TLR3), which also localize to lysosomes and FCGR1A, or in extracellular space (e.g., LGALS9 and CHIT1), or both (C1QTNF1, OLR1, ITGB2) (Fig. 3e). Intriguingly, many proteins (GPNMB, CD63, and HLA-DQA2) identified to be part of the endolysosomal pathway also localize to the plasma membrane and have a role in immune function19,20,21,22. Interestingly, a recent study found that a common noncoding variant of LRRK2 specifically contributes to risk of PD via microglial LRRK2 expression23. Our findings strongly support the previous observations regarding the importance of microglia and immune system function in the underlying pathobiology of PD. Independent of causality, this is the first time that all these proteins have been connected as part of the same pathways, and this helps to understand the mechanism by which LRRK2 common variants contribute to disease pathogenesis and pathophysiology.
GRN and HLA-DQA2 had discordant effect size direction between the PWAS/FUSION and MR (Supplementary Table 4 and Table 1). Both proteins had a positive direction (higher levels in PD) for the PWAS and a negative direction for the MR analyses. PWAS/FUSION infers the levels of a protein in disease vs controls by examining the genetically regulated association between the proteome level and disease status. PWAS/FUSION is not a causal-inference method, and this prediction can be confounded by secondary events leading to disease and pleiotropy. Our PWAS results were supported by the PPMI differential protein levels analyses, in which GRN and HLA-DQA2 had increased protein levels in the PD prodromal stages and LRRK2+ cases compared to controls (Supplementary Fig. 4). Although PWAS can predict genetic association with PD risk, PWAS/FUSION does not infer causality. MR infers the potential causal effect of proteins on PD exposure through links between the PD GWAS and pQTL summary statistics. MR does not, however, infer protein levels, and PWAS is able to include that missing biological context. For the MR results, both GRN and HLA-DQA2 had negative estimates, indicating that lower levels are leading to higher PD risk. GRN having a negative estimate is in line with the known biology, as lower GRN levels are found in GRN loss of function mutation carriers24. Thus, our PWAS results of positive estimates agree with the PPMI observational findings and our MR results of negative estimates are in line with the known biology. Our interpretation is that PWAS predicts protein levels that can be confound by other factors and not causality. For example, increased CSF protein levels in cases could arise from higher protein release in a dysfunctional lysosome and both GRN11 and HLA-DQA214 are involved in the endolysosomal pathway. Another example of protein levels being in the opposite direction of the causal effect is amyloid-β42 (Aβ42) in AD having higher Aβ42 levels in the brain and lower Aβ42 levels in CSF25. All and all, further work will be needed to address the directionality of GRN and HLA-DQA2 and the implications of discordant MR and PWAS direction results.
Finally, our study paves the way for clinical trials of new therapeutic approaches. The clinical trials targeting ITGB2 (NCT03812263), GAA (NCT00976352), and LCT (NCT02902016, NCT01145586, and NCT05100719) could be translated to PD in the future if found effective. Based on the UniProt DrugBank, there are druggable protein targets for CA1 (DB00819), CHIT1 (DB03539), FCGR1A (DB00112), FTL (DB09147), GAA (DB00284), GPNMB (DB05996), LCT (DB04779), LGALS9 (DB04472), and SRI (DB11348). Interestingly, atorvastatin26 and celastrol27 have been shown to reduce CD68 gene expression, which could similarly be applicable to PD.
In conclusion, trans-pQTL can identify protein interactions in an unbiased manner. This study linked, for the first time, LRRK2 variants with proteins that have support in our PWAS, MR, and PPMI differential protein-level analyses: some previously implicated in PD (GPNMB), and others not (C1QTNF1 and ITGB2). Our results suggest that the CSF concentrations of these proteins (such as GPNMB, C1QTNF1, and ITGB2) have the potential as biomarkers for target engagement and disease progression modification in clinical trials that target LRRK2.
Methods
Study design
We performed Phenome-wide Association Studies (PheWAS) of the sentinel SNP in LRRK2 (rs76904798) using a CSF protein QTL atlas that included more than 3000 samples to identify causal and druggable proteins for Parkinson’s disease (PD). We performed conditional analyses (GCTA-COJO) in this region, using those proteins, to identify additional independent SNPs modifying protein levels and PD risk. Next, we leveraged novel statistical approaches to identify genetically associated protein levels (Proteome-wide Association Studies (PWAS)) and causal proteins (MR). Finally, we performed pathway and cell-type enrichment analyses to determine the biological processes that these proteins are involved in, which major cell types they are expressed in, and how they interact with each other.
Cohorts
Protein QTL was performed in a total of 3107 unrelated, Non-Hispanic White samples recruited from six different cohorts: Dominantly Inherited Alzheimer Network (DIAN), the Charles F. and Joanne Knight Alzheimer Disease Research Center (Knight ADRC), Barcelona-1, Alzheimer’s Disease Neuroimaging Initiative (ADNI), Ace Alzheimer Center Barcelona (Fundació ACE), and Parkinson’s Progression Markers Initiative (PPMI) (Supplementary Table 1). DIAN is a longitudinal study of dominantly inherited AD mutation carriers and their family members including asymptomatic and mild to moderate AD. Knight ADRC Memory and Aging Project (MAP) is a longitudinal study of cognitive functioning in persons as they age, with the goal to advance AD research. Barcelona-1 is a longitudinal observational study of dementias such as AD dementia (ADD) and mild cognitive impairment (MCI) at the University Hospital Mutua de Terrassa, Barcelona, Spain. ADNI is a multisite study that aims to develop biomarkers that will improve clinical trials for the prevention and treatment of AD. PPMI is a longitudinal observational study of participants with and without PD, with comprehensive clinical and imaging data and biological samples aimed at identifying markers of disease progression for use in clinical trials of therapies to reduce the progression of PD disability.
Phenome-wide association study and conditional analyses
Proteins were measured using the SomaLogic aptamer-based Somascan platform. DIAN, Knight ADRC, Barcelona-1, ADNI, and Fundació ACE CSF samples were measured using SomaScan7K, which measured 7006 aptamers in 6139 protein targets. PPMI generated the CSF proteomic data using SomaScan5K, and 4785 aptamers in 4131 protein targets were measured. Genotyping and protein measurements have been described in detail elsewhere5. Proteomics generation was performed using the Somalogic platform in a total of 3107 unrelated non-Hispanic white individuals. Protein QC and processing followed the steps described elsewhere28. Initial normalization of the SomaScan7K data was performed by SomaLogic with median normalization to remove sample preparation biases and reference normalization to account for technical and biological variance. Aptamers were then removed if the aptamer’s scale factor diverged from the median scale factor >0.5 or if the median cross-plate coefficient of variation (CV) was >0.15. Aptamer values within 1.5-fold of the IQR (Interquartile range) were kept as the value. Samples and aptamers with call rate <65% were removed and the 85% call rate was applied to aptamers and samples separately after call rate recalculation. For the SomaScan5K data, scale factor and CV filtering was not performed due to the information not being available in the PPMI cohort. Using the same methods as the SomaScan7K dataset, IQR and call rate quality control was performed in the SomaScan5K dataset. Protein levels were then z-score normalized by log10-scale and values were centered to a mean around 0 and variance of 1 using the R function scale.
There were 3726 samples, with Hg38 GWAS and CSF proteomic data available, included in this study. After genetic principal component analysis (PCA) using the 1000 Genome to select European ancestry, 3328 samples remained. The 1000 Genome Project was used for ancestry grouping to select the non-Hispanic White population with gPC1 <0.005 and gPC2 >−0.01 in the PCA, and samples outside of three standard deviations from the mean were removed. After removing individuals with cryptic relatedness through identity by descent (IBD; PIHAT ≥ 0.25), 3107 samples remained. IBD was performed using certain variants (MAF ≥ 0.15 or HWE P ≥ 0.001, R2 = 0.2), and one sample from each cryptic-related pair (PIHAT ≥ 0.25) was removed. pQTL linear regression was then performed using PLINK29 with the normalized protein levels5, adjusting by age, sex, the first ten principal components, and technical variables (genotyping array). For PD risk, we used the latest published GWAS, which included 2,525,897 total samples in the PD GWAS8.
We first determined what proteins were associated with the sentinel SNP on the LRRK2 locus (rs76904798; chr12:40220632:C:T) by performing a PheWAS for rs76904798. Multiple test correction was performed at FDR 5% correction using R package FDRestimation30. We then performed conditional and joint association analysis (GTCA-COJO)31,32 on each significant aptamer to identify independently associated SNPs (Supplementary Table 2) on this locus. SNPs with r2 > 0.85 per LD block were pruned to keep independently associated SNPs without non-random association. Pearson correlation was then performed using the R package corrplot33 to determine correlation between proteins.
pQTL mapping was performed on the six independent cohorts using PLINK29 with linear regression on the normalized protein levels and adjusted by the same covariates as above (age, sex, PC 1-10, and genotyping array). Signal consistency was examined in each independent dataset using forest plots of effect sizes from the linear regression analyses and compared to the effect sizes of the joint and meta pQTL analyses. I2 heterogeneity P values were gathered from the fixed-effect meta pQTL analyses performed in METAL using inverse-variance weighting per aptamer. Effect sizes of the joint analyses was then compared to the effect sizes of shared aptamers in ref. 5 with SomaScan 1.3K measurements from 971 CSF samples to verify replicability.
Proteome-wide association study
A proteome-wide association study (PWAS) was performed to identify associations between PD risk and protein levels. PWAS7 can identify proteins potentially involved in PD etiology by examining genetically regulated association between proteome expression and PD status. Whereas these approaches were built using only cis signal expression data of the transcriptome, we modified the FUSION7 framework to include proteome expression weight files containing both cis and trans signals of the genes. Age, sex, the first ten genetic principal components, and genotyping array were included as covariates in the protein weight calculation model of the PWAS. The functional GWAS association statistic was calculated using the FUSION gusevlab GitHub R script7 and protein weights of signals on chromosome 12. FUSION post-processing was then performed to identify conditionally independent associated features by analyzing significant associations within a locus window boundary of 100,000 bp. Three strictly nonoverlapping loci were found with the second locus on the 39 to 41.5 Mb region of chromosome 12.
Validating protein-level association with the PPMI cohort
PPMI used DNA samples from blood or saliva for genetic mutation testing of LRRK2 (p.G2019S and p.R1441G), GBA (p.N370S), and SNCA mutations. PPMI generated the proteomic data on 1075 samples using the aptamer-based SomaScan5K platform. A total of 4783 aptamers remain after QC: outliers were removed by 1.5 IQR threshold, aptamer and individual call rate <65%, and aptamer and individual call rate <85%. There were 917 samples present with PPMI phenotypes including 185 healthy control, 545 PD, and 187 prodromal samples. Only samples with proteomic and clinical data were included in the analyses. Linear regression was performed on the normalized protein levels of PPMI using PLINK29 adjusted by age, sex, and cohortArray. Differential protein levels analyses were performed with control samples compared to case, prodromal, and mutation samples. Differential protein-level analyses by sex were also performed with male (n = 422) and female (n = 314) samples compared in all samples with PPMI cohort protein data and in only case, control, prodromal, and mutation samples.
Mendelian randomization
Mendelian randomization (MR) was performed using the summary statistics of each of the significant aptamer pQTLs generated here, and the PD GWAS8 to investigate the potential causal effect of the proteins on PD. Using the R package TwoSampleMR34,35, 21 aptamers from 17 proteins had independent genome-wide significant SNPs (P < 5 × 10−8) present in the pQTL and the PD GWAS. IVW (inverse-variance-weighted) P values, or Wald ratio P values for one SNP MR, were reported. Since LRRK2 is in a pleiotropic region and to avoid potential false positives, we performed the analyses again after removing the LRRK2 locus (39–41.5 Mb region), on aptamers with cis signal in the MR analyses, and then performed MR on only the cis-pQTL SNP. Cis-pQTL signal was defined as the SNP being within 1 Mb upstream or downstream of the hg38 coding region of the protein-encoding gene. F-statistics were calculated for SNPs as instruments to determine the instrument strengths are at least 10 (Supplementary Table 7).
Protein–protein interaction and pathway analysis
Pathway and interaction analyses were performed using STRING36, EnrichGO37, DisGeNET37, and GeneMANIA38. STRING was performed by entering the 25 genes that code for the proteins identified in this study together with LRRK2 for gene interactions. EnrichGO was performed with 6138 SomaScan7K genes as background and 26 genes of interest (25 proteins and LRRK2). Gene-disease enrichment was performed using DisGeNET of the 26 genes and 6138 SomaScan7K genes. Network analysis was performed using GeneMANIA with the default 20 affiliated genes in a network with the 25 genes and LRRK2.
Brain cell enrichment analysis
Cell-type enrichment analysis was performed on the 5706 genes in CSF SomaScan7K with cell-type data as background and the associated proteins as the target set. Brain cell types were determined using transcriptome expression data39 in microglial/macrophage, endothelial, mature astrocyte, neuronal, and oligodendrocyte cells isolated from human brains. Gene expression data was added for all the samples per gene, and percentiles were calculated per cell type. If one cell type had more than 50% of the sum-total expression of a gene, then that gene was considered as cell-type specific. Fold change (FC) was calculated using the ratio between cell percentage for each protein associated to LRRK2 and the SomaScan7K background set.
Data availability
Proteomic, pQTL, and raw data from the Knight ADRC participants are available at the NIAGADS and can be accessed at https://www.niagads.org/knight-adrc-collection. Data from the DIAN cohort is available to qualified investigators and can be requested at https://dian.wustl.edu/our-research/for-investigators/diantu-investigator-resources/dian-tu-biospecimen-request-form/. ADNI and PPMI proteomic data can be found at https://adni.loni.usc.edu/ and https://www.ppmi-info.org/, respectively.
References
Nalls, M. A. et al. Identification of novel risk loci, causal insights, and heritable risk for Parkinson’s disease: a meta-analysis of genome-wide association studies. Lancet Neurol. 18, 1091–1102 (2019).
Lake, J. et al. Coding and noncoding variation in LRRK2 and Parkinson’s disease risk. Mov. Disord. 37, 95–105 (2022).
Kaiser, S. et al. A proteogenomic view of Parkinson’s disease causality and heterogeneity. NPJ Parkinsons Dis. 9, 24 (2023).
Deming, Y., Filipello, F. & Cignarella, F. The MS4A gene cluster is a key modulator of soluble TREM2 and Alzheimer’s disease risk. Sci. Transl. Med. 11, eaau2291 (2019).
Yang, C. et al. Genomic atlas of the proteome from brain, CSF and plasma prioritizes proteins implicated in neurological disorders. Nat. Neurosci. 24, 1302–1312 (2021).
Gu, Z., Gu, L., Eils, R., Schlesner, M. & Brors, B. circlize implements and enhances circular visualization in R. Bioinformatics 30, 2811–2812 (2014).
Gusev, A. et al. Integrative approaches for large-scale transcriptome-wide association studies. Nat. Genet. 48, 245–52 (2016).
Kim, J. J. et al. Multi-ancestry genome-wide meta-analysis in Parkinson’s disease. Preprint at https://www.medrxiv.org/content/10.1101/2022.08.04.22278432v1 (2022).
Vidyadhara, D. J., Lee, J. E. & Chandra, S. S. Role of the endolysosomal system in Parkinson’s disease. J. Neurochem. 150, 487–506 (2019).
Schreij et al. LRRK2 localizes to endosomes and interacts with clathrin-light chains to limit Rac1 activation. EMBO Rep. 16, 79–86 (2015).
Kao, A. W., McKay, A., Singh, P. P., Brunet, A. & Huang, E. J. Progranulin, lysosomal regulation and neurodegenerative disease. Nat. Rev. Neurosci. 18, 325–333 (2017).
van der Lienden, M. J. C., Gaspar, P., Boot, R. Aerts, J. M. F. G. & van Eijk, M. Glycoprotein non-metastatic protein B: an emerging biomarker for lysosomal dysfunction in macrophages. Int. J. Mol. Sci. 20, 66 (2018).
Diaz-Ortiz, M. E. et al. GPNMB confers risk for Parkinson’s disease through interaction with α-synuclein. Science 377, eabk0637 (2022).
Naito, T. et al. Trans-ethnic fine-mapping of the major histocompatibility complex region linked to Parkinson’s Disease. Mov. Disord. 36, 1805–1814 (2021).
Fukuda M. Lysosomal membrane glycoproteins. Structure, biosynthesis, and intracellular trafficking. J. Biol. Chem. 266, 21327–21330 (1991).
Holness, C. L. & Simmons, D. L. Molecular cloning of CD68, a human macrophage marker related to lysosomal glycoproteins. Blood 81, 1607–13 (1993).
Bieri, G. et al. LRRK2 modifies α-syn pathology and spread in mouse models and human neurons. Acta Neuropathol. 137, 961–980 (2019).
Tan, E. K. et al. Parkinson disease and the immune system -associations, mechanisms and therapeutics. Nat. Rev. Neurol. 16, 303–318 (2020).
Saade, M., Araujo de Souza, G., Scavone, C. & Kinoshita, P. F. The role of GPNMB in inflammation. Front. Immunol. 12, 674739 (2021).
Petersen, S. H. et al. The role of tetraspanin CD63 in antigen presentation via MHC class II. Eur. J. Immunol. 41, 2556–61 (2011).
Rudy, G. B. & Lew, A. M. The nonpolymorphic MHC class II isotype, HLA-DQA2, is expressed on the surface of B lymphoblastoid cells. J. Immunol. 158, 2116–25 (1997).
Houser, M. C. et al. Progranulin loss results in sex-dependent dysregulation of the peripheral and central immune system. Front. Immunol. 13, 1056417 (2022).
Langston, R. G. et al. Association of a common genetic variant with Parkinson’s disease is mediated by microglia. Sci. Transl. Med. 14, eabp8869 (2022).
Meeter, L. H. et al. Progranulin levels in plasma and cerebrospinal fluid in granulin mutation carriers. Dement. Geriatr. Cogn. Dis. Extra 6, 330–340 (2016).
Sturchio, A. et al. High soluble amyloid-β42 predicts normal cognition in amyloid-positive individuals with Alzheimer’s disease-causing mutations. J. Alzheimers Dis. 90, 333–348 (2022).
Llaverias, G. et al. Atorvastatin reduces CD68, FABP4, and HBP expression in oxLDL-treated human macrophages. Biochem. Biophys. Res. Commun. 318, 265–74 (2004).
Cascão, R. et al. Decrease of CD68 synovial macrophages in celastrol treated arthritic rats. PLoS ONE 10, e0142448 (2015).
Timsina, J. et al. Comparative analysis of Alzheimer’s disease Cerebrospinal fluid biomarkers measurement by multiplex SOMAscan platform and immunoassay-based approach. J. Alzheimers Dis. 89, 193–207 (2022).
Purcell, S. et al. PLINK: a toolset for whole-genome association and population-based linkage analysis. Am. J. Hum. Genet. 81, 559–75 (2007).
Murray, M. H. & Blume, J. D. FDRestimation: flexible false discovery rate computation in R. F1000Research. 10, 441 (2021).
Yang, J. et al. Conditional and joint multiple-SNP analysis of GWAS summary statistics identifies additional variants influencing complex traits. Nat. Genet. 44, 369–375 (2012).
Yang, J., Lee, S. H., Goddard, M. E. & Visscher, P. M. GCTA: a tool for genome-wide complex trait analysis. Am. J. Hum. Genet 88, 76–82 (2011).
Wei, T. & Simko, V. R package ‘corrplot’: visualization of a Correlation Matrix. (Version 0.92). https://github.com/taiyun/corrplot. (2021).
Hemani, G. et al. The MR-Base Collaboration. The MR-Base platform supports systematic causal inference across the human phenome. eLife 7, e34408 (2018).
Hemani, G., Tilling, K. & Davey Smith, G. Orienting the causal relationship between imprecisely measured traits using GWAS summary data. PLoS Genet. 13, e1007081 (2017).
Warde-Farley, D. et al. The GeneMANIA prediction server: biological network integration for gene prioritization and predicting gene function. Nucleic Acids Res. 38, W214–20 (2010).
Yu, G., Wang, L., Han, Y. & He, Q. clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS: A J. Integr. Biol. 16, 284–287 (2012).
Szklarczyk, D. et al. The STRING database in 2021: customizable protein-protein networks, and functional characterization of user-uploaded gene/measurement sets. Nucleic Acids Res. 49, D605–D612 (2021).
Zhang, Y. et al. Purification and characterization of progenitor and mature human astrocytes reveals transcriptional and functional differences with mouse. Neuron 89, 37–53 (2016).
Acknowledgements
We thank all the participants and their families, as well as the many involved institutions and their staff. This work was supported by access to equipment made possible by the Hope Center for Neurological Disorders, the Neurogenomics and Informatics Center (NGI: https://neurogenomics.wustl.edu/) and the Departments of Neurology and Psychiatry at Washington University School of Medicine.
ADNI resources: Data collection and sharing for this project was funded by the Alzheimer’s Disease Neuroimaging Initiative (ADNI) (National Institutes of Health Grant U01 AG024904) and DOD ADNI (Department of Defense award number W81XWH-12-2-0012). ADNI is funded by the National Institute on Aging, the National Institute of Biomedical Imaging and Bioengineering, and through generous contributions from the following: AbbVie, Alzheimer’s Association; Alzheimer’s Drug Discovery Foundation; Araclon Biotech; BioClinica, Inc.; Biogen; Bristol-Myers Squibb Company; CereSpir, Inc.; Cogstate; Eisai Inc.; Elan Pharmaceuticals, Inc.; Eli Lilly and Company; EuroImmun; F. Hoffmann-La Roche Ltd and its affiliated company Genentech, Inc.; Fujirebio; GE Healthcare; IXICO Ltd.; Janssen Alzheimer Immunotherapy Research & Development, LLC.; Johnson & Johnson Pharmaceutical Research & Development LLC.; Lumosity; Lundbeck; Merck & Co., Inc.; Meso Scale Diagnostics, LLC.; NeuroRx Research; Neurotrack Technologies; Novartis Pharmaceuticals Corporation; Pfizer Inc.; Piramal Imaging; Servier; Takeda Pharmaceutical Company; and Transition Therapeutics. The Canadian Institutes of Health Research is providing funds to support ADNI clinical sites in Canada. Private sector contributions are facilitated by the Foundation for the National Institutes of Health (www.fnih.org). The grantee organization is the Northern California Institute for Research and Education, and the study is coordinated by the Alzheimer’s Therapeutic Research Institute at the University of Southern California. ADNI data are disseminated by the Laboratory for Neuro Imaging at the University of Southern California.
DIAN resources: Data collection and sharing for this project was supported by The Dominantly Inherited Alzheimer Network (DIAN, U19AG032438) funded by the National Institute on Aging (NIA), the Alzheimer’s Association (SG-20-690363-DIAN), the German Center for Neurodegenerative Diseases (DZNE), Raul Carrea Institute for Neurological Research (FLENI), Partial support by the Research and Development Grants for Dementia from Japan Agency for Medical Research and Development, AMED, and the Korea Health Technology R&D Project through the Korea Health Industry Development Institute (KHIDI), Spanish Institute of Health Carlos III (ISCIII), Canadian Institutes of Health Research (CIHR), Canadian Consortium of Neurodegeneration and Aging, Brain Canada Foundation, and Fonds de Recherche du Québec – Santé. This manuscript has been reviewed by DIAN Study investigators for scientific content and consistency of data interpretation with previous DIAN Study publications. We acknowledge the altruism of the participants and their families and the contributions of the DIAN research and support staff at each of the participating sites for their contributions to this study.
ACE Alzheimer Center Barcelona acknowledges all patients and their families for their collaboration. For CSF biomarker research, A.R. and M.B. received support from the European Union/EFPIA Innovative Medicines Initiative Joint undertaking ADAPTED and MOPEAD projects (grant numbers 115975 and 115985, respectively). M.B. and A.R. are also supported by national grants PI13/02434, PI16/01861, PI17/01474, PI19/01240, PI19/01301, PI22/01403 from the Acción Estratégica en Salud, integrated in the Spanish National RCDCI Plan and funded by Instituto de Salud Carlos III (ISCIII)—Subdirección General de Evaluación and the Fondo Europeo de Desarrollo Regional (FEDER—“Una manera de Hacer Europa”). A.R. and M.B. have also received support from CIBERNED (Instituto de Salud Carlos III (ISCIII). A.R. is also supported by the EXIT project, EU Euronanomed3 Program JCT2017, Grant No. AC17/00100 and PREADAPT project; the Joint Program for Neurodegenerative Diseases (JPND), Grant No. AC19/00097; Acción Estratégica en Salud, integrated in the Spanish National RCDCI Plan and funded by Instituto de Salud Carlos III (ISCIII)—Subdirección General de Evaluación and the Fondo Europeo de Desarrollo Regional (FEDER—“Una manera de Hacer Europa”). I. de Rojas is supported by a national grant from the Instituto de Salud Carlos III FI20/00215. *Data used in the preparation of this article were obtained from the Alzheimer’s Disease. Neuroimaging Initiative (ADNI) database (adni.loni.usc.edu). As such, the investigators within the ADNI contributed to the design and implementation of ADNI and/or provided data but did not participate in the analysis or writing of this report. A complete listing of ADNI investigators can be found at http://adni.loni.usc.edu/wp-content/uploads/how_to_apply/ADNI_Acknowledgement_List.pdf. This work was supported by grants from the National Institutes of Health (R01AG044546 (C.C.), P01AG003991(CC, JCM), RF1AG053303 (C.C.), RF1AG058501 (C.C.), U01AG058922 (C.C.), RF1AG074007 (Y.J.S.)), the Chuck Zuckerberg Initiative (C.Z.I.), the Michael J. Fox Foundation (L.I. and C.C.), and the Department of Defense (L.I.—W81XWH2010849). The recruitment and clinical characterization of research participants at Washington University were supported by NIH P30AG066444 (J.C.M.), P01AG03991 (J.C.M.), and P01AG026276 (J.C.M.).
Author information
Authors and Affiliations
Consortia
Contributions
B.P. wrote the manuscript with support from C.C. and performed the analyses. D.W. generated the CSF pQTL atlas. L.W. and C.Y. performed prior analyses on LRRK2 and GRN that conceptualized the analysis framework. J.T. performed proteomics quality control and data processing. Y.S. and L.I. processed the PPMI cohort data. P.G. performed data processing on the PD GWAS8. C.Y. performed genomics data processing. A.D. contributed to pathway analysis interpretation. NN contributed to protein cellular localization and endolysosomal pathway interpretations. I.A. and M.A. are part of the Memory Disorders Unit. P.P., S.E.S., A.M.F., A.R., and J.S.P. contributed to data processing supervision. R.P., P.G., I.R., M.M., M.B., and A.R. are part of Ace Alzheimer Center Barcelona and generated the Fundació ACE cohort data. J.C.M., A.M.F., and R.J.P. are part of the Dominantly Inherited Alzheimer Network (DIAN) Consortia. L.I. and Y.S.J. helped supervise the project. C.C. supervised the project. All authors reviewed the final manuscript.
Corresponding author
Ethics declarations
Competing interests
C.C. has received research support from: GSK and EISAI. The funders of the study had no role in the collection, analysis, or interpretation of the data; in the writing of the report; or in the decision to submit the paper for publication. C.C. is a member of the advisory board of Vivid Genomics and Circular Genomics. S.E.S. has analyzed data provided by C2N Diagnostics to Washington University, but she has not received any research funding or personal compensation from C2N Diagnostics or any other for-profit organizations. Authors B.P., D.W., L.W., J.T., Y.S., P.G., C.Y., A.D., N.N., I.A., M.A., P.P., J.C.M., A.M.F., R.P., P.G., I.R., M.M., M.B., A.R., J.S.P., L.I., R.J.P., and Y.S.J. declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Phillips, B., Western, D., Wang, L. et al. Proteome wide association studies of LRRK2 variants identify novel causal and druggable proteins for Parkinson’s disease. npj Parkinsons Dis. 9, 107 (2023). https://doi.org/10.1038/s41531-023-00555-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41531-023-00555-4
This article is cited by
-
Connectome-based modelling of neurodegenerative diseases: towards precision medicine and mechanistic insight
Nature Reviews Neuroscience (2023)