Abstract
Epigenetic gene silencing induced by expanded repeats can cause diverse phenotypes ranging from severe growth defects in plants to genetic diseases such as Friedreich’s ataxia in humans. The molecular mechanisms underlying repeat expansion-induced epigenetic silencing remain largely unknown. Using a plant model with a temperature-sensitive phenotype, we have previously shown that expanded repeats can induce small RNAs, which in turn can lead to epigenetic silencing through the RNA-dependent DNA methylation pathway. Here, using a genetic suppressor screen and yeast two-hybrid assays, we identified novel components required for epigenetic silencing caused by expanded repeats. We show that FOURTH ULP GENE CLASS 1 (FUG1)—an uncharacterized SUMO protease with no known role in gene silencing—is required for epigenetic silencing caused by expanded repeats. In addition, we demonstrate that FUG1 physically interacts with ALFIN-LIKE 3 (AL3)—a histone reader that is known to bind to active histone mark H3K4me2/3. Loss of function of AL3 abolishes epigenetic silencing caused by expanded repeats. AL3 physically interacts with the chromodomain protein LIKE HETEROCHROMATIN 1 (LHP1)—known to be associated with the spread of the repressive histone mark H3K27me3 to cause repeat expansion-induced epigenetic silencing. Loss of any of these components suppresses repeat expansion-associated phenotypes coupled with an increase in IIL1 expression with the reversal of gene silencing and associated change in epigenetic marks. Our findings suggest that the FUG1–AL3–LHP1 module is essential to confer repeat expansion-associated epigenetic silencing and highlight the importance of post-translational modifiers and histone readers in epigenetic silencing.
Similar content being viewed by others
Main
Variation in the length of short tandem repeats is associated with diverse phenotypic changes across organisms1,2,3,4,5,6. Extreme length variants exemplified by repeat expansions cause genetic defects such as Huntington’s disease and Friedreich’s ataxia in humans to growth defects in plants3,7,8. Recent explosion in genomic data has resulted in the discovery of novel repeat expansion-associated phenotypes in multiple organisms and new diseases including cancer9,10,11,12. Repeat expansions are seen in plant growth defects7 and, in diseases such as Friedreich’s ataxia, cause gene silencing, but the underlying mechanisms remain largely unknown13,14,15.
The Bur-0 accession of Arabidopsis carries a GAA/TTC repeat expansion in the third intron of the ISOPROPYLMALATE ISOMERASE LARGE SUBUNIT 1 (IIL1) gene associated with the epigenetic silencing at the IIL1 locus7. Triplet expansion-induced reduction in IIL1 expression leads to a temperature-sensitive growth defect referred to as the irregularly impaired leaves (iil) phenotype (narrower and twisted leaves) when grown at 27 °C (ref. 7). We have previously shown that the GAA/TTC repeat expansion at this locus leads to accumulation of 24 nt small interfering RNAs that map to IIL1, in a temperature-dependent manner, and through the RNA-directed DNA methylation (RdDM) pathway leads to epigenetic silencing16.
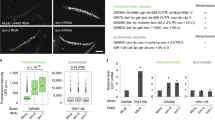
To decipher molecular pathways that mediate repeat expansion-associated gene silencing, we carried out a new large-scale ethyl methane sulfonate (EMS) mutagenesis screen in the Bur-0 (iil mutant) background (Fig. 1a). We quantified IIL1 expression to identify suppressors that act at the level of gene expression and identified a total of nine suppressors with elevated IIL1 levels (Fig. 1b). F1 progeny of all nine genetic suppressors in a cross with Bur-0 revealed that they were recessive in nature. We sequenced the mutants and used a combination of SHOrt REad mapping (SHOREmap)17 and positional cloning to identify the potential underlying mutations in these nine genetic suppressors of iil. For four mutants, we sequenced a pool of 500 segregating plants from an F2 population and mapped potential loss-of-function alleles at At3g57080 (RPB5B, an RNA Pol V subunit in 44-2), At2g40030 (NRPD1B, an RNA Pol V subunit in 49-9), At2g27040 (AGO4 in 57-3) and a mis-sense mutation at At3g23780 (NRPD2A, an RNA Pol IV subunit in 61-7) (Fig. 1c and Extended Data Fig. 1). We have previously shown that knocking down At2g40030, At3g23780 and At2g27040 in the Bur-0 background suppresses the iil phenotype, and therefore, we conclude that the observed mutations are causal. As SHOREmap analysis and our earlier findings16 confirmed the importance of the RdDM pathway for repeat expansion-induced gene silencing, we analysed the rest of the mutants by direct sequencing and looked for mutations in RdDM pathway genes. These analyses revealed that four other suppressors harboured mutations in At4g11130 (RDR2 in 71-9) or in At2g27040 (AGO4 in 72-6, 81-2 and 108-8) (Supplementary Table 1). In summary of the nine genetic suppressors, eight were mutants in RdDM pathway genes. We conclude that RdDM is the major pathway by which expanded repeats cause epigenetic silencing in Arabidopsis thaliana.
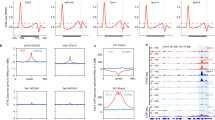
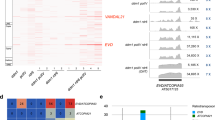
a, Phenotypes (denoted by their original screen identifiers) of the isolated suppressors compared to Bur-0. The irregularly impaired leaves are marked by white arrows in the Bur-0 wild type. Scale bars, 2 cm. b, Relative IIL1 expression levels in genetic suppressors identified through the genetic screen. The numbers represent the original screen identifiers, and the corresponding genes identified after cloning are shown below. Average expression levels based on three biological replicates for each line (except for Bur-0 and fug1, where n = 5 and 4, respectively) are shown. Asterisks (*) denote individual data points. P values are based on one-way analysis of variance with Tukey’s post hoc test, and lines with different letters are significantly different from each other (P < 0.05). Error bars represent s.e.m. c, An example of SHOREmap analysis with 44-2 identifies a mutation in Pol V. High-frequency alleles (>0.85) are coloured red, and red crosses show the putative causal alleles.
One mutant (4b) did not carry mutations in the candidates for the RdDM pathway but suppressed the temperature-dependent iil phenotype despite the repeat expansion (Fig. 1a and Extended Data Fig. 2). Sequence analysis failed to discover any major-effect mutations within a 250 kb region surrounding the IIL1 locus, which indicated that 4b is a second-site suppressor. We detected 34 potential loss-of-function mutations in 4b, but none of them were in obvious candidate genes (Supplementary Table 2). We crossed the 4b mutant with Pf-0 (IIL wild type), as the original repeat expansion-associated iil phenotype in the Bur-0 background segregated as a monogenic trait in this cross7. We then selected lines that are homozygous for IIL1 and still harbouring the repeat expansions and used the progeny of these lines for mapping 4b. By linkage mapping using a total of more than 750 mutant plants, we mapped 4b to a 90 kb region on chromosome 3, containing 20 protein-coding genes (Fig. 2a). We identified only a single EMS-type single-nucleotide polymorphism (SNP) within this 90 kb that was in the first exon of At3g48480 at position 17958141 (corresponding to The Arabidopsis Information Resource 10 (TAIR10) in Col-0), which changes Gln into a stop codon (Q152*) making it a potential candidate gene (Fig. 2a). At3g48480 encodes a protein referred to as FOURTH UBIQUITIN-LIKE PROTEASE GENE CLASS 1 (FUG1)18. To test whether FUG1 is 4b, we designed two independent artificial microRNAs (amiRNAs, 35S::amiR-FUG1) and generated knockdown lines in the Bur-0 (iil mutant) background (Fig. 2b). One hundred and ninety of the 200 independent 35S::amiR-FUG1 lines in the Bur-0 background showed suppression of the iil phenotype (Fig. 2b), confirming that the loss of function of FUG1 is sufficient to suppress the iil phenotype. Conversely, overexpression of FUG1 (35S::FUG1) in the 4b background restored the iil phenotype (Fig. 2b). Henceforth the 4b will be referred to as the fug1 mutant.
a, Positional cloning of the 4b/fug1 mutant. Analysis at F2 and F3 is shown in two stages, and subsequently the 90 kb interval was analysed through sequencing. Rf refers to the recombination frequency. b, At3g48480 is FUG1, and its protease domain is essential for its function. Phenotypes of plants harbouring 35S::amiR-FUG1 in Bur-0 and 35S::FUG1 in fug1 are shown. c, Western blot analysis with anti-SUMO antibodies on total protein extracts from Bur-0 and fug1. The pink asterisk represents a protein that is hyperSUMOylated in fug1 compared to Bur-0. The same protein extract probed with anti-actin is shown as the loading control. Two biological replicates were tested, which gave similar results. Protein markers are shown in kDa. d, In vitro deSUMOylation assay shows FUG1 is a deSUMOylase. HIS-SUMO-FLC (shown in blue dots) is incubated with or without HIS-FUG1, and the products were analysed through western blot with anti-SUMO antibodies. Free SUMO is shown by the red arrow, and HIS-SUMO-FLC is shown by the blue arrow. The experiment was repeated at least two times with similar results. e, FUG1 protease domain is required for its biological function. Phenotypes of plants transformed with the pFUG1::mFUG1 that harbours the mutation in the protease site along with the control pFUG1::FUG1 in the fug1 background are shown. Scale bars in b and e, 2 cm.
FUG1 belongs to a highly conserved superfamily of cysteine-type proteases, and its related members in Arabidopsis are potential small ubiquitin-like modifier (SUMO) proteases involved in deSUMOylation18,19. Among the SUMO proteases in Arabidopsis, FUG1 remains an uncharacterized protein19. Mutants for the rice orthologue of FUG1 show an increase in SUMOylation levels20. Western blots with anti-SUMO antibodies on total protein extracts revealed increased SUMOylation in fug1 mutants compared to Bur-0 (Fig. 2c), which suggests that FUG1 could function as a deSUMOylase in Arabidopsis. To directly test whether FUG1 can function as a deSUMOylase, we carried out an in vitro deSUMOylase assay in which purified FUG1 protein was tested for its ability to cleave SUMO from purified SUMOylated FLOWERING LOCUS C (HIS-SUMO-FLC) protein. We observed the release of SUMO upon incubation with purified HIS-FUG1, which showed that FUG1 can indeed function as a deSUMOylase (Fig. 2d).
To assess the impact of FUG1 on transcription, we compared the RNA expression profiles of plants Bur-0 and fug1. We detected a total of 3,505 out of 21,964 analysable genes to be differentially expressed between Bur-0 and fug1 (Supplementary Table 3), which indicated that FUG1 had a substantial (~16%) impact on the transcriptome. IIL1 was among the top upregulated genes in the fug1 background. Gene Ontology enrichment analysis revealed a connection between response to stimuli and post-translational modifications among the differentially expressed genes between Bur-0 and fug1 consistent with FUG1 as a deSUMOylase.
To test whether the protease function of FUG1 is required for the observed iil phenotype, we disrupted the catalytic triad in the FUG1 protein (C246S) and generated transgenic plants expressing either wild-type FUG1 (pFUG1:FUG1) or mutated FUG1 (pFUG1::mFUG1) under its native promoter and transformed them into the fug1 mutant background. pFUG1::FUG1 plants in the fug1 background showed the iil phenotype (restoring to Bur-0 situation) at 27 °C showing that the construct is functional (Fig. 2e). Conversely, pFUG1::mFUG1 plants failed to complement the fug1 mutant (normal-looking plant) confirming the requirement of the protease domain for FUG1 function (Fig. 2e). We conclude that FUG1 encodes a SUMO protease that is required to confer repeat expansion-associated gene silencing.
FUG1 was localized in the nucleus of 35S::GFP-FUG1 plants (Extended Data Fig. 3). To identify potential FUG1 interactors, we performed a yeast two-hybrid screen and recovered a plant homeodomain (PHD) finger containing nuclear protein ALFIN-LIKE 3 (AL3)21. PHD-finger-containing proteins are known to be histone H3K4me2/3 readers22,23. FUG1–AL3 interaction was confirmed in yeast cells (Fig. 3a). Bimolecular fluorescence complementation (BiFC) assays with co-transfection on tobacco leaf epidermal cells confirmed the in planta nuclear interaction of FUG1 and AL3 (Fig. 3b).
a, FUG1 interacts with AL3 but not LHP1 in yeast cells. AD and BD refer to activation and binding domains, respectively. b, FUG1 interacts with AL3 in planta in BiFC assays using nYFP and cCFP fused proteins in tobacco epidermal cells. PYL6 and CIS1 are used as negative controls in these assays, respectively, for C- and N-terminal fusion constructs. GFP, GFP channel; BF, bright field. c–e, Knocking down AL3 suppresses the iil phenotype in 35S::amiR-AL3 plants (c) with downregulation of AL3 expression (d) and corresponding increase in IIL1 expression (e). Each dot represents individual wild-type Bur-0 (n = 6) or 35S::amiR-AL3 primary independent (T1) transgenic lines (n = 27) and relative expression levels are normalized with tubulin levels and shown in arbitrary units. Scale bar, 2 cm. f, AL3 is SUMOylated in tobacco cells. SUMOylation assays with Myc-AL3 and HA-SUMO in tobacco cells. Blots were immunoprecipitated with anti-Myc antibodies and blotted with either anti-HA antibodies (top) or anti-Myc antibodies (bottom). Ponceau S staining of input is shown for reference. Marker labels refer to kDa. g, AL3 is more stable fug1 mutant background. GFP or GFP–AL3 or AL3 harbouring K178R mutation (GFP-mAL3) are in the Bur-0 background, and GFP–AL3/fug1 is in the fug1 background. Total protein from 10-day-old seedlings weighing equal amounts was extracted and immune-precipitated (IP) with anti-GFP magnetic beads and probed with anti-GFP antibody. Equal loading of protein samples is shown by Ponceau S staining of Rubisco. Marker labels refer to kDa. h, Disrupting the potential SUMOylation site in AL3 perturbs its nuclear localization. Representative maximum intensity projections showing the localization of GFP or GFP–AL3 or AL3 harbouring K178R mutation (GFP-mAL3) in Bur-0. Scale bars, 2.5 μm. i, Box and whisker plots of N:C (nuclear:cytoplasm) ratio of GFP (blue, n = 6) or GFP–AL3 (cyan, n = 16) or GFP-mAL3 (orange, n = 8) in Bur-0. Each dot represents the quantification from an individual protoplast expressing the corresponding transgene. Box plots show the 50th percentile, and the whiskers show the maximum and minimum with the mean shown by a line in the graph. Statistical comparisons were done with a two-sided Student’s t-test. *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001. Experiments shown in f–h were done at least twice with similar results.
We then asked whether this FUG1–AL3 interaction is linked with the role of FUG1 in repeat expansion-associated gene silencing. We observed AL3 among the top downregulated genes in the fug1 mutants along with an increase in IIL1 expression (Supplementary Table 3). To assess whether AL3 is required for epigenetic silencing, we generated knockdown lines (35S::amiR-AL3) in the Bur-0 background. More than 90% of the 35S::amiR-AL3 T1 transgenic lines (n = 70) showed suppression of the iil phenotype (Fig. 3c) coupled with a decrease in AL3 levels along with an increase in IIL1 expression (Fig. 3d,e and Extended Data Fig. 4a). These results confirmed the requirement of AL3 for the downregulation of IIL1 caused by gene silencing.
Members of the Alfin-like family including AL3 are known to be nuclear localized SUMOylated proteins22,24. Consistent with this, using transient assays in tobacco cells, we observed AL3 to be potentially poly-SUMOylated (Fig. 3f). SUMOylation can have multiple effects on protein function, which include promoting nuclear localization and stabilization25,26,27. To assess the impact of fug1 on AL3 at the protein level, we generated green fluorescent protein (GFP)-tagged AL3 (35S::GFP-AL3) and expressed them in Bur-0 or fug1 backgrounds. Analysis of the transgenic lines revealed that the different genotypes had roughly similar RNA expression levels of the transgene (Extended Data Fig. 4b). We found GFP–AL3 protein levels to be very low in the Bur-0 background, which is suggestive of instability of the AL3 protein, unlike GFP (Fig. 3g). However, the stability of GFP–AL3 was visibly higher in the fug1 background, which suggests that FUG1 may also be important in regulating AL3 protein levels, potentially by affecting its SUMOylation. We reasoned that perturbing the SUMOylation site in AL3 could allow us to assess its impact. SUMOylation site prediction tools GPS-SUMO 2.0 (ref. 28), JASSAv4 (ref. 29) and SUMOplot (https://www.abcepta.com/sumoplot) identified K178 to be a site of SUMOylation. However, our own in-house SUMO prediction tools failed to identify K178 as the site of SUMOylation. Nevertheless, we mutated lysine to arginine (K178R) and checked whether it would impact AL3 stability or localization using western blots (Fig. 3g) and quantitative fluorescence lifetime imaging microscopy (FLIM) (Fig. 3h,i and Extended Data Fig. 5). We found that K178R mutation also increased the protein stability but not to the same extent that we observed in thee fug1 mutants (Fig. 3g). This suggested that while K178R may be the site of SUMOylation, it is unlikely to be the sole site for SUMOylation at AL3. We also observed that K178R perturbed the nuclear localization of AL3 (Fig. 3h,i and Extended Data Fig. 5), which indicated a potential role for SUMOylation in the nuclear localization of AL3. However, we do not rule out other ways by which SUMOylation can affect AL3 function or other possible sites that might undergo SUMOylation.
AL3 is a known histone reader that binds to H3K4me3 (ref. 30), and Alfin-like family members are known to interact with components of the polycomb repressive complex to cause a chromatin state switch31. One of the chromodomain proteins is LIKE HETEROCHROMATIN 1 (LHP1), whose human orthologue HP1 is known to interact with SUMO proteases; this interaction is essential for epigenetic silencing32,33. Therefore, we considered whether FUG1 or AL3 could interact with LHP1. While we failed to detect any interaction between FUG1 and LHP1, we observed AL3–LHP1 interaction in yeast two-hybrid assays (Figs. 3a and 4a). We failed to detect an interaction between AL3 and other epigenetic regulators such as CLF and MSI1, which suggests that the AL3–LHP1 interaction was specific to LHP1 (Fig. 4a). BiFC assays in tobacco cells confirmed an in planta AL3–LHP1 interaction (Fig. 4b). To test whether LHP1 is required for epigenetic silencing caused by expanded repeats, we generated estradiol-inducible knockdown lines of LHP1 in the Bur-0 background (XVE::amiR-LHP1) and induced the amiRNAs against LHP1 at the 7th leaf stage at 27 °C under short-day conditions. Of the 40 independent transgenic lines, we observed a suppression of iil phenotype in 25 lines (Fig. 4c) coupled with an increase in IIL1 expression (Extended Data Fig. 6). These findings suggest that LHP1 is required for epigenetic silencing caused by expanded repeats.
a, AL3 interacts with LHP1 but not CLF or MSI in yeast cells. b, AL3 interacts with LHP1 in planta in BiFC assays using split-GFP in tobacco epidermal cells. PYL6 and CIS1 are used as negative controls in these assays, respectively, for C- and N-terminal fusion constructs. c, Mock-treated or estradiol-treated plants expressing amiRNAs against LHP1 (35S::amiR-LHP1) in the Bur-0 background. d, ChIP–seq profiles in an 8 kb region surrounding IIL1 for H3K4me3 or H3K27me3 marks compared with the input. Data from coverage plots of reads across the IIL1 locus (At4g13430, middle), normalized to the total number of mapped non-duplicate reads. The red asterisk in the gene model shows the location of the GAA/TTC tandem repeat. Replicates are overlayed. H3K4me3 ChIP–seq coverage is shown in green. The green rectangle shows the boundaries of the peak called at IIL1, which shows significantly increased coverage in the fug1 mutant. H3K27me3 ChIP–seq coverage is shown in orange. The red rectangle shows the gene body, which shows significantly decreased coverage in the fug1 mutant. Input for the ChIP–seq samples is shown in grey. e, Enrichment of H3K27me3 relative to agamous and H3K36me3 relative to actin in Bur-0 and fug1 at the IIL1 locus. f,g, Relative enrichment of H3K27me3 (f) and H3K4me3 (g) in 35S::amiR-AL3 and XVE::amiR-LHP1 lines compared to Bur-0. Three independent biological replicates for each sample were analysed in e–g. Statistical analysis was done with two-sided Student’s t-test. Error bars represent s.e.m. *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001.
The interaction between FUG1–AL3 and AL3–LHP1 raised the possibility that the epigenetic silencing might involve reading the H3K4me2/3 marks by AL3 followed by the recruitment of LHP1 to cause the chromatin state switch to H3K27me3 and condense the chromatin. An earlier study has shown the IIL1 locus to be a target for LHP1 (ref. 34). Therefore, we analysed the epigenetic status of the IIL1 locus in the fug1 mutant background by chromatin immunoprecipitation followed by sequencing (ChIP–seq) and by ChIP–PCR. We observed significantly high enrichment of H3K4me3 and H3K36me3 marks at the IIL1 locus in fug1 mutants compared to Bur-0 (Fig. 4d,e). The enrichment was higher at the 5′ region of the gene including the transcriptional start site. Conversely, we also observed an increase in the H3K27me3 mark in Bur-0 compared to fug1 mutants (P < 0.005; Fig. 4d,e). However, there is a diffused spread of the H3K27me3 throughout the IIL1 locus rather than a specific peak, consistent with the involvement of LHP1 and its role in spreading epigenetic silencing. These findings indicate the requirement of FUG1 in modulating the epigenetic status of IIL1 in the presence of the repeat expansion, potentially involving AL3/LHP1.
To assess whether AL3 and LHP1 can modulate the epigenetic status of the IIL1 locus, we carried out ChIP experiments in plants in which AL3 (35S::amiR-AL3) or LHP1 (XVE::amiR-LHP1) are compromised. Compromising either AL3 or LHP1 resulted in an increase in IIL1 expression (Fig. 3e and Extended Data Fig. 4a and 6). Consistent with this, we observed an increase in the H3K4me3 mark and a decrease in the H3K27me3 marks in both 35S::amiR-AL3 and XVE::amiR-LHP1 (Fig. 4f,g). These findings suggested that AL3 and LHP1 are required for the epigenetic silencing caused by expanded repeats, and their absence leads to a reversal of epigenetic signatures at the IIL1 locus.
Our earlier findings indicated that triplet repeat expansions lead to 24 nt small RNAs, which in turn recruit the RdDM components to the IIL1 locus resulting in epigenetic silencing (Fig. 5). An initial transcription is required to trigger this response, as knocking down RdDM components such as AGO4, DCL3 or POLV also abolishes the accumulation of 24 nt small RNAs16. We failed to detect small RNAs that map to the IIL1 locus in the fug1 mutant as well (Extended Data Fig. 7). Analysis of the expression levels of RdDM components failed to reveal any specific patterns. Some RdDM genes are upregulated (for example, AGO4), and some are downregulated (for example, NRPB6B (Pol V subunit)) suggesting that the fug1 effects cannot be explained by RNA expression levels of RdDM components (Extended Data Fig. 8). We, however, cannot rule out that FUG1 could target other RdDM components, as some of them are known to be SUMOylated24, although they were not recovered in our two-hybrid screen.
Expanded repeats lead to small RNAs, which through the RdDM pathway targets the IIL1 locus. This targeting involves changes in both DNA and histone methylations. When FUG1–AL3–LHP module is functional, it results in reduced IIL1 along with an increase in H3K27me3 and epigenetic silencing. When there are mutations in AL3 or FUG1 or LHP1, there is no switch to H3K27me3, and the plants retain high H3K36me3 or H3K4me3 marks coupled with an increase in IIL1 expression, resulting in a normal plant.
SUMO protease FUG1 interacts with AL3, a known histone H3K4me2/3 reader30. In the absence of FUG1 or AL3, there are higher levels of H3K4me3/H3K36me3 marks at the IIL1 locus, which suggests that removal of these marks at the IIL1 locus requires FUG1 and AL3. While FUG1 is likely to cause deSUMOylation of AL3, it is currently unclear how deSUMOylation may affect AL3 function. AL3 interacts with LHP1, and knocking down LHP1 results in an increase in H3K4me3 marks at IIL1 coupled with an increase in its expression (Fig. 5). In the presence of all these proteins, there is an increase in the repressive H3K27me3 mark coupled with a decrease in IIL1 expression, which suggests that there is a switch from active to inactive chromatin state at high temperatures. It is currently unclear whether the requirement of FUG1, AL3 and LHP1 for the epigenetic changes is a direct or indirect effect of these proteins at the IIL1 locus. While the mechanism is yet to be explored, we have demonstrated that SUMO protease FUG1, histone reader AL3 and the chromodomain protein LHP1 are required for epigenetic silencing caused by expanded repeats (Fig. 5).
In conclusion, we have demonstrated a key role for post-translational modifiers and histone readers in conferring epigenetic gene silencing in plants. In recent studies, there is an appreciation of the bivalent chromatin state in which both active (H3K4me3) and repressive (H3K27me3) marks are present, which play diverse roles in plant development and stress response35. Although the mechanisms remain unclear, our work suggests potential direct or indirect involvement of SUMO proteases, Alfin-like genes and LHP1 in chromatin switch changes. Given the importance of SUMOylation/deSUMOylation dynamics in stress response, it would be interesting to assess whether similar chromatin remodelling mechanisms operate in the context of plant defence or abiotic stress responses or transposon silencing.
Our studies draw parallels with the human system where sentrin-like protease-mediated deSUMOylation of HP1 has been shown to be critical in its enrichment at pericentric chromatin33. Interestingly, in this case, while we could not detect a direct interaction between the SUMO protease FUG1 and LHP1, a PHD finger protein AL3 acts as a conduit orchestrating a change in the chromatin. While studies on differing species reveal some species-specific aspects, our study once again suggests that core principles of gene regulation may be conserved across eukaryotic systems. It will be interesting to see whether post-translational modifiers and histone readers have any role in the epigenetic silencing caused by expanded intronic repeats in Friedreich’s ataxia and other human diseases.
Methods
Plant material and growth conditions
A. thaliana accessions Bur-0 (and Pf-0 have been previously described7. Plants were grown at 23 °C and 27 °C under short-day (8 h light, 16 h darkness) conditions for DNA sequencing, CHIP–seq and small-RNA profiling. For the genetic suppressor screens, approximately 32,000 Bur-0 seeds were mutagenized with EMS as described previously7. Approximately we pooled 10 M1 plants per family and the plants were grown at the Monash greenhouse in long-day conditions for seed collection. In total we screened 120 M2 families at 27 °C under short-day conditions to identify potential suppressors of the iil phenotype.
Yeast two-hybrid screening library
Yeast two-hybrid screening was done as described previously36. The coding region (CDS) of FUG1 was cloned into the pBridge vector (Clonetech), and FUG1-pBridge was used as the bait for yeast two-hybrid screening. FUG1-pBridge and the plasmid library containing pACT2 (from J. Ecker, Salk Institute, California) were co-transformed into Y190 yeast strain for yeast two-hybrid screening. Diluted samples of the Y190 yeast strain were spread on appropriate synthetic defined (SD) selection agar plates (SD-Trp/-His/-Leu containing 30 mM 3-amino-1,2,4-triazole) and grown at 28 °C for 6–8 days. The mono-yeast clones with large volume were selected for strewing and X-Gal chromogenic reaction. The clones with blue X-Gal chromogenic reaction were used for yeast PCR, and the products of PCR were sequenced by Sanger sequencing; NCBI Blastn was used to identify the target genes.
To test the interaction with FUG1, the CDS of AL3 was cloned into the pGADT7 vector (Clonetech). We then cloned AL3 into the pBridge vector and tested interactions with the CDS of LHP1, CLF and MSI1 cloned into the pGADT7 vector. The different combinations were co-transformed into yeast AH109 or Y190 and plated on SD-Leu/-Trp medium for growth. Then the AH109 strains were transferred to SD-/-Trp/-His/-Leu/-Ala plate, and the Y190 strains were subjected to X-Gal chromogenic reaction to test interaction.
BiFC assay
BiFC assays were done according to Walter et al.37 as described previously36. The CDS of AL3 was cloned into the pUC-SPYNE (nYFP-X) vector. FUG1 and LHP1 were cloned into the pUC-SPYCE (cYFP-X) vector. Each vector was transformed into Agrobacterium GV3101, and isolated single clones were incubated overnight in Luria–Bertani medium at 28 °C. MES mixing buffer (10 mM MgCl2, 10 mM MES, 100 µM acetobutylone) was used to dilute the culture to a suitable OD600 (0.8–1.0). The different combinations of GV3101 containing N-terminal yellow fluorescent protein (nYFP) and C-terminal cyan fluorescent protein (cCFP) fusion constructs were mixed, respectively, and infected the abaxial side of 3-week-old tobacco leaves, and the fluorescence signals were observed by confocal microscopy 3 days later using an Olympus Fluorescence Microscope (Olympus CellSens Standard: excitation wavelength, 488 nm; emission wavelength, 505 nm).
Subcellular localization analysis by FLIM
Protoplasts and homozygous transgenic plants containing either enhanced green fluorescent protein (EGFP), EGFP–AL3, EGFP-FUG1 or EGFP–AL3K178R were imaged using an SP8 Falcon (Leica Microsystems) with an 86 × 1.2 numerical aperture objective. Protoplasts were extracted using the whole leaf tape sandwich method as per Wu et al.38, and transfections were carried out according to Yoo et al.39. The protoplasts and leaves were imaged as volumes with slices taken at 0.5 μm and 2 μm, respectively, using a 1.0 AU pinhole. Images were acquired with 488 nm excitation at 10% transmission using a tuneable pulsed white-light laser. Emissions were collected from 500 nm to 560 nm using a hybrid detector (HyD). The GFP and auto-fluorescence were separated using pixel-wise two-component fitting with τ1 = 2.600 ns and τ1 ≈ 0.2 ns. These images were analysed as volumes, and the representative images were displayed as maximum-intensity projections.
SHOREmap analysis and sequencing of mutants
For bulk segregant analysis, we pooled leaf tissues from more than 500 plants that showed the suppressor phenotype (normal looking) from the segregating F2 population derived from a backcross with the Bur-0 parent. Other mutants were directly sequenced. Genomic DNA was extracted by using DNeasy Plant Maxi kit (catalogue number 68163) from Qiagen. All samples that passed quality control were processed for sequencing on the Illumina Novaseq PE 150 platform by GENEWIZ-China. Approximately 30 million reads per sample were analysed.
Quality-controlled sequences were aligned to the TAIR10 reference genome using Bowtie2 (v2.4.4)40. SNPs were called using samtools (v1.13; mpileup -E -uf), and VCF files were generated with bcftools (v1.13; call -c). We then ran SHOREmap (v3.6) to identify SNPs most enriched in suppressor-phenotype F2 plants (foreground) compared to the Bur-0 parent (background). We found variant SNPs in suppressor-phenotype F2 VCFs using SHOREmap (v3.6; convert command). We were unable to run the SHOREmap extract command, so we developed a python script (filter_subtract_variants_vcf.py) to calculate the allele frequencies as follows: Allele frequency = Frequency of SNP in foreground − Frequency of SNP in background. We only considered SNPs if they were EMS-type C to T mutations or G to A mutations. SNPs were considered putative losses of function if they generated a stop codon or altered the core sequence (GT or AG) of an annotated splice site, or otherwise if one of the most enriched SNPs was predicted to cause a change in amino acid sequence.
Given that our first four suppressors mapped to genes in the RdDM pathway, we adapted our pipeline to identify candidate genes without requiring a backcross. For the suppressor mutants themselves, DNA was extracted, sequenced, aligned and processed to VCF files as above. We then ran SHOREmap (v3.6) to identify SNPs most enriched in the suppressors themselves (foreground) compared to the Bur-0 parent (background). This meant that all homozygous EMS SNPs unique to the suppressor were assigned an allele frequency of 1.0. We then took only those SNPs putatively causing a loss of function. We then asked whether any of the dozen or so remaining SNPs belonged to known RdDM pathway genes.
Small RNA sequencing
Small RNA sequencing has been previously described16. Leaf tissue from 45-day-old fug1 mutant plants were collected 5 h after the beginning of the day (light exposure) and snap frozen immediately using liquid nitrogen. Total RNA was extracted using TRIzol reagent (Ambion). Quality-controlled reads were mapped to the TAIR10 Arabidopsis genome using SCRAM41 and analysed as described previously16.
ChIP–PCR and ChIP–seq
For ChIP–seq, Bur-0 and fug1 mutant plants were grown for 5 weeks at 27 °C under short-day conditions. Leaf tissues (1.5 g) from two independent biological replicates were used for chromatin preparation. Further chromatin preparation steps were followed as previously described42. We assessed histone marks H3K4me3, H3K27me3 and H3K36me3 and the ChIP-grade antibodies against these marks procured from Diagenode (H3K4me3, C15410003-50; H3K27me3, C15410195) or Abcam (H3K36me3, Ab9050). Antibody (5 μg) was used for each ChIP reaction. ChIP–PCR analysis was done for the IIL1 region or for GFP with primers described in Supplementary Table 3. We used AGAMOUS as a positive control for H3K27me3 and ACTIN as a positive control for H3K4me3 and H3K36me3 and normalized the samples using these positive controls, similar to a normalization in quantitative reverse transcription PCR (RT-PCR) where housekeeping genes are used as controls. This process allows to control for variation within biological replicates. Subsequently, the samples were further normalized to the Bur-0 control allowing direct comparison of samples.
For ChIP–seq, purified DNA quality was checked by Bioanalyzer, and samples that passed quality control were subjected to library preparation and processed for sequencing using Illumina Novaseq PE150 sequencing platform. Roughly 20 million reads (6 gigabases) per sample raw data were used for analysis. Clean reads were aligned to the TAIR10 genome using Bowtie2 (ref. 40) with default parameters. For H3K4me3 samples, peaks were identified using MACS3 (ref. 43) v3.0.0a7; parameters: -f BAMPE, –broad, –broad-cutoff 0.1, -g 1.35e8). We took all peaks observed across all samples; where two peaks in different replicates overlapped, we considered them to be the same peak and took the maximal boundaries. For H3K27me3 samples, we simply measured within TAIR10 annotated gene boundaries. Reads across regions (either peaks or genes) were counted using featureCounts from the Subread package v2.0.1 (ref. 44). Differentially covered regions were identified using DESeq2 (ref. 45).
Transient in vivo SUMOylation assay of AL3
To analyse the SUMOylation of AL3, PCR-amplified full-length complementary DNA (cDNA) of AL3 was cloned into pGWB621. The hemagglutinin (HA)-tagged SUMO (HA-SUMO) construct has been previously described46. Agrobacterium harbouring AL3-myc and HA-SUMO1 were diluted in 10 mM MgCl2 supplemented with 150 µM acetosyringone with a final OD600 of 0.2 and co-infiltrated in 4-week-old tobacco leaves as described previously47. After 3 days post infiltration, approximately 2 g of the leaf samples was collected and frozen in liquid nitrogen and stored at −80°C until use. Total protein was extracted in SUMO extraction buffer (50 mM Tris pH 8.5, 150 mM NaCl, 1 mM EDTA, 0.1% SDS, 1% NP-40, 20 mM N-ethylmaleimide, 0.5% sodium deoxycholate and 1× protease inhibitor cocktail) in a cold room. The supernatant was incubated with anti-Myc magnetic beads for 30 min in a rotary mixer in a cold room. The bound protein was washed twice with extraction buffer and eluted in 1× sample-loading buffer pre-heated at 95 °C. Eluted protein was further analysed in 10% SDS–PAGE gel and further transferred to polyvinylidene difluoride membrane. After blocking with 5% skimmed milk, membrane was further incubated with anti-HA (Abcam)/anti-SUMO1 (in-house produced) and anti-Myc (Sigma) overnight (all 1:1,000 dilutions). After incubating the membrane with secondary antibodies for 1 h, the membrane was vigorously washed, and the X-ray films were developed by automated developer machine.
In vitro SUMOylation assay
In vitro deSUMOylation assays were performed as described previously48. SUMO protease substrate His-SUMO-FLC1 and FUG1-HIS SUMO protease were expressed in bacteria and purified. His-SUMO1-FLC1 was incubated alone or with increasing concentration of FUG1-HIS protease in SUMO protease buffer (50 mM Tris–HCl, pH 8.0, 0.2% Igepal, 1 mM DTT) at 30°C for 4 h. Reaction was stopped by adding 1× SDS loading buffer and heated at 98 °C for 5 min. Protein samples were separated on 12% SDS–PAGE gel. HIS-SUMO1-FLC1 and free SUMO were detected using anti-SUMO1 antibody (1:1,000 dilution).
Western blots
Total protein was extracted as per standard protocols49 and used for western blots. Briefly, around 1 gm of plant tissue was ground into fine powder and mixed with 1 ml of extraction buffer (50 mM Tris acetate [pH 7.9], 100 mM potassium acetate, 1 mM EDTA, 1 mM DDT, 20% glycerol and protease inhibitors) and centrifuged at 13,000 r.p.m. at 4 °C for 10 min to separate the soluble total protein. Protein concentration was determined using Bradford assay (Bio-Rad), and 100 ng of total proteins was loaded per sample on the SDS gel (Bio-Rad). Proteins were transferred to a polyvinylidene difluoride membrane (Thermo Scientific), blocked with 5% skimmed milk powder at room temperature and probed with the anti-sumo1 (Abcam, ab5316; 1:1,000) or anti-actin (Abcam, ab197345; 1:1,000) antibody overnight at 4 °C. After washing, the membrane was probed with a horseradish peroxidase (HRP)-conjugated secondary antibody and incubated further for 1 h at room temperature. The membrane was washed and visualized by chemiluminescence using an enhanced chemiluminescense reagent (Thermo Scientific) under an automated developer.
Immunoprecipitation experiments
Approximately 1 g of 10-day-old seedlings of 35S::GFP-AL3 in the Bur-0 and fug1 backgrounds, and 35S::GFP-mAL3 and 35S::GFP in Bur-0 plants were collected for protein extractions. The immunoprecipitation method was done as described previously50 with slight modifications. The exact amounts of proteins from each genotype were loaded onto a 4% to 12% standard SDS–PAGE gel (Bio-Rad). Proteins were transferred to a polyvinyl difluoride membrane (Bio-Rad) for immunoblot analysis. The membrane was blocked in Tween 20 and Tris-buffered saline (TTBS)–milk (5% (w/v) dry non-fat milk, 10 mM Tris–HCl, pH 8, 150 mM NaCl, and 0.1% (v/v) Tween 20) before incubation with anti-GFP antibody (Abcam), diluted to 1:1,000. Furthermore, the membrane was washed in TTBS and incubated with secondary antibody (anti-rabbit HRP conjugate (Sigma Aldrich)) diluted to 1:10,000 in TTBS–milk. Finally, the membrane was washed and incubated with the HRP substrate (Immobilon Western; Millipore) and taken for image analysis (ChemiDoc imaging systems, Bio-Rad).
Artificial microRNAs
Online tool WMD3 (http://wmd3.weigelworld.org/cgi-bin/webapp.cgi) was used to design amiRNAs against FUG1, Al3 and LHP1. FUG1 and LHP1 amiRNAs were generated by site-directed mutagenesis as per Schwab et al.51; the LHP1 amiRNA construction details have been explained previously16. AL3 amiRNA constructs were commercially synthesized in plasmids (IDT) and then sub-cloned into Gateway-compatible entry vector (pDONR221). The final amiRNA precursors were sub-cloned into the plasmid pFK210 by conducting LR reactions according to the manufacturer’s protocol (Gateway LR Clonase II Enzyme mix, Invitrogen). The LHP1 amiRNA was cloned into the Gateway-compatible pGREAT XVE-system vector16. Sequence-verified constructs were transformed into Agrobacterium tumefaciens GV3103 further transformed into Col-0, Bur-0 and fug1 genotypes using the floral dip method52. The first generation (T1) of transformants were grown at 27 °C under short-day conditions and continuously watered with nutrient water containing 120 mg l−1 BASTA (glufosinate ammonium, Bayer) for selection. For the inducible system, plants were sprayed with 50 µM estradiol with 0.1% Silwet L-77 every 2 days after the emergence of the 7th leaf, as previously described16. We obtained multiple independent primary transgenic lines, and most of the transgenic lines showed phenotypic suppression. The number of primary independent transformants that showed complete phenotypic suppression for each of the transgenes is: 35S::amiR-FUG1 –190, 35S::amiR-AL3 –68, 35S::XVE::amiR-LHP1 –25. The presence of each transgene was confirmed by PCR using the primers listed in Supplementary Table 4, and the expression level of each amiRNA’s target gene was determined by quantitative RT-PCR (qRT-PCR). Expanded GAA/TTC repeat tracts at the IIL1 locus were confirmed by PCR (primer pair: oSKB_608 and oSKB_561; Supplementary Table 4). After phenotypic analysis at 27 °C under short-day conditions, primary transformants were transferred to 23 °C long-day conditions for seed collection.
RNA-seq and gene expression analysis
RNA-seq analysis was carried out as described previously42. Five-week-old Bur-0 and fug-1 plants grown at 27 °C were collected and flash frozen for RNA extractions. Three biological replicates for each genotype was used. Total RNA was quality controlled, and paired-end sequencing was carried out by GENEWIZ (Azenta USA). Sequences were processed and analysed by GENEWIZ, and the differentially expressed genes were identified through DESeq2 (ref. 45). For qRT-PCR, total RNA was extracted using TRIzol reagent (Ambion). cDNA was synthesized with Anchored-oligo (dT)18 primers and the Transcriptor First Strand cDNA Synthesis Kit (Roche). cDNA was diluted fivefold with water, and 2–4 μl of the diluted cDNA served as the template for each qRT-PCR as described previously16. Variation in gene expression was analysed through qRT-PCR analysis using the 2−ΔΔcT method53. The statistical significance of the difference in gene expression between specific samples was analysed through Student’s t-test or analysis of variance. For ChIP experiments, the data are expressed after normalizing with a positive control, and the statistical significance was analysed through a Student’s t-test.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All sequencing data are available at the National Center for Biotechnology Information short reads archive. The ChIP–seq reads are available in Bioproject ID PRJNA1080228, the small RNA sequence from fug1 is in Bioproject ID PRJNA1080246 and the RNA-seq data are in Bioproject ID PRJNA1080085. Source data are provided with this paper.
Code availability
The python script that we used to extract allele frequencies is available upon request to the corresponding authors.
References
Zhang, G., Wang, Y. & Andersen, E. C. Natural variation in C. elegans short tandem repeats. Genome Res. 32, 1852–1861 (2022).
Sawyer, L. A. et al. Natural variation in a Drosophila clock gene and temperature compensation. Science 278, 2117–2120 (1997).
Garg, P. et al. A phenome-wide association study identifies effects of copy-number variation of VNTRs and multicopy genes on multiple human traits. Am. J. Hum. Genet. 109, 1065–1076 (2022).
Barre, B. P. et al. Intragenic repeat expansion in the cell wall protein gene HPF1 controls yeast chronological aging. Genome Res. 30, 697–710 (2020).
Han, Z. et al. Natural variation underlies differences in ETHYLENE RESPONSE FACTOR17 activity in fruit peel degreening. Plant Physiol. 176, 2292–2304 (2018).
Press, M. O., McCoy, R. C., Hall, A. N., Akey, J. M. & Queitsch, C. Massive variation of short tandem repeats with functional consequences across strains of Arabidopsis thaliana. Genome Res. 28, 1169–1178 (2018).
Sureshkumar, S. et al. A genetic defect caused by a triplet repeat expansion in Arabidopsis thaliana. Science 323, 1060–1063 (2009).
Depienne, C. & Mandel, J. L. 30 years of repeat expansion disorders: what have we learned and what are the remaining challenges? Am. J. Hum. Genet. 108, 764–785 (2021).
Gall-Duncan, T., Sato, N., Yuen, R. K. C. & Pearson, C. E. Advancing genomic technologies and clinical awareness accelerates discovery of disease-associated tandem repeat sequences. Genome Res. 32, 1–27 (2022).
Rafehi, H. et al. An intronic GAA repeat expansion in FGF14 causes the autosomal-dominant adult-onset ataxia SCA50/ATX-FGF14. Am. J. Hum. Genet. 110, 105–119 (2023).
Pellerin, D. et al. Deep intronic FGF14 GAA repeat expansion in late-onset cerebellar ataxia. N. Engl. J. Med. 388, 128–141 (2023).
Erwin, G. S. et al. Recurrent repeat expansions in human cancer genomes. Nature https://doi.org/10.1038/s41586-022-05515-1 (2022).
Malik, I., Kelley, C. P., Wang, E. T. & Todd, P. K. Molecular mechanisms underlying nucleotide repeat expansion disorders. Nat. Rev. Mol. Cell Biol. 22, 589–607 (2021).
Finsterer, J. Phenotype and management of neurologic intronic repeat disorders (NIRDs). Rev. Neurol. https://doi.org/10.1016/j.neurol.2022.09.004 (2022).
Gottesfeld, J. M. Molecular mechanisms and therapeutics for the GAA.TTC expansion disease Friedreich ataxia. Neurotherapeutics 16, 1032–1049 (2019).
Eimer, H. et al. RNA-dependent epigenetic silencing directs transcriptional downregulation caused by intronic repeat expansions. Cell 174, 1095–1105 e1011 (2018).
Schneeberger, K. et al. SHOREmap: simultaneous mapping and mutation identification by deep sequencing. Nat. Methods 6, 550–551 (2009).
Castro, P. H. et al. Arabidopsis thaliana SPF1 and SPF2 are nuclear-located ULP2-like SUMO proteases that act downstream of SIZ1 in plant development. J. Exp. Bot. 69, 4633–4649 (2018).
Morrell, R. & Sadanandom, A. Dealing with stress: a review of plant SUMO proteases. Front. Plant Sci. 10, 1122 (2019).
Rosa, M. T. G. et al. Insights into the transcriptional and post-transcriptional regulation of the rice SUMOylation machinery and into the role of two rice SUMO proteases. BMC Plant Biol. 18, 349 (2018).
Wei, W. et al. The Alfin-like homeodomain finger protein AL5 suppresses multiple negative factors to confer abiotic stress tolerance in Arabidopsis. Plant J. 81, 871–883 (2015).
Lee, W. Y., Lee, D., Chung, W. I. & Kwon, C. S. Arabidopsis ING and Alfin1-like protein families localize to the nucleus and bind to H3K4me3/2 via plant homeodomain fingers. Plant J 58, 511–524 (2009).
Wysocka, J. et al. A PHD finger of NURF couples histone H3 lysine 4 trimethylation with chromatin remodelling. Nature 442, 86–90 (2006).
Miller, M. J., Barrett-Wilt, G. A., Hua, Z. & Vierstra, R. D. Proteomic analyses identify a diverse array of nuclear processes affected by small ubiquitin-like modifier conjugation in Arabidopsis. Proc. Natl Acad. Sci. USA 107, 16512–16517 (2010).
Augustine, R. C. & Vierstra, R. D. SUMOylation: re-wiring the plant nucleus during stress and development. Curr. Opin. Plant Biol. 45, 143–154 (2018).
Wen, D., Wu, J., Wang, L. & Fu, Z. SUMOylation promotes nuclear import and stabilization of polo-like kinase 1 to support its mitotic function. Cell Rep. 21, 2147–2159 (2017).
Park, I. S. et al. SUMOylation regulates nuclear localization and stability of TRAIP/RNF206. Biochem. Biophys. Res. Commun. 470, 881–887 (2016).
Zhao, Q. et al. GPS-SUMO: a tool for the prediction of sumoylation sites and SUMO-interaction motifs. Nucleic Acids Res. 42, W325–W330 (2014).
Beauclair, G., Bridier-Nahmias, A., Zagury, J. F., Saib, A. & Zamborlini, A. JASSA: a comprehensive tool for prediction of SUMOylation sites and SIMs. Bioinformatics 31, 3483–3491 (2015).
Zhao, S., Zhang, B., Yang, M., Zhu, J. & Li, H. Systematic profiling of histone readers in Arabidopsis thaliana. Cell Rep. 22, 1090–1102 (2018).
Molitor, A. M., Bu, Z., Yu, Y. & Shen, W. H. Arabidopsis AL PHD-PRC1 complexes promote seed germination through H3K4me3-to-H3K27me3 chromatin state switch in repression of seed developmental genes. PLoS Genet. 10, e1004091 (2014).
Romeo, K. et al. The SENP7 sumo-protease presents a module of two HP1 interaction motifs that locks HP1 protein at pericentric heterochromatin. Cell Rep. https://doi.org/10.1016/j.celrep.2015.01.004 (2015).
Maison, C. et al. The SUMO protease SENP7 is a critical component to ensure HP1 enrichment at pericentric heterochromatin. Nat. Struct. Mol. Biol. 19, 458–460 (2012).
Veluchamy, A. et al. LHP1 regulates H3K27me3 spreading and shapes the three-dimensional conformation of the Arabidopsis genome. PLoS ONE 11, e0158936 (2016).
Faivre, L. & Schubert, D. Facilitating transcriptional transitions: an overview of chromatin bivalency in plants. J. Exp. Bot. 74, 1770–1783 (2023).
Zhao, Z. et al. CRY2 interacts with CIS1 to regulate thermosensory flowering via FLM alternative splicing. Nat. Commun. 13, 7045 (2022).
Walter, M. et al. Visualization of protein interactions in living plant cells using bimolecular fluorescence complementation. Plant J. 40, 428–438 (2004).
Wu, F. H. et al. Tape–Arabidopsis sandwich—a simpler Arabidopsis protoplast isolation method. Plant Methods 5, 16 (2009).
Yoo, S. D., Cho, Y. H. & Sheen, J. Arabidopsis mesophyll protoplasts: a versatile cell system for transient gene expression analysis. Nat. Protoc. 2, 1565–1572 (2007).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Fletcher, S. J., Boden, M., Mitter, N. & Carroll, B. J. SCRAM: a pipeline for fast index-free small RNA read alignment and visualization. Bioinformatics https://doi.org/10.1093/bioinformatics/bty161 (2018).
Tasset, C. et al. POWERDRESS-mediated histone deacetylation is essential for thermomorphogenesis in Arabidopsis thaliana. PLoS Genet. 14, e1007280 (2018).
Zhang, Y. et al. Model-based analysis of ChIP-seq (MACS). Genome Biol. 9, R137, https://doi.org/10.1186/gb-2008-9-9-r137 (2008).
Liao, Y., Smyth, G. K. & Shi, W. The Subread aligner: fast, accurate and scalable read mapping by seed-and-vote. Nucleic Acids Res. 41, e108 (2013).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Conti, L., Kioumourtzoglou, D., O’Donnell, E., Dominy, P. & Sadanandom, A. OTS1 and OTS2 SUMO proteases link plant development and survival under salt stress. Plant Signal Behav. 4, 225–227 (2009).
Ewan, R. et al. Deubiquitinating enzymes AtUBP12 and AtUBP13 and their tobacco homologue NtUBP12 are negative regulators of plant immunity. New Phytol. 191, 92–106 (2011).
Bhagat, P. K., Roy, D. & Sadanandom, A. Expression, purification, and enzymatic analysis of plant SUMO proteases. Methods Mol. Biol. 2581, 109–119 (2023).
Weigel, D. & Glazebrook, J. Arabidopsis: A Laboratory Manual (Cold Spring Harbor Laboratory Press, 2002).
Orosa, B. et al. SUMO conjugation to the pattern recognition receptor FLS2 triggers intracellular signalling in plant innate immunity. Nat. Commun. 9, 5185 (2018).
Schwab, R., Ossowski, S., Riester, M., Warthmann, N. & Weigel, D. Highly specific gene silencing by artificial microRNAs in Arabidopsis. Plant Cell 18, 1121–1133 (2006).
Clough, S. J. & Bent, A. F. Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J. 16, 735–743 (1998).
Pfaffl, M. W. A new mathematical model for relative quantification in real-time RT-PCR. Nucleic Acids Res. 29, e45 (2001).
Acknowledgements
We thank R. Clark (Utah, USA) for help with sequencing the fug1 mutant, A. Seleznev for help with bioinformatic analysis and H. Sako for initial characterization of suppressors. We thank J. Bowman, J. Coutts, Y. Guo, D. Smyth and L. Wong for discussions and comments on this manuscript. This work is supported by a Canadian Institute for Health Research grant PJT-178112 (E.R.), Department of Science and Technology – Science and Engineering Research Board Core grant (R.K.Y.), a National Health and Medical Research Council project grant APP1182090 (S.B. and S.S.), Australian Research Council Discovery Projects DP1095325 (S.B.) and DP190101818 (S.B.), and Australian Research Council Future Fellowships FT100100377 (S.B.), FT190100403 (S.S.), Australia–India Strategic Research Fund–Early and Mid-Career Fellowship (S.S.) and a Monash Larkins Fellowship (S.B.).
Author information
Authors and Affiliations
Contributions
Conceptualization: S.S. and S.B.; methodology: S.S., H.L., S.A., A.S., E.R. and S.B.; software: C.I.D., S.S. and S.B.; formal analysis: S.S., C.B., J.L., C.I.D., P.K.B., S.M., R.S., C.A., H.M.Y., N.S., G.F., P.T., A.S.Y., B.G.B., S.K., P.S., O.S.G. and S.B.; investigation: S.S., J.L., S.M., R.S., C.A., H.M.Y., N.S. and C.B.; writing—original draft: S.S.; writing—review and editing: S.S., A.S., S.A., H.L. and S.B.; visualization: S.S., S.A., H.M.Y. and S.B.; supervision: S.S., R.K.Y., S.A., A.S., H.L. and S.B.; project administration: S.S. and S.B.; funding acquisition: S.S. and S.B.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Plants thanks Jixian Zhai and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Genetic mapping of the suppressors of Bur-0 by SHOREMAP.
Allele frequency of EMS-type SNPs in the fraction of F2 pooled plants that display the suppression of the iil phenotype. a) 49-9 b) 61-7 and c) 57-3. High frequency alleles (>0.85) are coloured red and red crosses show the putative causal alleles.
Extended Data Fig. 2 fug1 mutant suppresses the temperature-dependent iil phenotype despite the presence of repeat expansion.
a) fug1 phenotype compared with Bur-0 at 23 °C and 27 °C. b) Suppression of the iil phenotype in fug1 is not due to the loss of a repeat expansion. Gels showing retention of the expanded repeat in fug1 plants compared with Bur-0 plants. The red boxes display the typical banding pattern of the repeat expansion. 1.5Kb band on the 1Kb plus (Invitrogen) is shown as the molecular marker. Each lane represents DNA analysed from individual plants with the Col-0 control in the first lane after the marker showing a non-expanded repeat. This is analysed multiple times with similar results.
Extended Data Fig. 3 FUG1 is localised to the nucleus.
Representative maximum intensity projections (MIPs) of protoplasts of transgenic Bur-0 leaves harbouring either GFP (left) or GFP-FUG1 (right). Autofluorescence (purple) was separated using fluorescence life time. Scale bar=10 µM. The experiment is replicated more than twice with similar results.
Extended Data Fig. 4 Analysis of AL3 expression in transgenic lines.
a) Knocking down AL3 leads to increase in IIL1 expression. Independent plants representing multiple T2 lines (shown in X-axis) were analysed for IIL1 expression. Data were analysed through ANOVA and the statistical significance was tested with Tukey’s posthoc test. Different labels indicate significant differences (p < 0.05) in gene expression n = 5 in Bur-0, n = 4 in T2 Line 28 and n = 3 in all other T2 lines. b) Comparison of the expression levels of various fusion constructs of GFP-AL3. Primers against the GFP region allowed the measurement of transgenic AL3 levels in different backgrounds. Anlysis by ANOVA revealed that there were no significant differences in gene expression n = 3 in all samples except 35 S::GFP-AL3, where n = 6. Error bars represent standard errors of mean.
Extended Data Fig. 5 Perturbing the SUMOylation site of AL3 affects its sub-cellular localisation.
a) A schematic of the Fluorescence Lifetime Imaging (FLIM) assay using transformed protoplasts. Scale bar = 5 μm b) Representative maximum intensity projections showing the localisation of GFP or GFP-AL3 or AL3 harbouring K178R mutation (GFP-AL3-K178R) in Bur-0 and fug1 mutant backgrounds 2.5 μm. c) Box and whisker plots of nuclear:cytoplasm ratio of GFP (blue, n = 6 in both Bur-0 and fug1) or GFP-AL3 (cyan, n = 16 in Bur-0 and n = 24 in fug1) or GFP-AL3-K178R (orange, n = 8 in Bur-0 and n = 14 in fug1) in Bur-0 and fug1 mutant backgrounds. Box plots show the 50th percentile and the whiskers show the max and minimum with the mean shown by a line in the graph. Each dot represents the quantification from an individual protoplast expressing the corresponding transgene. Statistical comparisons were done with a two sided Student’s t-test. P-values: **<0.01, ***<0.001, ****<0.0001. The first lane of panel b and the left half (Bur) of panel c are also used in the Main Fig. 3(h & i). The protoplast experiments were carried out at least twice with similar results.
Extended Data Fig. 6 XVE::amiR-LHP1 lines show an increase in IIL1 levels upon estradiol induction.
a) Relative IIL1 expression levels. Comparison is made between Bur-0, empty vector controls as well as mock treated plants with that of estradiol treated samples in one-way ANOVA with the samples being separated as controls vs test. b) Comparison of IIL1 expression in plants that show the suppression of the iil phenotype and those that do not analysed together independent of the specific transgenic lines in one-way ANOVA c) Comparison of IIL1 expression in plants that show the suppression of the iil phenotype and those that do not from independent T2 lines. Expression analysis is from 2–3 independent plants as shown by individual data points on each of the bar graphs. The percentage of plants that show the suppression and the number of plants analysed are shown above each of the transgenic lines. The comparison of gene expression in this dataset is by two-sided Student’s T-test. Error bars represent standard errors of mean. p-values: *<0.05, **<0.01, ***<0.001, ****<0.0001.
Extended Data Fig. 7 Abundance of siRNAs that map to IIL1 locus in fug1 mutant.
Small RNAs that are typically found in Bur-0 at 27 °C (Fig. 2 of Eimer et al, Cell, 20184 were not observed in the fug 1 mutant. Small RNA profiles are generated as previously described4 using Small Complementary RnA Mapper (SCRAM)37. The genic region is shown in the bottom with the black boxes and lines representing exons and introns, respectively. Normalised coverage along with standard error shown as shadows is shown for different types of small RNAs as indicated by the colour code. Only small RNAs that mapped to the non-triplet repeat sequences of IIL1 are shown. Quantification of the sense and antisense small RNAs are shown in the positive and negative dimensions, respectively, along the y-axis. The green arrow indicates the position of the GAA/TTC repeat in the intron 3 of IIL1.
Extended Data Fig. 8 Effects of fug1 on gene silencing cannot be explained by changes in the RNA expression levels of the RdDM components.
A volcano plot depicting the changes in the RNA expression levels of genes in the RdDM pathway. Genes that show significant differential expression are marked in red. Differential expression was assessed by DESeq2 and only samples with an adjusted p-value after multiple correction (p-adj) are shown. An example for upregulated (AGO4) and downregulated (AGO9) are highlighted in the figure.
Supplementary information
Supplementary Information
Supplementary Tables 1–4.
Source data
Source Data Fig. 2
Unprocessed actin blot of the lower panel of Fig. 2c.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Sureshkumar, S., Bandaranayake, C., Lv, J. et al. SUMO protease FUG1, histone reader AL3 and chromodomain protein LHP1 are integral to repeat expansion-induced gene silencing in Arabidopsis thaliana. Nat. Plants (2024). https://doi.org/10.1038/s41477-024-01672-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41477-024-01672-5