Abstract
Prevention of autonomous division of the egg apparatus and central cell in a female gametophyte before fertilization ensures successful reproduction in flowering plants. Here we show that rice ovules of Polycomb repressive complex 2 (PRC2) Osfie1 and Osfie2 double mutants exhibit asexual embryo and autonomous endosperm formation at a high frequency, while ovules of single Osfie2 mutants display asexual pre-embryo-like structures at a lower frequency without fertilization. Earlier onset, higher penetrance and better development of asexual embryos in the double mutants compared with those in Osfie2 suggest that the autonomous endosperm facilitated asexual embryo development. Transcriptomic analysis showed that male genome-expressed OsBBM1 and OsWOX8/9 were activated in the asexual embryos. Similarly, the maternal alleles of the paternally expressed imprinted genes were activated in the autonomous endosperm, suggesting that the egg apparatus and central cell convergently adopt PRC2 to maintain the non-dividing state before fertilization, possibly through silencing of the maternal alleles of male genome-expressed genes.
Similar content being viewed by others
Main
In flowering plants, seed propagation requires double fertilization, in which the haploid egg cell and the homodiploid central cell in a female gametophyte (embryo sac) are fertilized with two genetically identical sperms to form a diploid embryo and triploid endosperm1,2. The fertilized egg or zygote starts embryogenesis following a serial division stage to form a basic body plan for a stereotyped seedling. Genes with de novo expression in the fertilized egg (zygote) are important for embryogenesis2,3,4. Among these genes with known embryogenic functions in rice are transcription factors, such as OsBBM1, OsBBM2, OsWOX8/9 and OsWOX2 (refs. 4,5,6,7). OsBBM1, OsBBM2 and OsWOX8/9 are specifically expressed from the paternal alleles after fertilization and ectopic expression of OsBBM1 in the egg can trigger embryogenesis3,4, suggesting that the imprinted state of the maternal allele is important for egg quiescence. Little is known about the mechanism that maintains this quiescent state before fertilization.
The fertilized central cell nucleus starts to divide without cytokinesis to form a multinucleate cell called syncytial endosperm, followed by cellularization. A body of knowledge that explains how the central cell is prevented from autonomous division has been built up in Arabidopsis and rice8,9,10,11,12,13,14,15. Mutations in members of the Polycomb repressive complex 2 (PRC2) genes cause autonomous endosperm development potentially by activating the auxin biosynthesis pathway in the central cell in the absence of fertilization16,17. PRC2, an important epigenetic regulator, catalyses histone H3 lysine27 trimethylation (H3K27me3)17,18, which regulates imprinting in endosperm by preferentially targeting maternal alleles of paternally expressed imprinted genes (PEGs)18,19,20,21,22,23,24.
Despite progress in understanding the role of PRC2 in regulating endosperm development in rice, studies on how the two PRC2 FERTILIZATION INDEPENDENT ENDOSPERM (FIE) homologues suppress central cell initiation report inconsistent results14,25,26,27. By analysing multiple mutants of the closely linked Osfie1 and Osfie2 mutations induced via CRISPR/Cas9, we uncovered a novel function of PRC2 in suppressing cell division of the egg apparatus in the absence of fertilization, and provided further insight on the role of the OsFIE genes in reproductive development. Transcriptomic analysis revealed the activation of pluripotency factor genes OsBBM1 and OsWOX8/9 in asexual embryos and PEGs in autonomous endosperm. This study provides evidence of the pivotal roles of PRC2 in maintaining the repressive state of the egg (or egg apparatus) and central cell from autonomous division to ensure reproductive integrity in rice.
Results
CRISPR/Cas9-induced mutants at OsFIE1 and OsFIE2
Among ~150 T0 plants transformed with the CRISPR/Cas9 vector targeting the two FIE genes (Fig. 1a), we focused on plants with normal pollen dehiscence, as heterozygous fie mutants and other Polycomb mutants are not expected to give pollen sterility in heterozygous condition7,8,9,10,11,12,13,14. Four plants that showed distinct ~50% shrivelled seed formation without other morphological phenotypes were isolated as Osfie double or Osfie2 mutant candidates on the basis of phenotypic similarity to previously reported Polycomb mutants. Among other normal-looking plants, with random Sanger sequencing, we identified three putative homozygous Osfie1 (Fig. 1b,c and Table 1; designated as Osfie1-single1, 2 and 3) and three putative heterozygous Osfie2 mutants at the expected positions (Fig. 1b,c and Table 1; designated as Osfie2-single1, 2 and 3). The transgene-free homozygous Osfie1 plants isolated at T1 displayed no apparent phenotype as expected25 (Fig. 2a,b and Extended Data Fig. 1a–d). The transgene-free Osfie2 heterozygotes at T1 also had normal seed formation and set comparable to wild-type Nipponbare (Fig. 2c and Extended Data Fig. 1e–g), in contrast to the previous finding where an Osfie2 mutation caused seeds to shrivel14. By analysing T2 progeny derived from the transgene-free T1 heterozygotes of Osfie2-single1, we could not find homozygous mutants but found wild type (WT) and heterozygotes segregating at ~1:2 ratio (28:55) in the progeny (Table 1 and Fig. 2e,l,m), suggesting that the Osfie2 mutations probably caused homozygous lethality of embryos without causing seed shrivelling. We further analysed germinating seeds harvested from the transgene-free heterozygote of Osfie2-single1 to verify the embryo lethal phenotype. The OsFIE2 PCR fragments from the embryo and endosperm of non-germinated seeds were sequenced and showed that many of these seeds were homozygous for the Osfie2-2 mutation (Fig. 2n). The self-pollinated seeds of Osfie2-single1 containing arrested embryos were confirmed by confocal microscopy of developing seeds and cryo-section of mature seeds (Fig. 2f,g,i,j). We further hand sectioned the mature seeds from the heterozygote and found that ~25% of seeds (27/121) had abnormally differentiated embryos, while the other ~75% of the embryos appeared normal (Table 1 and Fig. 2e). Similarly, another two Osfie2 single mutants displayed similar genetic segregation and embryo abortion phenotype at T2, lacking homozygous mutant segregants (Table 1). This result suggests that OsFIE2 functions alone in embryo development but redundantly with OsFIE1 in endosperm development, suggesting that there is partial subfunctionalization between OsFIE1 and OsFIE2, leading to differential controls on embryo development (only by OsFIE2) and endosperm development (by both FIE genes).
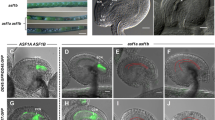
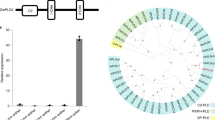
a, Positions at the exons of two OsFIE loci targeted by three gRNAs (g1, g2 and g3), two gRNAs targeting OsFIE1 and OsFIE2, respectively (g1 and g3) and one targeting an identical region between two genes (g2); OsFIE1 is ~9 kb away from OsFIE2. b, Mutant alleles induced by CRISPR/Cas9 at the two gene loci; black underline indicating gRNA target; red dotted line indicating the deletion and red letter indicating addition or substitution. c, Genotypes and associated gRNA targets (g1, g2 and g3) of three Osfie1, three Osfie2 and four double mutants used for analysis (Table 1).
a–d, Panicles with de-husked seeds: Nipponbare (a), Osfie1-single1 homozygote (b), Osfie2-single1 heterozygote (c) and Osfie-double1 heterozygote (d). Red asterisks show the aborted seeds; red arrowheads show the spiklets containing aborted seeds. e, T2 plants (83) from an Osfie2-single1 T1 heterozygote segregated into two genotypes: WT and heterozygote at a ~1:2 (28:55) ratio. Hand dissection of 121 T2 seeds shows that ~25% (27/121) had aborted embryos, which were considered to be Osfie2 homozygotes (Table 1). f, Normal 5-day-old embryo from an Osfie2-single1 heterozygote under confocal microscopy (n = 3). g, Abnormal 5-day-old embryo from an Osfie2-single1 heterozygote under confocal microscopy (n = 3). h, T2 plants (31) from an Osfie-double1 T1 heterozygote segregated into two genotypes: the double heterozygotes similar to the parent showing aborted seeds and Osfie1 homozygous segregants showing no aborted seeds with a ~1:1 (18:13) ratio. Detailed phenotyping of the aborted seeds by dissection shows ~50% (137/245) collapsed seeds containing aborted embryos (k), which were shown to be haploid (Fig. 5b–f). i, Well-differentiated embryos from Osfie2-single1 heterozygote by cryo-section at maturation (n = 3). j, Abnormal embryos from Osfie2-single1 heterozygote by cryo-section at maturation (n = 3). k, Aborted seeds from Osfie-double1 at maturation showing deformed embryo (red arrowhead) and endosperm (yellow arrowhead) (n = 137). l, Sanger sequencing of segregants (n = 28) of Osfie2-single1 heterozygote showing the WT allele at OsFIE2; red asterisks show 2 bp ‘AA’, which are deleted at the Osfie2-2 locus. m, Sanger sequencing of segregants (n = 55) of Osfie2-single1 heterozygote showing the heterozygote allele at OsFIE2. n, Sanger sequencing of embryos and endosperm (n = 11) of non-germinated seeds from Osfie2-single1 heterozygote showing the mutant allele with two base-pair deletion (−AA, red arrowhead) at the Osfie2-2 locus. Scale bars, 50 μm.
For the four T0 plants showing ~50% seed shrivelling, we found simultaneous mutations at both OsFIE1 and OsFIE2 loci (Figs. 1c and 2d, and Table 1; designated as Osfie-double1, 2, 3 and 4), with three having heteroallelic mutations and one being heterozygous at OsFIE1. All the Osfie1 mutations were predicted to cause loss of function. At OsFIE2, three plants were heterozygous and all the mutations were predicted to cause loss of function, while the fourth plant (termed Osfie-double3) had heteroallelic mutations (Fig. 1c), with Osfie2-7 being predicted to cause loss of function and Osfie2-6 predicted to be a weak allele that retains function, as it resulted in a serine to glycine substitution in a non-conserved region. We speculated that if Osfie2-6 caused loss of function, the heteroallelic Osfie2-6/Osfie2-7 genotype would have caused embryo lethality and this plant would not have been isolated. We selected transgene-free mutants at T1, which all showed shrivelled seed phenotypes (Extended Data Fig. 1h–k). From the mutant Osfie-double1 (Fig. 1c), we could identify 13 plants homozygous for Osfie1-4 showing no shrivelled seeds, and 18 plants heterozygous for Osfie2-4 and heteroallelic for Osfie1 displaying similar shrivelled seeds as the T1 parent, without double homozygous mutants being identified, suggesting that the linked loss-of-function mutations at both OsFIE loci could not be transmitted to T2 by the mutant female gametophyte (Fig. 2h). The double mutations could be transmitted by pollen and the mutant ovules carrying double mutations gave aborted seed upon fertilization (see below) (Table 1 and Fig. 2h). Using a progeny of the double mutant at T2, we confirmed the 1:1 segregation of the two genotypes at T3, with no other genotypes being identified. Similar segregation and seed abortion were seen in the other three double mutants at T4 for Osfie-double2 and T2 for Osfie-double3 and 4 (Table 1 and Fig. 1c). By dissecting the aborted seeds, we observed abnormally differentiated embryos of various morphologies, many with aborted endosperm (Fig. 2k and Extended Data Fig. 2a–p). The closely linked fie mutations caused seed abortion in a maternal gametophytic manner, a similar phenotype observed in the Arabidopsis FIE and other fis-class mutants9.
Asexual embryos and autonomous endosperm in double mutants
The shrivelled seed formation in the Osfie double mutants is similar to that in the Arabidopsis fie mutant and prompted us to investigate autonomous endosperm formation in the double mutants. We examined emasculated ovules of Osfie-double1 heterozygotes, with the Osfie1 homozygous segregants as control at T2 and sampled the ovules at different stages for confocal microscopy (Supplementary Table 1). Autonomous endosperms were seen in ovules at a frequency of ~50% (26/57) at 0 d post emasculation (DPE) (Figs. 3c,d and 5a, and Supplementary Table 1), with some ovules showing slight elongation (Extended Data Fig. 2q), while the other half exhibited a wild-type morphology (Fig. 3a,b). Interestingly, ~10% (6/57) of ovules that had autonomous endosperm also had globular embryo-like structures (Figs. 3c and 5a, and Supplementary Table 1), with no signs of eggs and synergids. Other ovules that had autonomous endosperm but no embryo always contained intact eggs characterized by having large nuclei and vacuoles but no synergids (Fig. 3d). We speculated that synergids might be disintegrated by autonomous endosperm in a similar way in which early sexually derived endosperm fuses to one remaining synergid soon after fertilization28. Ovules at 1, 2 and 3 DPE showed similar autonomous development of endosperm in ~50% of the ovules and the frequency of these asexually derived embryos occurring gradually increased (Figs. 3e and 5a, and Supplementary Table 1). From 6 DPE, we started to observe a few ovules with asexual embryos without clear presence of autonomous endosperm. We reckoned that the endosperm might degrade in these ovules. Therefore, we defined the ovules with embryos and/or endosperm as autonomous seeds for scoring the frequency of autonomous development (Fig. 5a and Table 1). At 6, 9 and 14 DPE, ovules showing autonomous seed formation accounted for ~50% of the total emasculated ovules, with the ratios of asexual embryos being elevated (5/27 at 6 DPE, 17/39 at 9 DPE and 9/26 at 14 DPE; Supplementary Table 1 and Fig. 5a). Ovules at 9 DPE contained structures resembling embryos and/or cellularized endosperm in ~50% of the ovules examined (Figs. 3f and 5a, and Supplementary Table 1), suggesting that the later asexual embryos were probably derived from eggs with the possibility that the embryos in early emasculated ovules could originate from synergids. At day 14, the asexual embryo-like structures exhibited various morphologies, with some showing signs of vascular bundles (Fig. 3g and Supplementary Table 1). At maturity, the autonomous seeds contained arrested structures, with some being accompanied by abnormally developed starchy endosperm as shown by staining with an iodine and potassium-iodide solution (Fig. 3i). The ovules (n = 59) of Osfie1 homozygous segregants did not display asexual embryo or autonomous endosperm formation after emasculation (Fig. 3j and Supplementary Table 1). We then pollinated WT with Osfie-double1 at T3 and analysed the autonomous phenotype in the F1 plants. All emasculated double heterozygous segregants in the progeny of the cross exhibited the asexual embryos (4/14) and autonomous seed formation (7/14), while the Osfie1 heterozygous segregants in the cross did not (0/58) (Supplementary Table 1 and Extended Data Fig. 3s,t).
a, Ovules of Nipponbare, with an egg (white arrowhead), two synergids (sy, one in focus by blue arrowhead), two central nuclei (cc) and antipodal cells (ap, grey arrowhead) at 0 DPE (n = 35). b, WT-looking ovules of Osfie-double1, with an egg (white arrowhead), two synergids (one in focus by blue arrowhead), two central nuclei and antipodal cells (insert) at 0 DPE (n = 31). c, Asexual globular embryos (em, red arrowhead) and syncytial endosperm (yellow asterisks) in Osfie-double1 at 0 DPE (n = 6). d, Ovules with autonomous endosperm (yellow asterisks) in Osfie-double1 at 0 DPE, with an egg cell (white arrowhead) (n = 26). e, Asexual globular embryos (red arrowhead) and cellularized endosperm (circled by yellow dashed line) in Osfie-double1 at 3 DPE (n = 5). f, Asexual globular embryos (red arrowhead) and cellularized endosperm (circled by yellow dashed line) in Osfie-double1 at 9 DPE (n = 17). g, Asexual embryos (red arrowhead) and autonomous endosperm (insert) in Osfie-double1 at 14 DPE, showing vascular structure (pink arrowheads) (n = 9). h, Asexual embryo (red arrowhead) and autonomous endosperm (en, yellow arrowhead) in a dissected seed of Osfie-double1 at 15 DPE (n = 17). i, Asexual embryo (red arrowhead) and autonomous endosperm (circled by yellow dashed line) in a dissected seed of Osfie-double1 at 15 DPE, showing starch granules (dark) being stained with iodine potassium-iodide solution (n = 3). j, Embryo sacs with an egg (white arrowhead) and central cell (insert) of an Osfie1 segregant of Osfie-double1 at 12 DPE, showing no autonomous development (n = 22). Scale bars, 50 μm.
We analysed three more independently isolated double mutant lines to further characterize asexual embryo formation and exclude the possibility that the phenotypes were caused by certain guide RNA (gRNA) combinations, off-targeting gRNA or tissue culture-induced effects. We examined ovules from Osfie-double2 at 0 DPE to investigate the origins of asexual embryos. We found one ovule (1/56) showing a four-celled pre-embryo with a degenerating synergid and limited autonomous endosperm nuclei, while others are either WT or ovules with more advanced asexual globular embryos as shown in other lines (Extended Data Fig. 3b and Supplementary Table 1). This four-celled embryo is full of vacuoles and resembles a dividing zygote, suggesting that it originated from an egg29. Among the progeny of Osfie-double2 at T4 and F1 of WT × Osfie-double4 (T3) (Fig. 1c), the emasculated double heterozygous segregants exhibited autonomous seed formation in ovules (53/99 for Osfie-double2; 40/78 for Osfie-double4) and asexual embryos (39/74 for Osfie-double2; 26/78 for Osfie-double4), while the WT or Osfie1 homozygous segregants did not (WT n = 68, Osfie1 n = 64; Table 1, Extended Data Fig. 3a–j,x–z and Supplementary Table 1). Similarly, among the F1 progeny of WT × Osfie-double3 (Fig. 1c), the autonomous phenotype only occurred in the ovules of double heteroallelic segregants (20/42 for autonomous seed formation; 13/42 for asexual embryo formation), while the ovules of the fie1-3 and Osfie2-6 double heterozygous segregants (n = 59) did not show autonomous phenotype as Osfie2-6 is a weak allele (Extended Data Fig. 3u–w, Supplementary Table 1 and Table 1). By analysing the Osfie-double3 and 4 at T2, we observed similar autonomous phenotypes in the ovules of double mutants carrying loss-of-function Osfie2 alleles, while the control segregants did not show autonomous growth (Supplementary Table 1 and Extended Data Fig. 3k–r). We further emasculated three independently isolated Osfie1 homozygotes and Nipponbare and found no asexual embryo or autonomous endosperm formation (Table 1, Supplementary Table 1 and Extended Data Fig. 4a–d). As all the mutants were generated by the same gene construct and were obtained by the same transformation process, the three independent Osfie1 homozygous lines and the four independent double mutant lines should have borne the same chance of having or not having the asexual embryo phenotype if off-targeting or tissue culture-induced effects were responsible. However, only the four double mutant lines exhibited the phenotype but not the Osfie1 lines, suggesting that the double mutations are probably responsible for the asexual embryo formation. Since all the 25 segregants, which did not carry loss-of-function Osfie2 alleles, exhibited no asexual embryo formation in the progeny of the selfed or out-crossed double mutant lines (Supplementary Table 1), if the gRNA off-targeting or tissue culture effects had caused the phenotype, the probability of all the 25 segregants showing no asexual embryo formation would be close to zero (50% to the power of 25, assuming each segregant had 50% equal chance of inheriting the off-targeting-induced mutations). Therefore, it is highly unlikely that gRNA off-targeting or tissue culture effects were involved in inducing the asexual embryo formation in the double mutants.
The gametophytic nature of the seed abortion and the autonomous seed formation, and the 1:1 segregation of two genotypes in the heterozygous double mutant lines, as in the Arabidopsis fie mutant, imply that meiosis must have occurred normally to give rise to the viable WT (or Osfie1) embryo sac and the Osfie1/Osfie2 embryo sac (Fig. 2h). Therefore, the egg apparatus-derived asexual embryos are probably haploid. To test this, we were able to induce callus formation from 3 of ~100 asexual embryos with enough biomass for detection of DNA content by flow cytometry30,31. As expected, all the induced calli (n = 3) were haploid, while the DNA contents from WT leaves and calli induced from sexually derived embryos were diploid (Fig. 5b–e, Extended Data Fig. 5 and Supplementary Table 2). Given that the collapsed seeds of the double mutants under self-pollination morphologically resemble the autonomous seeds (Fig. 3h,i and Extended Data Fig. 2) and ~50% ovules of double mutants showed autonomous development at 0 DPE, we reasoned that ovules with the double mutations must have initiated autonomous development before pollen dehiscence and that the defective embryos were of asexual origin. We successfully induced 3 calli from ~100 abnormal embryos from self-pollinated ovules and found that these calli contained similar DNA contents as the asexual embryos (Fig. 5b–e). We further verified the maternal genotype of aborted embryos by pollinating Osfie-double1 with an indica rice 9311. By analysing the genotypes of both viable and aborted seeds via sequencing of the PCR products amplified from a region containing single nucleotide polymorphisms (SNPs) at OsYUCCA11 (Fig. 5f), we observed that the aborted embryos (n = 11) only showed the maternal SNP, while the viable embryos (n = 16) had two parental SNPs, suggesting that the aborted seeds were of asexual origin.
Asexual pre-embryo-like structures in Osfie2 mutants
These novel asexual embryo-like structures in the four independent Osfie double mutants prompted us to investigate whether this phenotype also occurred in the three independent Osfie2 mutants. This would clarify whether the two OsFIE genes play a redundant role in modulating the autonomous phenotype. We emasculated all three independent OsFIE2 +/− heterozygous lines at T2 and T3, using WT segregants as control (Supplementary Table 1). In Osfie2-single3, we found the embryo sacs from 2 to 4 DPE (n = 48) displaying a WT morphology (Fig. 4a), as in Nipponbare (Fig. 3a), and giving neither asexual embryo nor autonomous endosperm phenotypes. At 6 DPE, we observed one multicellular structure resembling an early embryo at the micropylar end of the emasculated ovule, accompanied by an egg-like cell characterized by having a large nucleus with clear nucleolus and vacuoles, two central cell nuclei and proliferated antipodal from 22 ovules (Fig. 4b and Supplementary Table 1). At 11 DPE, we observed more asexual pre-embryo-like structures (5/33), sometimes without an egg (Fig. 4c and Supplementary Table 1). At 12 DPE, we observed more asexual pre-embryo-like structures in ovules (5/23), while the WT segregants gave no such structures after emasculation (n = 72; Fig. 4d and Supplementary Table 1). At T3, 12 asexual pre-embryos were observed in 63 ovules from three heterozygous segregants (Supplementary Table 1). In Osfie2-single1, the embryo sacs from 0 to 6 DPE (n = 85) exhibited no asexual embryo formation but a WT morphology (Fig. 4e). At 9 DPE, we observed one asexual pre-embryo-like structure (1/8) at the micropylar end of the emasculated ovule. At 15 DPE, we observed more similar structures (8/34), with 6 more spotted without scoring the total ovules (Fig. 4f,g and Supplementary Table 1). At T3, 22 asexual pre-embryos were observed in 99 ovules from three heterozygous segregants. The ovules of WT segregants at T2 gave no asexual embryo formation after emasculation (n = 42) (Supplementary Table 1 and Fig. 4h). In Osfie2-single2, we observed similar asexual embryo-like structures with a similar frequency in the 8 and 12 DPE ovules (6/33) at T2 and 21 DPE ovules (5/22) at T3, while the ovules (n = 78) of WT segregants did not display asexual embryo formation (Extended Data Fig. 4e–h, Table 1 and Supplementary Table 1). We did not find any autonomous division of the central cell in all Osfie2 mutants, in contrast to the previous finding of a low frequency of autonomous endosperm formation in Osfie2 mutants14. The absence of asexual embryo formation in all 10 WT segregants of the selfed Osfie2 heterozygous lines and the presence of asexual embryo formation in all the single Osfie2 and double mutant lines support the idea that the single Osfie2 mutation is responsible for the asexual embryo formation. The female gametophyte carrying the Osfie2 single mutation could be fertilized and gave rise to normal-looking heterozygous plants (Fig. 2e), suggesting that these asexual embryos derived from the Osfie2 embryo sacs are also haploid. We further scored the asexual pre-embryos of these lines from quality confocal images for the presence and absence of accompanying eggs and found that ~40% (28/71) were accompanied by egg-like cells, while others were not, indicating that these structures might well be derived from the eggs and synergids. The asexual embryo-like structures in the Osfie2 mutants remained small compared with those in the double mutants and arrested without further development, suggesting that the autonomous endosperm in the double mutants facilitated the asexual embryo development.
a, WT-looking embryo sacs in Osfie2-single3 at 2 DPE, with an egg (white arrowhead), two synergids (one in focus by blue arrowhead) and two central nuclei (n = 25). b, Asexual pre-embryo-like structure (red arrowhead) in Osfie2-single3 at 6 DPE, with an egg-like cell (white arrowhead) and central cell nuclei (insert) (n = 1). c, Asexual pre-embryo-like structures (red arrowhead) in Osfie2-single3 at 11 DPE, with central cell nuclei and without an egg (n = 5). d, Embryo sacs of a WT segregant in Osfie2-single3 at 12 DPE, showing an egg (white arrowhead) and two central nuclei (n = 18). e, WT-looking ovules of Osfie2-single1 at 0 DPE, showing an egg (white arrowhead) and two central nuclei (n = 8). f, Asexual embryo-like structures (red arrowhead) in Osfie2-single1 at 15 DPE, accompanied by an egg-like cell (white arrowhead) and two central cell nuclei (insert) (n = 14 in f and g). g, Asexual embryo-like structures (red arrowhead) in Osfie2-single1 at 15 DPE, with central cell nuclei and without an egg. h, Embryo sacs of WT segregants of Osfie2-single1 at 12 DPE, showing an egg (white arrowhead) and two central nuclei (n = 20). i, Emasculated ovules of Osfie2-single1 treated with 2,4-d containing a proliferated asexual embryo-like structure at 7 DPE, with central cell nuclei (insert) (n = 3). j, Emasculated ovules of Osfie2-single3 treated with 2,4-d containing a proliferated asexual embryo-like structure at 5 DPE, with central cell nuclei (insert) (n = 6). Scale bars, 50 μm.
Previous studies suggest that FIS PRC2 represses auxin production in the central cell, and fertilization brings in the paternally expressed auxin biosynthesis genes that trigger central cell division. Exogenous application of auxin 2,4-d induces autonomous endosperm development in Arabidopsis16. Emasculated florets were treated with a synthetic auxin (2,4-d) of different concentrations from 10 µm to 200 µm, but no endosperm or embryo formation was induced in wild type or Osfie1, while the pericarps elongated as expected (Extended Data Fig. 4i–l). Interestingly, by treating emasculated Osfie2-single3 and Osfie2-single1, we observed proliferated cell masses with early emergence at the micropylar ends of the emasculated ovules at a frequency of ~20% (9/50 for Osfie2-single3 at 5 and 6 DPE, and 8/43 at 2–4 DPE; 11/60 at 15 DPE for Osfie2-single1) similar to the asexual pre-embryos in the Osfie2 mutants without treatment (Supplementary Table 1 and Fig. 4i,j), suggesting that the external application of auxin facilitated the early onset and cell division of the asexual pre-embryos. However, these proliferated cell masses did not show any signs of differentiation.
Transcriptomic analysis of asexual embryos and autonomous endosperm
To understand how the FIE function represses autonomous development, we isolated RNAs from the autonomous endosperm and asexual embryos from selfed Osfie-double1 with three biological repeats to generate transcriptome data using the Illumina platform (Supplementary Table 3). We also included the transcriptomic datasets from the sexually derived embryo and endosperm in our previous study32, the egg, the zygotes of different stages3,33 and somatic embryos34. A principal component analysis (PCA) of these transcriptomes showed that autonomous endosperm and asexual embryos are grouped closer to each other than to other tissues including embryos, somatic embryos, endosperm, egg and zygotes (Extended Data Fig. 6a), suggesting that the maternal origin and/or severely perturbed development of these autonomous structures with compromised H3K27me3 might affect similar target genes and contribute to the close positioning of these two sets of transcriptomic data.
a, Frequency of asexual embryo and autonomous seed formation at different days post emasculation (extracted from Supplementary Table 1). b–d, Flow cytometric DNA histograms for ploidy levels of Nipponbare (Nip) calli (b), calli from aborted embryo of self-pollinated seeds (c) and asexual embryo (d) of Osfie-double1; the gating borders and peaks for internal standards (Bellis perennis) are shown in Extended Data Fig. 5. The positions of peaks for the internal control were slightly different between the histograms (Extended Data Fig. 5) and would not affect measurement of the absolute DNA contents for each sample (Supplementary Table 2). e, Cellular DNA content of aborted embryos under pollination, asexual embryos and wild-type tissues by flow cytometry (Supplementary Table 2). f, Sanger sequencing of OsYUCCA11 PCR fragments amplified from embryos of the double heterozygote (in Nip background) pollinated with 9311 (indica), showing that viable embryos were hybrids with double peaks (left, n = 16) and aborted embryos are maternal with a single Nip peak (right, n = 11); red asterisks indicate the SNPs between Nip and 9311. g, Expression of OsFIE1 and OsFIE2 in different tissues using publicly available transcriptomic data. CC, central cell. Data are presented as mean ± s.d. of 3 (egg, zygote2.5h, zygote5h, zygote9h, somaticEM1, somaticEM2), 2 (SexEM and SexEN) and 1 (CC) biological replicate.
We identified over 17,000 commonly expressed genes in the transcriptomes of asexual, sexual and somatic cell-induced embryos (Extended Data Fig. 6b), which include genes that modulate the early pattern formation of the somatic embryo in rice3,34,35,36,37,38,39,40,41,42,43,44. Those genes are expressed at a comparable level to that in the sexual embryo (Extended Data Fig. 6c), suggesting that these asexual structures had acquired some degree of the characteristics of a sexual embryo at the molecular level. We identified 8,352 differentially expressed genes, including 5,535 upregulated and 2,817 downregulated genes between asexual embryos and sexual embryos (Supplementary Table 4 and Extended Data Fig. 7a). Gene Ontology (GO) term enrichment analysis of the differentially expressed genes in asexual embryos showed that they comprised diverse molecular functions and biological processes, consistent with the extensive disruption of the development and differentiation of the asexual embryos (Supplementary Table 5). We then focused on genes with possible functions in embryogenesis. The genes with de novo expression in zygote compared with egg may be involved in the initiation of embryogenesis3,33, among which some are male genome-expressed genes upon fertilization, while the female alleles in the egg and zygote are silenced. We asked how many of these genes with de novo expression in zygote are also expressed in the asexual embryo. We also extended the comparison to other BBM and WUSCHEL family members which are not included in the list of refs. 3,33. We found 36 genes activated in the asexual embryo (Supplementary Table 6), including 1 out of 4 BBM-like and 8 out of 11 WUS family genes, suggesting that the rice FIE genes may directly or indirectly repress expression of these genes in the egg apparatus and that they became activated without involvement of the paternal genome due to the mutations at the FIE loci. Among these genes with known embryogenic functions in plants are transcription factors, OsBBM1 (LOC_Os11919060) and WUS-LIKE HOMEOBOX GENES5,6,7,45 (Fig. 6a).
a, Heat map showing expression levels of BBM1 and WUS family genes in asexual embryo (asexEM) egg3,33, embryo32, somatic embryo (somaticEM;34) and zygote3,33 (Supplementary Table 6). b, Heat map showing the expression levels of paternal alleles (EN_pat) and maternal alleles (EN_mat) of MEGs in hybrid endosperm (Supplementary Table 13; ref. 32) and expression levels of the same genes in autonomous endosperm (auto_EN; Supplementary Table 13). Lane EN shows the total expression level of combined parental alleles of MEGs in hybrid endosperm. Small bars on the left (in b and c) indicate which genes are marked with H3K27me3 at the maternal allele (green bars) or the paternal allele (purple bar), or have no marks detected (no bars) (Supplementary Table 15). c, Heat map showing the expression levels of paternal alleles (EN_pat) and maternal alleles (EN_mat) of PEGs in hybrid endosperm (Supplementary Table 12; ref. 32), and expression levels of the same genes in autonomous endosperm (auto_EN; Supplementary Table 12). Lane EN shows the expression levels of combined parental alleles of PEGs in hybrid endosperm. Red asterisk represents OsYUCCA11.
Similarly, GO-term enrichment analysis on the differentially expressed genes (Supplementary Tables 7 and 8, and Extended Data Fig. 7b) between autonomous and sexual endosperm indicated that a wide range of molecular functions and biological processes of the autonomous endosperm had been altered (Supplementary Table 8). As auxin production and signalling in the primary endosperm are characteristic events after fertilization, we further analysed the expression of genes that are involved in auxin biosynthesis, transportation and signalling in autonomous endosperm (Supplementary Table 9). Most genes are expressed in autonomous endosperm, with YUCCA genes being expressed at a lower level than in sexual endosperm, indicating a possible low activity of auxin biosynthesis in the autonomous endosperm. The staining with an iodine and potassium-iodide solution of starch granules in autonomous endosperm prompted us to analyse the activity of genes involved in starch biosynthesis (Supplementary Table 10). Most of the starch biosynthesis genes have lower expression in autonomous endosperm than in sexual endosperm, consistent with the poor accumulation of starch in autonomous seeds, and this might be due to the lower gene dosage in autonomous endosperm. In rice, type I MADS-box transcription factor genes have been speculated to be involved in endosperm cellularization as is the case for the counterpart AGL62 in Arabidopsis. We found that OsMADS77 and 87 and several other MADS-box genes are activated in the autonomous endosperm as observed in Osemf2a15,46 (Supplementary Table 11 and Extended Data Fig. 7c–k).
Endosperm is characterized by expression of maternally or paternally expressed imprinted genes. De-repression of paternally expressed imprinted genes YUCCA10 and TAR1 in auxin biosynthesis is thought to trigger autonomous endosperm development in Arabidopsis PRC2 fis-class mutants18,19,32,47,48,49,50. We compared the relative expression levels of all the maternally expressed imprinted genes (MEGs)32 between autonomous endosperm and sexual endosperm and found that most of the MEGs are expressed in autonomous endosperm (Fig. 6b and Supplementary Table 13). Interestingly, 100% (57/57) of the PEGs32 were also expressed in the autonomous endosperm, showing comparable levels to those in sexual endosperm (Fig. 6c and Supplementary Table 12). As the autonomous endosperm is maternally derived, we conclude that loss of FIE function activates the maternal alleles of the PEG loci. It is worth noting that the PEG OsYUCCA11 was maternally activated in the autonomous endosperm (Fig. 6c and Supplementary Table 12). We used chromatin immunoprecipitation (ChIP-seq) with antibody against H3K27me3, followed by deep sequencing to identify H3K27me3-marked regions in the maternal or paternal genomes of endosperm from the reciprocal crosses between Nip and 9311 (Supplementary Method). The ChIP assay result suggests that the PRC2-modulated H3K27me3 targeted the maternal alleles of PEGs in endosperm (Fig. 6c, Supplementary Tables 14–17, Extended Data Figs. 8 and 9, and Supplementary Method), consistent with the idea that PEGs are activated in the autonomous endosperm16.
We then investigated the gene expression patterns of OsFIE1 and OsFIE2 by analysing transcriptomic data in egg, zygote, embryo, somatic embryo, endosperm3,32,33,34 and central cell (an unpublished transcriptome deposited in NCBI (DRR000623)). OsFIE1 is relatively highly expressed in endosperm and central cell but very lowly expressed in egg and zygote (Fig. 5g), while OsFIE2 is ubiquitously expressed in all tissues, consistent with the mutant phenotypes that OsFIE1 and OsFIE2 redundantly repress central cell division and regulate endosperm development, while OsFIE2 functions independently in the embryo to repress egg apparatus division.
Discussion
Our study uncovers a novel role for the PRC2 component OsFIE2 in suppressing the autonomous division of the egg apparatus, in addition to the expected role of OsFIE1 and OsFIE2 in suppressing central cell division before fertilization in rice. The asexual pre-embryo-like structures in Osfie2 mutants started to emerge at the micropylar end of emasculated ovules ~1 week after emasculation, with higher frequency of these structures being observed at later stages. Some of the asexual pre-embryos co-located with egg-like cells, while others existed without accompanying eggs, suggesting that asexual embryo formation might be derived from synergids and eggs. Further study is required to characterize the exact cell lineage of these asexual embryos. However, the analysis of the asexual embryo formation phenotypes in the Osfie1Osfie2 double mutants suggests that asexual embryos are mainly egg derived. In the early ovules with autonomous endosperm, only a small portion of ovules contained asexual globular embryos, with the rest containing egg-like cells without accompanying synergids. The synergids might be disintegrated by autonomous endosperm in a way similar to the typical fusing of the early sexually derived endosperm with one remaining synergid after fertilization28. This allowed the egg to become the major cell type to develop into parthenogenetic embryos in the ovules of later stages. Rarely, a four-celled pre-embryo resembling a dividing zygote was observed, coexisting with a synergid and endosperm nuclei, further supporting the egg origins of most asexual embryos. We reasoned that the lack of autonomous endosperm formation in the Osfie2 mutants might have allowed the synergids to survive without being consumed by endosperm28 and facilitated the formation of synergid-derived embryos. However, there is a possibility that the asexual embryos in early ovules might also be synergid derived in the double mutants as those in Osfie2 mutants—a possibility requiring further investigation.
In contrast to the late emergence, low penetrance and early arrest of asexual pre-embryos in Osfie2, the earlier onset, higher frequency and better development of the asexual embryos in the double mutants suggest that the autonomous endosperm facilitates the asexual embryo development of the Osfie2-activated egg apparatus potentially by providing signals or nutrients. This is supported by the observation of starch granule formation and expression of genes that are involved in auxin biosynthesis, transportation, signalling and starch biosynthesis in autonomous endosperm. The asexual pre-embryo structures emerged earlier and grew into bigger cell masses in the Osfie2 mutants when the emasculated ovules were treated with synthetic auxin 2,4-d, indicating that auxin may be one of the important signals to support embryo growth. A similar phenomenon has been observed in Arabidopsis where autonomous endosperm derived from the PRC2 fis1/mea mutant was able to support haploid embryo development51.
Our finding contrasts with a similar study where single knockout of OsFIE2 and double knockout of the two FIE genes both caused autonomous endosperm formation at a very low frequency without asexual embryo formation phenotype14. In our study, the double linked mutations at both OsFIE loci induced very high penetrance of asexual embryo formation and autonomous endosperm phenotype, while the Osfie2 single mutations caused asexual pre-embryo-like structures at a lower frequency. We used the same vector containing three guides, with two guides targeting the two OsFIE genes and one common guide targeting both genes to generate all mutants in the same transformation process. Three independently isolated Osfie1 homozygotes, two induced by the specific OsFIE1 guide and one by the common guide, did not show any autonomous phenotypes, while the three independent Osfie2 heterozygotes by the common guide and the four double mutants by different guide combinations exhibited asexual embryo formation, with the control segregants displaying none of the phenotypes. It is unlikely that the phenotype was caused by gRNA off-targeting or tissue culture effects during transformation, as each of the 10 analysed lines should have borne the same chance to have or not have the phenotype if the Osfie2 mutations were not responsible for the phenotype. The fact that there was no asexual embryo formation in the segregants lacking the Osfie2 loss-of-function mutations in the progeny of the Osfie2 or double mutant lines further support the idea that the Osfie2 mutations caused the asexual embryo formation. The possible reason for the discrepancy between the two studies is that the mutants were generated in different genetic backgrounds where unknown modifiers may exist to suppress the autonomous phenotype13,14. The low penetrance of the autonomous endosperm phenotype might obscure the observation of the embryo phenotype in ref. 14, as we showed that the high penetrance of the autonomous endosperm facilitates the prevalence of the asexual embryos. In Arabidopsis, a likely parthenogenetic pre-embryo phenotype has been observed in the mutant of the non-canonical PRC2 member AtMSI1; however, this phenotype has not been convincingly demonstrated except for the autonomous endosperm phenotypes in other PRC2 fis-class mutants10,52. It remains to be investigated whether asexual embryo formation triggered by the loss of the PRC2 only occurs in certain species or even only exists in certain genetic backgrounds in rice. It is also tempting to revisit whether the egg in the Arabidopsis fie mutant would undergo limited division as in the Atmsi1 mutant. Similar to the findings in the moss Physcomitrella patens that mutations in PRC2 genes result in fertilization-independent production of a sporophyte-like body on side branches of the gametophytic protonema filaments53,54, the formation of asexual embryos in the rice mutants suggests that the PRC2 complex may function as a universal mechanism via H3K27me3 to maintain reproductive integrity.
The transcriptomic data were generated from tissues in which the development programme was severely perturbed by the loss-of-function mutations of the essential epigenetic modification genes. Therefore, the results of the transcriptomic analysis should be interpreted with caution, especially when compared with transcriptomes from sexually derived embryos or endosperm. Nevertheless, we found that the rice BBM and WUS homologues which are silenced in egg3,4 are expressed in the asexual embryos, suggesting that these genes must be activated at some stages during asexual embryo formation without the involvement of male genome. Ectopic expression of the BBM and WUS genes in Arabidopsis, brassicas and cereals has been shown to promote somatic embryogenesis and shoot regeneration from tissue culture, suggesting that they act as pluripotency factors55,56,57. In rice, OsBBM1, OsBBM2 and OsWOX8/9 are initially male genome-expressed in the early zygote and ectopic expression of OsBBM1 in egg triggers parthenogenesis, suggesting that the requirement for fertilization in embryogenesis is mediated by paternal genome transmission of pluripotency factors3,4. We propose that OsBBM1 and other pluripotency factors, such as WUS homologues, are repressed by the PRC2 complex directly or indirectly in the egg apparatus until fertilization, which brings the male genome-expressed pluripotency factors. Loss of the FIE function leads to de-repression of the pluripotency factors via the loss of the H3K27me3, which triggers asexual embryo formation.
Similarly, maternal activation of the known PEGs in autonomous endosperm indicates that the maternal alleles of those PEGs were de-repressed by the fie mutations. The expression of both MEGs and PEGs in the autonomous endosperm suggests that the fie-activated central cells may have sufficient stimuli for autonomous division as the fie-activated egg apparatus. The repressed state of the central cell due to PRC2-mediated H3K27me3 can be overridden by the introduction of the active paternal alleles of PEGs with fertilization24, as occurs in the zygote where fertilization introduces the male genome-expressed pluripotency factors (such as OsBBM1) and these paternally derived PEG products act as pluripotency factors (including auxin biosynthesis genes24) to trigger endosperm formation. This suggests that the embryo and endosperm progenitor cells convergently adopt PRC2 to suppress asexual embryo and autonomous endosperm formation possibly through silencing of maternal alleles of male genome-expressed genes before fertilization. Our study sheds light on the interplay between epigenetic regulation and fertilization to ensure proper seed development. The mechanism of FIS-PRC2 repression of asexual embryo and autonomous endosperm formation and the high penetrance of the autonomous phenotype in the mutant may help to induce haploids for speed breeding58 or be harnessed to engineer synthetic apomictic crops—an emerging revolutionary tool for fixing heterosis and enhancing yield4,59,60,61,62.
Methods
Generation of CRISPR/Cas9-edited mutants
The rice genome has two FIE homologues (LOC_Os08g04270 and LOC_Os08g04290) that are closely linked (Fig. 1a). The CRISPR/Cas9 editing method was used for mutant generation as previously described63. We cloned three single guide RNA (sgRNA) sequences in the transformation binary vector. The target seed sequences (~20 bp) of the three sgRNAs were selected from the OsFIE1 (LOC_Os08g04290) and OsFIE2 (LOC_Os08g04270) coding regions, with two specifics for OsFIE1 (g3: GTCACCGACACGAAGTACT) and for OsFIE2 (g1: TCGTTCTACACACTGAGTT), respectively, and the third one targeting both genes (g2: CTCATCATTTCTGCAAGCA) (Fig. 1a). This would maximize the chance of generating mutations respectively and simultaneously at both closely linked loci. The rice small nuclear promoters OsU3, OsU6a and OsU6b were used to drive g1, g2 and g3 sgRNA, respectively, and the DNA for the three sgRNA expression were synthesized by IDT, with Type II restriction enzyme Bsa I sites attached (Supplementary Table 18). These synthesized sequences were inserted into the binary vector pYLCRISPR/Cas9-MH using the GoldenGate strategy, with Type II restriction enzyme Bsa I for digestion and T4 DNA ligase for ligation63.
The calli were induced from mature rice seeds (Oryza sativa ssp. japonica cv. Nipponbare) and transformation was performed by using Agrobacterium tumefaciens strain GV3101. Positive transformed calli were screened by hygromycin, and then used to regenerate transgenic plants64.
Growth condition and mutation detection
The wild type of rice (O. sativa ssp. japonica cv. Nipponbare) and transgenic plants were grown in a glasshouse at 28 °C during the day and at 20 °C during the night under natural light. Gene-specific primers were designed to identify the mutations around target sites. Four sets of primer pairs flanking the target sequences were used to amplify the DNA isolated from transgenic plants for mutation detection. They are OsFIE2-T1-F: ACCTACAGCTGCCTCAAGGA and OsFIE2-T1-R: TATCAGCCACGTAGCAAGCA for target 1 at OsFIE2, OsFIE2-T2-F: GGTGGAAGATGTAGAACCTAGTGG and OsFIE2-T2-R: ATCCTATGCAATGCCATGTGAAA for target 2 at OsFIE2, OsFIE1-T2-F: TGTGGTTTCAGTGGGTCTTTAGC and OsFIE1-T2-R: TAAGATCCCTGTCTGCACATTCC for target 2 at OsFIE1, and OsFIE1-T3-F: CTGTGGAATGTGCAGACAGGGATC and OsFIE1-T3-R: GTGACATCAGAAGCTGGATGAGT for target 3 at OsFIE1 (Supplementary Table 18). The PCR products were directly used for Sanger sequencing and mutations were deduced from the sequencing traces. The transgenic plants, or transgene-free plants, were used for DNA extraction (Qiagen DNeasy Plant Pro Kit) and PCR amplified with above primers for Sanger sequencing to score the genotype.
Emasculation and microscopy
Florets predicted to flower within 1–2 d were used for emasculation after removing other younger florets and opened florets. The emasculation was performed by removing the un-opened stamens after cutting off the top end of florets during late afternoon. The treated panicles were protected in paper bags to avoid being outpollinated.
Emasculated florets or pollinated florets were immediately fixed in freshly prepared FAA (formaldehyde:acetic acid:glycerol:alcohol at 5:6:5:50 + 34% distilled water)65 for at least 24 h, then washed with 50% ethanol and stored in 70% ethanol at 4 °C. The samples were hydrated sequentially in 50% ethanol, 30% ethanol and distilled water for 30 min at each stage. Hydrated caryopses were pre-treated in 2% aluminum potassium sulfate for 20 min and stained with 10 mg l−1 eosin B solution for 12 h at room temperature. The samples were post-treated in 2% aluminum potassium sulfate for 20 min, rinsed in distilled water three times and dehydrated in a series of ethanol solutions (30%, 50%, 70%, 90%, 100%). The samples were transferred to 50% ethanol:methyl salicylate for 2 h and then cleared in pure methyl salicylate for at least 2 h before confocal microscopy imaging. Images were collected using a Leica SP8 laser scanning confocal microscope (Leica) equipped with ×10 (NA = 0.3), ×20 (NA = 0.75) and ×40 (NA = 1.1) water immersion objectives. Excitation wavelength was 543 nm and an emission range of 510–650 nm66 was collected with a PMT detector at a pixel resolution of 2,048 × 2,048. All images were collected and pseudo-coloured with the Green LUT using Leica Application Suite v.3.5 (LASX, Leica Microsystems). The autonomous phenotypes were scored by the presence of asexual embryos and/or autonomous endosperm in the emasculated ovules of the analysed Osfie mutants to avoid the occasional pseudo-parthenocarpy in emasculated ovules. The identity of the egg cell was determined by the characteristic presence of a large nucleus with a large nucleolus and the formation of vacuoles, while synergids were identified by being denser, having a smaller nucleus and the presence or absence of a smaller nucleolus.
For starch granule detection, the autonomous seeds were stained using an iodine and potassium-iodide solution67 (2 mg I2, 20 mg KI ml−1). Images were collected using a Leica m205c dissecting microscope equipped with a ×0.63 objective and a Leica IC90e digital camera (Leica).
For cryo-sectioning, developing seeds were cut in half and fixed in FAA for 48 h. Fixed seeds were embedded in OCT (Tissue-Tek, Sakura Finetek) medium and stored at −20 °C for over 2 h before sectioning. Sections (10 µm thick) were cut at −20 °C using a Leica CM1850 cryostat (Leica) and mounted on microscope slides (Fisher Scientific) at room temperature. Images were collected using a Zeiss AxioImager M1 fluorescence microscope equipped with a Zeiss Axiocam 712 colour CCD camera and plan-apochromat ×5 (NA = 0.5) objective, using ZEN 3.2 acquisition software (Carl Zeiss). All images were processed using Photoshop CC (Adobe).
The parent-of-origin analysis of asexual embryo
The embryos from the double heterozygous mutants pollinated with indica rice 9311 were isolated for DNA extraction individually. A pair of primers flanking an SNP at OsYUCCA11 (ref. 32) between Nipponbare (Nip) and 9311 was used to amplify DNA isolated from the aborted embryos. The PCR products were used for Sanger sequencing.
Autonomous seeds were harvested, sterilized in 20% bleach and washed 5 times with sterilized water. The asexual embryos were dissected out and used for callus induction using N6D medium64. The induced calli (~4 weeks induction) and internal control leaves of Bellis prennis (50 mg) were placed in a Petri dish on ice. The samples were gently chopped in 500 μl of modified Galbraith’s buffer68 for 30 s and gently mixed. After adding another 500 μl buffer, the samples were filtered through a two-step filter (42 µm first, then 20 µm) and collected in a flow cytometry sample cup. RNase (50 μl, 10 mg ml−1), 50 μl propidium iodide stock (1 mg ml−1) and 2 μl beta-mercapethanol were added to each sample for flow cytometry assay (Beckman Coulter). The DNA content and ploidy level were analysed30,31.
RNA isolation and transcriptomic analysis
Asexual embryos and autonomous endosperm from ~20 ovules at 9 d post emasculation were harvested and collected in 1.5 ml RNase-free Eppendorf tubes for RNA isolation using TRIzol (Thermo Fisher) on ice. RNA quality was assessed using the Agilent Bioanalyzer with RNA integrity number values over 7. Freeze-dried triplicate RNA samples were sent to Novogene for transcriptome sequencing on an Illumina platform. The reads for the six transcriptomic data were deposited in NCBI under BioProject PRJNA786704. The public transcriptome data were downloaded from the NCBI FTP site (Supplementary Table 3). All data were aligned to the reference genome sequence of O. sativa ssp. japonica cv. Nipponbare69 (MSU 7.0) using HISAT (v.2.2.0)70. The alignment results were output as bam files. Samtools (v.0.1.19)71 was used to sort and index the bam files containing the aligned reads. The alignments were visualized using IGV genome browser72. The reference genome mapping ratio of the alignments was also counted by Samtools (Supplementary Table 3). Read counts were generated by featureCounts (v.2.0.1)73. The gene expression value was calculated using the transcript per million (TPM) method, which was based on reads counts and transcript length. After log2 normalization, heat mapping and clustering analysis were performed using TBtools74. PCA was performed using the online platform Majorbio Cloud (www.majorbio.com)75.
On the basis of the RNA-seq raw counts, differentially expressed genes (DEGs) analysis was performed using DESeq2 (ref. 76) in the R package for comparison between asexual embryo and sexual embryo, and between asexual endosperm and sexual endosperm. Genes with |log2(fold change)| > 2 and adjusted P value (Padj) < 0.01 were selected as DEGs (Supplementary Tables 4 and 7). DEGs were visualized using volcano plots in TBtools, which effectively displays the significance against the log2 fold change of the genes and highlights genes that are most differentially expressed (Extended Data Fig. 7a,b).
GO annotation of the whole genome was downloaded from Biomart (http://www.biomart.org). Biological Network Gene Ontology (BiNGO v.3.0.5)77, a Cytoscape78 plugin, was used to analyse GO enrichment and display the GO network diagram. Enrichment significance was determined using a hypergeometric test, with terms having a corrected P value below 0.05 being considered as enriched (Supplementary Tables 5 and 8).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Transcriptomic and ChIP-seq data generated in this study are deposited in the NCBI database under BioProject PRJNA786704, accession numbers: SRR17151221, SRR17151220, SRR17151219, SRR17151224, SRR17151223, SRR17151222, SRR17210911, SRR17210910, SRR25655515, SRR25655516, SRR25678596, SRR25678595. Other transcriptomic data used in the analysis were downloaded from NCBI including the BioProject PRJNA218883 (accession numbers: SRR976336, SRR976337, SRR976338, SRR976339, SRR976340, SRR976341, SRR976335, SRR976342, SRR976343), BioProject PRJNA295002 (accession numbers: SRR2295903, SRR2295904, SRR2295905, SRR2295906, SRR2295907, SRR2295908), BioProject PRJNA412710 (accession numbers: SRR6122716, SRR6122707, SRR6122710, SRR6122706, SRR6122708, SRR6122722, SRR6122709, SRR6122720, SRR6122704, SRR6122715), BioProject PRJDA51201 (accession number: DRR000623). Source data are provided with this paper.
Code availability
Scripts specifically for the integrated data analysis of ChIP-seq datasets and SNP bias in hybrid rice from endosperm from the reciprocal crosses between Nipponbare (Nip) and 9311 are available on GitHub (https://github.com/biz007/NPLANTS-211112076-CSIRO).
References
Dresselhaus, T. & Jurgens, G. Comparative embryogenesis in angiosperms: activation and patterning of embryonic cell lineages. Annu. Rev. Plant Biol. 72, 641–676 (2021).
Palovaara, J., de Zeeuw, T. & Weijers, D. Tissue and organ initiation in the plant embryo: a first time for everything. Annu. Rev. Cell Dev. Biol. 32, 47–75 (2016).
Anderson, S. N. et al. The zygotic transition is initiated in unicellular plant zygotes with asymmetric activation of parental genomes. Dev. Cell 43, 349–358.e4 (2017).
Khanday, I., Skinner, D., Yang, B., Mercier, R. & Sundaresan, V. A male-expressed rice embryogenic trigger redirected for asexual propagation through seeds. Nature 565, 91–95 (2019).
Kwong, R. W. et al. LEAFY COTYLEDON1-LIKE defines a class of regulators essential for embryo development. Plant Cell 15, 5–18 (2003).
Conner, J. A., Mookkan, M., Huo, H., Chae, K. & Ozias-Akins, P. A parthenogenesis gene of apomict origin elicits embryo formation from unfertilized eggs in a sexual plant. Proc. Natl Acad. Sci. USA 112, 11205–11210 (2015).
Chen, B. et al. BABY BOOM regulates early embryo and endosperm development. Proc. Natl Acad. Sci. USA 119, e2201761119 (2022).
Chaudhury, A. M. et al. Fertilization-independent seed development in Arabidopsis thaliana. Proc. Natl Acad. Sci. USA 94, 4223–4228 (1997).
Ohad, N. et al. Mutations in FIE, a WD polycomb group gene, allow endosperm development without fertilization. Plant Cell 11, 10 (1999).
Kohler, C. et al. Arabidopsis MSI1 is a component of the MEA/FIE Polycomb group complex and required for seed development. EMBO J. 22, 4804–4814 (2003).
Luo, M. et al. Genes controlling fertilization-independent seed development in Arabidopsis thaliana. Proc. Natl Acad. Sci. USA 96, 296–301 (1999).
Kiyosue, T. et al. Control of fertilization-independent endosperm development by the MEDEA polycomb gene in Arabidopsis. Proc. Natl Acad. Sci. USA 96, 4186–4191 (1999).
Luo, M., Bilodeau, P., Dennis, E. S., Peacock, W. J. & Chaudhury, A. Expression and parent-of-origin effects for FIS2, MEA, and FIE in the endosperm and embryo of developing Arabidopsis seeds. Proc. Natl Acad. Sci. USA 97, 10637–10642 (2000).
Cheng, X. et al. Functional divergence of two duplicated Fertilization Independent Endosperm genes in rice with respect to seed development. Plant J. 104, 124–137 (2020).
Tonosaki, K. et al. Mutation of the imprinted gene OsEMF2a induces autonomous endosperm development and delayed cellularization in rice. Plant Cell 33, 85–103 (2021).
Figueiredo, D. D., Batista, R. A., Roszak, P. J. & Kohler, C. Auxin production couples endosperm development to fertilization. Nat. Plants 1, 15184 (2015).
Derkacheva, M. & Hennig, L. Variations on a theme: Polycomb group proteins in plants. J. Exp. Bot. 65, 2769–2784 (2014).
Hsieh, T.-F. et al. Regulation of imprinted gene expression in Arabidopsis endosperm. Proc. Natl Acad. Sci. USA 108, 1755–1762 (2011).
Wolff, P. et al. High-resolution analysis of parent-of-origin allelic expression in the Arabidopsis endosperm. PLoS Genet. 7, e1002126 (2011).
Weinhofer, I., Hehenberger, E., Roszak, P., Hennig, L. & Kohler, C. H3K27me3 profiling of the endosperm implies exclusion of polycomb group protein targeting by DNA methylation. PLoS Genet. 6, e1001152 (2010).
Moreno-Romero, J., Jiang, H., Santos-Gonzalez, J. & Kohler, C. Parental epigenetic asymmetry of PRC2-mediated histone modifications in the Arabidopsis endosperm. EMBO J. 35, 1298–1311 (2016).
Zhang, M. et al. Genome-wide high resolution parental-specific DNA and histone methylation maps uncover patterns of imprinting regulation in maize. Genome Res. 24, 167–176 (2014).
Dong, X. et al. Dynamic and antagonistic allele-specific epigenetic modifications controlling the expression of imprinted genes in maize endosperm. Mol. Plant 10, 442–455 (2017).
Figueiredo, D. D. & Köhler, C. Auxin: a molecular trigger of seed development. Genes Dev. 32, 13 479–490 (2018).
Luo, M., Platten, D., Chaudhury, A., Peacock, W. J. & Dennis, E. S. Expression, imprinting, and evolution of rice homologs of the polycomb group genes. Mol. Plant 2, 711–723 (2009).
Nallamilli, B. R. R. et al. Polycomb group gene OsFIE2 regulates rice (Oryza sativa) seed development and grain filling via a mechanism distinct from Arabidopsis. PLoS Genet. 9, e1003322 (2013).
Li, S. et al. OsFIE2 plays an essential role in the regulation of rice vegetative and reproductive development. New Phytol. 201, 66–79 (2014).
Maruyama, D. et al. Rapid elimination of the persistent synergid through a cell fusion mechanism. Cell 161, 907–918 (2015).
You, L. et al. Identification and analysis of genes involved in double fertilization in rice. Int. J. Mol. Sci. 22, 12850 (2021).
Dolezel, J., Greilhuber, J. & Suda, J. Estimation of nuclear DNA content in plants using flow cytometry. Nat. Protoc. 2, 2233–2244 (2007).
Cousin, A., Heel, K., Cowling, W. A. & Nelson, M. N. An efficient high-throughput flow cytometric method for estimating DNA ploidy level in plants. Cytometry A 75, 1015–1019 (2009).
Luo, M. et al. A genome-wide survey of imprinted genes in rice seeds reveals imprinting primarily occurs in the endosperm. PLoS Genet. 7, e1002125 (2011).
Anderson, S. N. et al. Transcriptomes of isolated Oryza sativa gametes characterized by deep sequencing: evidence for distinct sex-dependent chromatin and epigenetic states before fertilization. Plant J. 76, 729–741 (2013).
Indoliya, Y. et al. Decoding regulatory landscape of somatic embryogenesis reveals differential regulatory networks between japonica and indica rice subspecies. Sci. Rep. 6, 23050 (2016).
Sato, Y. et al. A rice homeobox gene, OSH1, is expressed before organ differentiation in a specific region during early embryogenesis. Proc. Natl Acad. Sci. USA 93, 8117–8122 (1996).
Sentoku, N. et al. Regional expression of the rice KN1-type homeobox gene family during embryo, shoot, and flower development. Plant Cell 11, 1651–1663 (1999).
Sato, Y., Sentoku, N., Nagato, Y. & Matsuoka, M. Isolation and characterization of a rice homebox gene, OSH15. Plant Mol. Biol. 38, 983–998 (1998).
Ito, M. et al. Position dependent expression of GL2-type homeobox gene, Roc1: significance for protoderm differentiation and radial pattern formation in early rice embryogenesis. Plant J. 29, 497–507 (2002).
Huang, X., Peng, X. & Sun, M.-X. OsGCD1 is essential for rice fertility and required for embryo dorsal-ventral pattern formation and endosperm development. New Phytol. 215, 1039–1058 (2017).
Yi, J. et al. OsMPK6 plays a critical role in cell differentiation during early embryogenesis in Oryza sativa. J. Exp. Bot. 67, 2425–2437 (2016).
Kamiya, N. et al. Rice globular embryo 4 (gle4) mutant is defective in radial pattern formation during embryogenesis. Plant Cell Physiol. 44, 875–883 (2003).
Ito, Y., Eiguchi, M. & Kurata, N. Expression of novel homeobox genes in early embryogenesis in rice. Biochim. Biophys. Acta 1444, 445–450 (1999).
Horst, N. A. et al. A single homeobox gene triggers phase transition, embryogenesis and asexual reproduction. Nat. Plants 2, 15209 (2016).
Yao, L. et al. OsMATL mutation induces haploid seed formation in indica rice. Nat. Plants 4, 530–533 (2018).
Conner, J. A., Podio, M. & Ozias-Akins, P. Haploid embryo production in rice and maize induced by PsASGR-BBML transgenes. Plant Reprod. 30, 41–52 (2017).
Cheng, X. et al. The maternally expressed polycomb group gene OsEMF2a is essential for endosperm cellularization and imprinting in rice. Plant Commun. 2, 100092 (2021).
Gehring, M., Missirian, V. & Henikoff, S. Genomic analysis of parent-of-origin allelic expression in Arabidopsis thaliana seeds. PLoS ONE 6, e23687 (2011).
Du, M., Luo, M., Zhang, R., Finnegan, E. J. & Koltunow, A. M. G. Imprinting in rice: the role of DNA and histone methylation in modulating parent-of-origin specific expression and determining transcript start sites. Plant J. 79, 232–242 (2014).
Waters, A. J. et al. Comprehensive analysis of imprinted genes in maize reveals allelic variation for imprinting and limited conservation with other species. Proc. Natl Acad. Sci. USA 110, 19639–19644 (2013).
Hater, F., Nakel, T. & Gross-Hardt, R. Reproductive multitasking: the female gametophyte. Annu. Rev. Plant Biol. 71, 517–546 (2020).
Nowack, M. K. et al. A positive signal from the fertilization of the egg cell sets off endosperm proliferation in angiosperm embryogenesis. Nat. Genet. 38, 63–67 (2006).
Guitton, A.-E. & Berger, F. Loss of function of MULTICOPY SUPPRESSOR OF IRA 1 produces nonviable parthenogenetic embryos in Arabidopsis. Curr. Biol. 15, 750–754 (2005).
Mosquna, A. et al. Regulation of stem cell maintenance by the polycomb protein FIE has been conserved during land plant evolution. Development 136, 2433–2444 (2009).
Okano, Y. et al. A polycomb repressive complex 2 gene regulates apogamy and gives evolutionary insights into early land plant evolution. Proc. Natl Acad. Sci. USA 106, 16321–16326 (2009).
Boutilier, K. et al. Ectopic expression of BABY BOOM triggers a conversion from vegetative to embryonic growth. Plant Cell 14, 1737–1749 (2002).
Lowe, K. et al. Morphogenic regulators Baby boom and Wuschel improve monocot transformation. Plant Cell 28, 1998–2015 (2016).
Zhang, T.-Q. et al. A two-step model for de novo activation of WUSCHEL during plant shoot regeneration. Plant Cell 29, 1073–1087 (2017).
Dunwell, J. M. Haploids in flowering plants: origins and exploitation. Plant Biotechnol. J. 8, 377–424 (2010).
Hand, M. L. & Koltunow, A. M. G. The genetic control of apomixis: asexual seed formation. Genetics 197, 441–450 (2014).
Ozias-Akins, P. & van Dijk, P. J. Mendelian genetics of apomixis in plants. Annu. Rev. Genet. 41, 509–537 (2007).
Khush, G. S. (ed.) Apomixis: Exploiting Hybrid Vigor in Rice (International Rice Research Institute, 1994).
Wang, K. Fixation of hybrid vigor in rice: synthetic apomixis generated by genome editing. aBIOTECH 1, 15–20 (2019).
Ma, X. et al. A robust CRISPR/Cas9 system for convenient, high-efficiency multiplex genome editing in monocot and dicot plants. Mol. Plant 8, 1274–1284 (2015).
Hiei, Y. & Komari, T. Agrobacterium-mediated transformation of rice using immature embryos or calli induced from mature seed. Nat. Protoc. 3, 824–834 (2008).
Wu, X., Liu, J., Li, D. & Liu, C.-M. Rice caryopsis development II: dynamic changes in the endosperm. J. Integr. Plant Biol. 58, 786–798 (2016).
Zeng, Y.-X., Hu, C.-Y., Lu, Y.-G., Li, J.-Q. & Liu, X.-D. Abnormalities occurring during female gametophyte development result in the diversity of abnormal embryo sacs and leads to abnormal fertilization in indica/japonica hybrids in rice. J. Integr. Plant Biol. 51, 3–12 (2009).
Morrison, W. R. & Laignelet, B. An improved colorimetric procedure for determining apparent and total amylose in cereal and other starches. J. Cereal Sci. 1, 9–20 (1983).
Galbraith, D. W. et al. Rapid flow cytometric analysis of the cell cycle in intact plant tissues. Science 220, 1049–1051 (1983).
Kawahara, Y. et al. Improvement of the Oryza sativa Nipponbare reference genome using next generation sequence and optical map data. Rice 6, 4 (2013).
Kim, D., Paggi, J. M., Park, C., Bennett, C. & Salzberg, S. L. Graph-based genome alignment and genotyping with HISAT2 and HISAT-genotype. Nat. Biotechnol. 37, 907–915 (2019).
Danecek, P. et al. Twelve years of SAMtools and BCFtools. Gigascience 10, giab008 (2021).
Robinson, J. T., Thorvaldsdottir, H., Wenger, A. M., Zehir, A. & Mesirov, J. P. Variant review with the Integrative Genomics Viewer. Cancer Res. 77, e31–e34 (2017).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Chen, C. et al. TBtools: an integrative toolkit developed for interactive analyses of big biological data. Mol. Plant 13, 1194–1202 (2020).
Ren, Y. et al. Majorbio Cloud: a one-stop, comprehensive bioinformatic platform for multiomics analyses. iMeta https://doi.org/10.1002/imt2.12 (2022).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Maere, S., Heymans, K. & Kuiper, M. BiNGO: a Cytoscape plugin to assess overrepresentation of gene ontology categories in biological networks. Bioinformatics 21, 3448–3449 (2005).
Shannon, P. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504 (2003).
Acknowledgements
We thank R. White for assistance with microscopy, C. Miller and L. Li for DNA flow cytometry; T. Loan and L. Dennis for critical reading of the manuscript. This work was funded by a CSIRO-Chinese Academy of Sciences Collaborative Fund to M.L. and Xiaoba Wu at CSIRO, the CAS strategic Priority Research programme XDB27030201 to F.L., B.Z. and X.C., the Sichuan Science and Technology fund 2022YFH0029 to H.Z. and Xianjun Wu, and a visiting scholarship from China Scholarship Council to L.X.
Funding
Open access funding provided by CSIRO Library Services.
Author information
Authors and Affiliations
Contributions
M.L. proposed the project. M.L., Xiaoba Wu and L.X. designed the study. L.X., Xiaoba Wu, X.S., N.W., H.Z., P.H., F.L., L.M., B.Z. and J.Y. performed all experiments. M.L., Xiaoba Wu, L.X., B.Z., J.F., C.H., A.C., L.M., Xianjun Wu and X.C. analysed the data. M.L., X.W. and L.X. wrote and revised the paper. All authors commented, discussed and provided input on the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Plants thanks Venkatesan Sundaresan and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 De-husked seeds of WT and self-pollinated Osfie mutants.
a, Seeds from Nipponbare (WT). b–d, Seeds from Osfie1-single1, 2, and 3 homozygotes. e–g, Seeds from Osfie2-single1, 2, and 3 heterozygotes. i–k, Seeds from Osfie-double1, 2, 3 and 4 heterozygotes (Bar = 0.5mm for all figures).
Extended Data Fig. 2 Aborted embryos and endosperm in Osfie double mutants under self-pollination or emasculation.
a–d, Self-pollinated Osfie-double1 at maturity; n = 137 aborted seeds dissected. e–h, Self-pollinated Osfie-double2 at maturity; n = 126 aborted seeds dissected. i–l, Self-pollinated Osfie-double3 at maturity; n = 23 aborted seeds dissected. m–p, Self-pollinated Osfie-double4 at maturity; n = 27 aborted seeds dissected. q, Slightly elongated ovules with intact anthers from Osfie-double1 on the day for emasculation (n = 4). r–t, Asexual embryos and autonomous endosperm in Osfie-double1 at 15 DPE. Noting that endosperm is visually absent in t; n = 17 aborted seeds dissected. In all figures except q, the red arrowheads indicate embryos and yellow arrowheads indicate endosperm; In q, red arrowhead indicates ovary (ov); Scale bar = 0.5mm.
Extended Data Fig. 3 Asexual embryos and autonomous endosperm in different double mutants.
a, Embryo sacs of WT segregants of Osfie-double2 at 12 DPE, showing an egg (white arrowhead), and two central nuclei (cc) (n = 68). b, Asexual four celled pre-embryo (circled by white dashed line, with another two out of focus) with a possible degenerating synergid cell (circled by yellow dashed line) and autonomous endosperm nuclei (inserted) in Osfie-double2 at 0 DEP. em: four-celled asexual embryo (red arrowhead); sy: synergid cell; en: autonomous endosperm; ap: antipodal (n = 1). c, Asexual globular embryos (red arrowhead) and cellularized endosperm (circled by yellow dashed line) in Osfie-double2 at 7 DPE (n = 31 in c-f). d, Asexual globular embryos (red arrowhead) and endosperm (circled by yellow dashed line) in Osfie-double2 at 7 DPE. e, Asexual embryos (red arrowhead) and endosperm (circled by yellow dashed line) in Osfie-double2 at 7 DPE. f, Cellularized autonomous endosperm for the ovule in e. g & h, Elongated asexual embryos (g) and starchy endosperm (h) from ovules in Osfie-double2 at 12 DPE (n = 8 in g-j). i & j, Elongated asexual embryos (i) and starchy endosperm (j) from ovules in Osfie-double2 at 12 DPE. k, Embryo sac of an Osfie1 homozygous segregant of Osfie-double4 at 10 DPE, showing an egg (white arrowhead), and two central nuclei (cc) (n = 24). l, Autonomous endosperm in Osfie-double4 at 0 DPE (egg circled by red dashed line and endosperm nuclei marked by yellow asterisks) (n = 19). m, Autonomous development in Osfie-double4 at 5 DPE (red arrowhead for embryo and yellow circle for endosperm) (n = 5). n, Asexual embryo (red arrowhead) in Osfie-double4 at 12 DPE (n = 3). Endosperm is absent. o, Embryo sacs showing no autonomous development in double homozygous segregants with Osfie2-6 weak allele of Osfie-double3 at 9 DPE (n = 41); white arrowhead for egg; cc for central cell nuclei. p, Autonomous development in Osfie-double3 at 0 DPE (egg circled by red dots and endosperm nuclei marked by yellow asterisks) (n = 9). q, Autonomous development in Osfie-double3 at 4 DPE (red arrowhead for embryo and yellow dashed line circle for endosperm) (n = 8). r, Autonomous development in Osfie-double3 at 12 DPE (red arrowhead for embryo, yellow dashed line circle for endosperm, pink arrowhead for vascular structure) (n = 5). s, Embryo sacs showing an egg cell (white arrowhead) and central cell nuclei (cc) in Osfie1 heterozygous segregants of (WT X Osfie-double1)F1 at 12 DPE (n = 58). t, Embryo sacs showing autonomous development in double heterozygous segregants of (WT X Osfie-double1)F1 at 12 DPE (red arrowhead for embryo and yellow dashed line circle for endosperm) (n = 4). u, Embryo sacs showing an egg cell (white arrowhead), synergid (blue arrowhead) and central cell nuclei (cc) in double heterozygous segregants with Osfie2-6 weak allele of (WT X Osfie-double3)F1 at 12 DPE (n = 59). v & w, Embryo sacs showing autonomous development in double heterozygous segregants of (WT X Osfie-double3)F1 at 12 DPE (red arrowhead for embryo and endosperm in W) (n = 13). x, Embryo sacs showing an egg cell (white arrowhead) and central cell nuclei (cc) in Osfie1 heterozygous segregants of (WT X Osfie-double4)F1 at 12 DPE (n = 64). y & z, Embryo sacs showing autonomous development in double heterozygous segregants of (WT X Osfie-double4)F1 at 12 DPE (red arrowhead for embryo) (n = 26). Bar = 50 μm.
Extended Data Fig. 4 Emasculated ovules of WT, Osfie1, and Osfie2.
a, Embryo sacs in Nipponbare at 15 DPE, showing an egg (white arrowhead), and two central nuclei (cc) (n = 49). b, Embryo sacs in Osfie1-single1 at 15 DPE, showing an egg (white arrowhead), and two central nuclei (cc) (n = 7). c, Embryo sacs in Osfie1-single2 at 10 DPE, showing an egg (white arrowhead), and two central nuclei (cc) (n = 5). d, Embryo sacs in Osfie1-single3 at 12 DPE, showing an egg (white arrowhead), and two central nuclei (cc) (n = 24). e, WT-looking embryo sacs in Osfie2-single2 at 0 DEP, showing an egg (white arrowhead), two synergids (one in focus by blue arrowhead, sy), two central nuclei (cc), and antipodal cells (grey arrowhead, ap) (n = 14). f, Asexual pre-embryo like structures (red arrowhead) at 8 DPE and a likely egg (white arrowhead) in Osfie2-single2 (n = 2 in f and g). g, Asexual pre-embryo like structures (red arrowhead) at 8 DPE and a likely egg (white arrowhead) in Osfie2-single2. h, Embryo sacs of WT segregants in Osfie2-single2 at 12 DPE, showing an egg (white arrowhead), and two central nuclei (cc) (n = 78). i, Embryo sacs of emasculated Nipponbare 4 days after auxin treatment, showing an egg (white arrowhead), and two central nuclei (cc) (n = 11). j, Embryo sacs of emasculated fie1-single1 14 days after auxin treatment (n = 10). k, Embryo sacs of emasculated fie1-single1 7 days after auxin treatment, showing two central nuclei (cc) (n = 10). l, Embryo sacs of emasculated Nipponbare 7 days after auxin treatment; central cell (insert), showing an egg (white arrowhead), and two central nuclei (cc) (n = 11) Bar = 50 μm.
Extended Data Fig. 5 The flow cytometric DNA histograms showing the gating borders (bars above the Propidium iodide florescence intensity peak) and peaks of rice samples (olive and pink) and internal control Bellis prennis (green).
a, Nipponbare callus showing a 2C (olive) and 4C (pink) peaks. b, Callus from aborted embryo of a self-pollinated seed showing 1C (olive) and 2C (pink) peaks. c, Callus from asexual embryo showing 1C (olive) and 2C (pink) peaks.
Extended Data Fig. 6 Transcriptomic analysis of asexual embryo and endosperm of Osfie-double1.
a, Principal Component Analysis (PCA) analysis for the transcriptomic samples, asexEM (asexual embryo with triplicates), autoEN (autonomous endosperm with triplicates), EM (5 days sexual embryo), EN (5 days sexual endosperm), SE (Somatic embryo), egg, Zygote (zygote 9 hours post fertilization). b, Venn diagram showing overlapping of expressed genes in asexual (asex), sexual (sex) and somatic embryo (EM) transcriptomes. c, Expression of genes potentially involved in embryo pattern formation in various tissues, using transcriptome data (OSH1, OSH6, OSH15, OSH71, OsROC1, OsGCD1, OsMPK6, OsPNH1, OsSCR, OsHOS66, OsLEC1B, OsPLT2/OsBBM2, OsBLH1, OsBLH4 and OsMTL; VC: Vegetative cell). Data are presented as mean values ± s.d. of 3 (AsexEM, somaticEM1, somticEM2, Egg, zygote2.5h, zygote5h, zygote9h,) and 2 (SexEM) biological replicates.
Extended Data Fig. 7 Differential gene expression, and visualization of gene expression and H3K27me3 enrichment of type I MADS-box genes.
a, Volcano plot showing genome-wide differential gene expression between asexual (asexEM) and sexual embryo (sexEM) (Supplementary Table 4, using DESeq276 R package). X axis showing Log2 values of fold change. Y axis showing the statistical significance (-Log10(padj)) of the expression change and the padj was selected as 0.05. The grey dots represent genes with expression of no more than 4 times fold change. Red dots represent genes with 4 times higher expression (n = 5535), while blue dots represent 4 times lower expression in asexual embryo (n = 2817). b, Volcano plot showing genome-wide differential gene expression between autonomous (asexEn) and sexual endosperm (sexEN) (Supplementary Table 7; using DESeq276 R package). X axis showing Log2 values of fold change. Y axis showing the statistical significance (-Log10(padj)) of the expression change and the padj was selected as 0.05. The grey dots represent genes with expression of no more than 4 times fold change. Red dots represent genes with 4 times higher expression (n = 5887) while blue dots represent 4 times lower expression in autonomous endosperm (n = 2796). c–k, Type I MADS-box genes (OsMADS1, OsMADS7, OsMADS8, OsMADS17, OsMADS65, OsMADS71, OsMADS77, OsMADS87 and OsMADS89) with higher expression in autonomous endosperm than in sexual endosperm showing enrichment of H3K27me3 in sexual endosperm. Data are presented as mean values ± s.d. of 3 (AsexEN) and 2 (SexEN) biological replicates shown.
Extended Data Fig. 8 Chromatin immunoprecipitation assay with anti-body against H3K27me3 followed by deep sequencing (ChIP-seq) to identify H3K27me3 marked regions in the maternal or paternal genomes of endosperm from the reciprocal crosses between Nipponbare (Nip) and 9311.
a, Dot plot indicating the parental bias status of H3K27me3 -enriched peaks based on the parental SNP counts in each of the reciprocal crosses, Nipx9311 and 9311xNip. Each dot represents a peak. Zero indicates no bias. Negative indicates H3K27me3 enrichment biased to maternal alleles. Positive indicates H3K27me3 enrichment biased to paternal alleles. Nip: biased to Nip allele in both crosses (436 peaks); Paternal: biased to paternal parents in both crosses (94 peaks); Paternal@Nip: biased to Nip in 9311xNip but no bias in Nipx9311 (136 peaks); paternal@9311: biased to 9311 in Nipx9311 but no bias in 9311xNip (140 peaks); 9311: biased to 9311 in both crosses (547 peaks); non-bias: no significant bias in both crosses (cutoff < 2^0.5) (577 peaks); Maternal: biased to maternal parents in both crosses (3602 peaks); Maternal@9311: biased to 9311 in 9311xNip but no bias in Nipx9311 (1454 peaks); maternal@Nip: biased to Nip in Nipx9311 but no bias in 9311xNip (1283 peaks). b, qPCR confirmation of H3K27me3 enrichment at OsYUCCA11 locus using ChIP DNA prepared from hybrid endosperm harvested from the reciprocal crosses between Nip and 9311 six days after pollination. H3: ChIP performed with antibody against histone 3 in 9311xNip; NX9: ChIP performed with antibody against H3K27me3 in Nipx9311; 9XN: ChIP performed with antibody against H3K27me3 in 9311xNip; Input: input DNA in 9311xNip. Data are presented as mean values ± s.d. of 3 biological replicates. c, Sanger sequencing of the PCR products amplified on the above ChIP DNA and input DNA showing DNA enriched by antibody against histone 3 and input DNA have both parental SNPs, while the DNA enriched by antibody against H3K27me3 on the Nipx9311 or 9311XNip endosperm only gave the maternal SNPs. Asterisks indicate the SNPs. Similar experiments were performed for other PEGs and results are shown in Extended Data Fig. 9a. d, Transcription and H3K27me3 enrichment around 4 SNPs at OsYUCCA11 locus for the NipX9311 endosperm, and transcription for the autonomous endosperm. For gene expression, m represents maternal normalized reads; p represents paternal normalized reads. For H3K27me3 ChIP, m represents maternal normalized reads; p represents paternal normalized reads. For autonomous endosperm, expression is shown by the normalized reads. More examples of other PEGs are in Extended Data Fig. 9.
Extended Data Fig. 9 Sanger sequencing of PCR products amplified on H3K27me3 enriched DNA, and visualization of transcription and H3K27me3 enrichment for selected PEGs.
a, Sanger sequencing of the PCR products amplified on the ChIP DNA and input for selected PEGs, showing input DNA samples have both parental SNPs (asterisks), while the DNA samples enriched by antibody against H3K27me3 on the Nipx9311 or 9311XNip endosperm only gave the maternal SNPs (asterisks). b, Transcription and H3K27me3 enrichment around 4 SNPs at selective PEGs for the NipX9311 endosperm, and transcription for the autonomous endosperm. For gene expression, m represents maternal normalized reads; p represents paternal normalized reads. For H3K27me3 ChIP, m represents maternal normalized reads; p represents paternal normalized reads. For autonomous endosperm, expression is shown by the normalized reads. c, Venn diagram showing genes marked with maternal H3k27me3 overlapping with imprinted genes. d, Venn diagram showing genes marked with paternal H3k27me3 overlapping with imprinted genes.
Supplementary information
Supplementary Information
Supplementary Method for ChIP assay.
Supplementary Tables
Supplementary Tables 1–18.
Source data
Source Data Fig. 5a
. Data for frequency of asexual embryo and autonomous seed formation at different stages. Source Data Fig. 5g, Source Data Extended Data Fig. 6c, Source Data Extended Data Fig. 7c–k and Source Data Extended Data Fig. 8b. Transcriptomic data for relevant gene expression. Source Data Fig. 6a. Data for heat map showing expression levels of BBM1 and WUS family genes in asexual embryos and other tissues.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wu, X., Xie, L., Sun, X. et al. Mutation in Polycomb repressive complex 2 gene OsFIE2 promotes asexual embryo formation in rice. Nat. Plants 9, 1848–1861 (2023). https://doi.org/10.1038/s41477-023-01536-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41477-023-01536-4
This article is cited by
-
Polycomb repression of the asexual embryo
Nature Plants (2023)