Abstract
Global warming threatens crop production. G proteins mediate plant responses to multiple abiotic stresses. Here we identified a natural quantitative trait locus, TT2 (THERMOTOLERANCE 2), encoding a Gγ subunit, that confers thermotolerance in rice during both vegetative and reproductive growth without a yield penalty. A natural allele with loss of TT2 function was associated with greater retention of wax at high temperatures and increased thermotolerance. Mechanistically, we found that a transcription factor, SCT1 (Sensing Ca2+ Transcription factor 1), functions to decode Ca2+ through Ca2+-enhanced interaction with calmodulin and acts as a negative regulator of its target genes (for example, Wax Synthesis Regulatory 2 (OsWR2)). The calmodulin–SCT1 interaction was attenuated by reduced heat-triggered Ca2+ caused by disrupted TT2, thus explaining the observed heat-induced changes in wax content. Beyond establishing a bridge linking G protein, Ca2+ sensing and wax metabolism, our study illustrates innovative approaches for developing potentially yield-penalty-free thermotolerant crop varieties.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Intact RNA-seq datasets were deposited in the NCBI Gene Expression Omnibus (GEO) under accession number GSE168650. The Rice SNP-Seek Database is accessible at https://snp-seek.irri.org/; the EnsemblPlants database is accessible at http://plants.ensembl.org/index.html. Source data are provided with this paper.
References
Lobell, D. B., Schlenker, W. & Costa-Roberts, J. Climate trends and global crop production since 1980. Science 333, 616–620 (2011).
Zhao, C. et al. Temperature increase reduces global yields of major crops in four independent estimates. Proc. Natl Acad. Sci. USA 114, 9326–9331 (2017).
You, J. & Chan, Z. ROS regulation during abiotic stress responses in crop plants. Front. Plant Sci. 6, 1092 (2015).
Li, X.-M. et al. Natural alleles of a proteasome α2 subunit gene contribute to thermotolerance and adaptation of African rice. Nat. Genet. 47, 827–833 (2015).
Zhu, J.-K. Abiotic stress signaling and responses in plants. Cell 167, 313–324 (2016).
Zhang, J., Li, X.-M., Lin, H.-X. & Chong, K. Crop improvement through temperature resilience. Annu. Rev. Plant Biol. 70, 753–780 (2019).
Pandey, S. Heterotrimeric G-protein signaling in plants: conserved and novel mechanisms. Annu. Rev. Plant Biol. 70, 213–238 (2019).
Urano, D., Chen, J.-G., Botella, J. R. & Jones, A. M. Heterotrimeric G protein signalling in the plant kingdom. Open Biol. 3, 120186 (2013).
Ma, Y. et al. COLD1 confers chilling tolerance in rice. Cell 160, 1209–1221 (2015).
Ferrero-Serrano, Á. & Assmann, S. M. The α-subunit of the rice heterotrimeric G protein, RGA1, regulates drought tolerance during the vegetative phase in the dwarf rice mutant d1. J. Exp. Bot. 67, 3433–3443 (2016).
Sun, S. et al. A G-protein pathway determines grain size in rice. Nat. Commun. 9, 851 (2018).
Kunihiro, S. et al. Rice DEP1, encoding a highly cysteine-rich G protein γ subunit, confers cadmium tolerance on yeast cells and plants. J. Exp. Bot. 64, 4517–4527 (2013).
Liu, Q. et al. G-protein βγ subunits determine grain size through interaction with MADS-domain transcription factors in rice. Nat. Commun. 9, 852 (2018).
Sun, H. et al. Heterotrimeric G proteins regulate nitrogen-use efficiency in rice. Nat. Genet. 46, 652–656 (2014).
Swain, D. M. et al. Function of heterotrimeric G-protein γ subunit RGG1 in providing salinity stress tolerance in rice by elevating detoxification of ROS. Planta 245, 367–383 (2017).
Wang, L. & Chong, K. in Integrated G Proteins Signaling in Plants (eds Yalovsky, S., et al.) 1–25 (Springer, 2010).
Herlitze, S. et al. Modulation of Ca2+ channels by G-protein beta gamma subunits. Nature 380, 258–262 (1996).
Yatani, A. et al. A G protein directly regulates mammalian cardiac calcium channels. Science 238, 1288–1292 (1987).
Gelli, A., Higgins, V. J. & Blumwald, E. Activation of plant plasma membrane Ca2+-permeable channels by race-specific fungal elicitors. Plant Physiol. 113, 269–279 (1997).
Chen, Y.-L. et al. Extracellular calmodulin-induced stomatal closure is mediated by heterotrimeric G protein and H2O2. Plant Physiol. 136, 4096–4103 (2004).
Jeon, B. W., Acharya, B. R. & Assmann, S. M. The Arabidopsis heterotrimeric G-protein β subunit, AGB1, is required for guard cell calcium sensing and calcium-induced calcium release. Plant J. 99, 231–244 (2019).
Wu, Y. et al. Heterotrimeric G-protein participation in Arabidopsis pollen germination through modulation of a plasmamembrane hyperpolarization-activated Ca2+-permeable channel. New Phytol. 176, 550–559 (2007).
Hao, L.-H. et al. Extracellular ATP promotes stomatal opening of Arabidopsis thaliana through heterotrimeric G protein α subunit and reactive oxygen species. Mol. Plant 5, 852–864 (2012).
Zhang, W., Jeon, B. W. & Assmann, S. M. Heterotrimeric G-protein regulation of ROS signalling and calcium currents in Arabidopsis guard cells. J. Exp. Bot. 62, 2371–2379 (2011).
Finka, A., Cuendet, A. F. H., Maathuis, F. J. M., Saidi, Y. & Goloubinoff, P. Plasma membrane cyclic nucleotide gated calcium channels control land plant thermal sensing and acquired thermotolerance. Plant Cell 24, 3333–3348 (2012).
Saidi, Y. et al. The heat shock response in moss plants is regulated by specific calcium-permeable channels in the plasma membrane. Plant Cell 21, 2829–2843 (2009).
Du, L., Yang, T., Puthanveettil, S. V. & Poovaiah, B. W. in Coding and Decoding of Calcium Signals in Plants (ed. Luan, S.) 177–233 (Springer, 2011).
Reddy, A. S. N., Ali, G. S., Celesnik, H. & Day, I. S. Coping with stresses: roles of calcium- and calcium/calmodulin-regulated gene expression. Plant Cell 23, 2010–2032 (2011).
Doherty, C. J., van Buskirk, H. A., Myers, S. J. & Thomashow, M. F. Roles for Arabidopsis CAMTA transcription factors in cold-regulated gene expression and freezing tolerance. Plant Cell 21, 972–984 (2009).
Kidokoro, S. et al. Different cold-signaling pathways function in the responses to rapid and gradual decreases in temperature. Plant Cell 29, 760–774 (2017).
Xie, Y., Shen, R., Chen, L. & Liu, Y.-G. Molecular mechanisms of hybrid sterility in rice. Sci. China Life Sci. 62, 737–743 (2019).
Shepherd, T. & Wynne Griffiths, D. The effects of stress on plant cuticular waxes. New Phytol. 171, 469–499 (2006).
Lewandowska, M., Keyl, A. & Feussner, I. Wax biosynthesis in response to danger: its regulation upon abiotic and biotic stress. New Phytol. 227, 698–713 (2020).
Zhou, X. et al. Overexpression of transcription factor OsWR2 regulates wax and cutin biosynthesis in rice and enhances its tolerance to water deficit. Plant Mol. Biol. Rep. 32, 719–731 (2014).
Tanaka, T., Tanaka, H., Machida, C., Watanabe, M. & Machida, Y. A new method for rapid visualization of defects in leaf cuticle reveals five intrinsic patterns of surface defects in Arabidopsis. Plant J. 37, 139–146 (2004).
Fan, C. et al. GS3, a major QTL for grain length and weight and minor QTL for grain width and thickness in rice, encodes a putative transmembrane protein. Theor. Appl. Genet. 112, 1164–1171 (2006).
Du, L. et al. Ca2+/calmodulin regulates salicylic-acid-mediated plant immunity. Nature 457, 1154–1158 (2009).
Yang, T. & Poovaiah, B. W. A calmodulin-binding/CGCG box DNA-binding protein family involved in multiple signaling pathways in plants. J. Biol. Chem. 277, 45049–45058 (2002).
Cui, Y. et al. CYCLIC NUCLEOTIDE-GATED ION CHANNELs 14 and 16 promote tolerance to heat and chilling in rice. Plant Physiol. 183, 1794–1808 (2020).
Clapham, D. E. Calcium signaling. Cell 131, 1047–1058 (2007).
Thor, K. et al. The calcium-permeable channel OSCA1.3 regulates plant stomatal immunity. Nature 585, 569–573 (2020).
Wang, J. et al. Transcriptional activation and phosphorylation of OsCNGC9 confer enhanced chilling tolerance in rice. Mol. Plant 14, 315–329 (2021).
Krebs, M. et al. FRET-based genetically encoded sensors allow high-resolution live cell imaging of Ca2+ dynamics. Plant J. 69, 181–192 (2012).
Ma, L. et al. The SOS2-SCaBP8 complex generates and fine-tunes an AtANN4-dependent calcium signature under salt stress. Dev. Cell 48, 697–709.e5 (2019).
Choi, M. S. et al. Isolation of a calmodulin-binding transcription factor from rice (Oryza sativa L.). J. Biol. Chem. 280, 40820–40831 (2005).
Pan, Y. et al. Dynamic interactions of plant CNGC subunits and calmodulins drive oscillatory Ca2+ channel activities. Dev. Cell 48, 710–725.e5 (2019).
Haiech, J. et al. Restoration of the calcium binding activity of mutant calmodulins toward normal by the presence of a calmodulin binding structure. J. Biol. Chem. 266, 3427–3431 (1991).
Hayes, S., Schachtschabel, J., Mishkind, M., Munnik, T. & Arisz, S. A. Hot topic: thermosensing in plants. Plant Cell Environ. 44, 2018–2033 (2021).
DeFalco, ThomasA., Bender, KyleW. & Snedden, WayneA. Breaking the code: Ca2+ sensors in plant signalling. Biochem. J. 425, 27–40 (2009).
Zhang, W. et al. Molecular and genetic evidence for the key role of AtCaM3 in heat-shock signal transduction in Arabidopsis. Plant Physiol. 149, 1773–1784 (2009).
Chakravorty, D. et al. An atypical heterotrimeric G-protein γ-subunit is involved in guard cell K+-channel regulation and morphological development in Arabidopsis thaliana. Plant J. 67, 840–851 (2011).
Szymanski, J., Brotman, Y., Willmitzer, L. & Cuadros-Inostroza, Á. Linking gene expression and membrane lipid composition of Arabidopsis. Plant Cell 26, 915–928 (2014).
Sadok, W., Lopez, J. R. & Smith, K. P. Transpiration increases under high-temperature stress: potential mechanisms, trade-offs and prospects for crop resilience in a warming world. Plant Cell Environ. https://doi.org/10.1111/pce.13970 (2020).
Liu, N., Karunakaran, C., Lahlali, R., Warkentin, T. & Bueckert, R. A. Genotypic and heat stress effects on leaf cuticles of field pea using ATR-FTIR spectroscopy. Planta 249, 601–613 (2019).
Wang, Z. et al. The E3 ligase DROUGHT HYPERSENSITIVE negatively regulates cuticular wax biosynthesis by promoting the degradation of transcription factor ROC4 in rice. Plant Cell 30, 228–244 (2018).
Chen, X. et al. Arabidopsis HSP70-16 is required for flower opening under normal or mild heat stress temperatures. Plant Cell Environ. 42, 1190–1204 (2019).
Chen, J.-H. et al. Nuclear-encoded synthesis of the D1 subunit of photosystem II increases photosynthetic efficiency and crop yield. Nat. Plants 6, 570–580 (2020).
Shen, H. et al. Overexpression of receptor-like kinase ERECTA improves thermotolerance in rice and tomato. Nat. Biotechnol. 33, 996–1003 (2015).
Wang, D. et al. Nucleolar DEAD-box RNA helicase TOGR1 regulates thermotolerant growth as a pre-rRNA chaperone in rice. PLoS Genet. 12, e1005844 (2016).
Mickelbart, M. V., Hasegawa, P. M. & Bailey-Serres, J. Genetic mechanisms of abiotic stress tolerance that translate to crop yield stability. Nat. Rev. Genet. 16, 237–251 (2015).
Wang, W. et al. Genomic variation in 3,010 diverse accessions of Asian cultivated rice. Nature 557, 43–49 (2018).
Hasanuzzaman, M., Nahar, K., Alam, M. M., Roychowdhury, R. & Fujita, M. Physiological, biochemical, and molecular mechanisms of heat stress tolerance in plants. Int. J. Mol. Sci. 14, 9643–9684 (2013).
Nishimura, A., Aichi, I. & Matsuoka, M. A protocol for Agrobacterium-mediated transformation in rice. Nat. Protoc. 1, 2796–2802 (2006).
Mortazavi, A., Williams, B. A., McCue, K., Schaeffer, L. & Wold, B. Mapping and quantifying mammalian transcriptomes by RNA-seq. Nat. Methods 5, 621–628 (2008).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Bray, N. L., Pimentel, H., Melsted, P. & Pachter, L. Near-optimal probabilistic RNA-seq quantification. Nat. Biotechnol. 34, 525–527 (2016).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Wang, Y. et al. An ethylene response factor OsWR1 responsive to drought stress transcriptionally activates wax synthesis related genes and increases wax production in rice. Plant Mol. Biol. 78, 275–288 (2012).
Zhang, H. et al. Metabolomics reveals how cucumber (Cucumis sativus) reprograms metabolites to cope with silver ions and silver nanoparticle-induced oxidative stress. Environ. Sci. Technol. 52, 8016–8026 (2018).
Bartlett, A. et al. Mapping genome-wide transcription-factor binding sites using DAP-seq. Nat. Protoc. 12, 1659–1672 (2017).
Li, W. et al. A natural allele of a transcription factor in rice confers broad-spectrum blast resistance. Cell 170, 114–126.e15 (2017).
Cui, L.-G., Shan, J.-X., Shi, M., Gao, J.-P. & Lin, H.-X. DCA1 acts as a transcriptional co-activator of DST and contributes to drought and salt tolerance in rice. PLoS Genet. 11, e1005617 (2015).
Acknowledgements
We thank Dr G. Zhang (South China Agricultural University) for providing CSSL plant materials; Drs J. Kudla (University of Heidelberg, Germany), K. Chong and X. Guo (Institute of Botany, CAS, China) for providing YC3.6 vectors and technical support; Drs Y. Guo and L. Ma (China Agriculture University, China) for providing aequorin vectors and technical support; Drs X. Li, M. Shi, W. Cai, S. Yin, Z. Zhang, X. Gao, J. Li and L. Xu (CAS Centre for Excellence in Molecular Plant Sciences, Shanghai Institute of Plant Physiology and Ecology, CAS) for technical support; Dr Y. Liu (South China Agriculture University) for donating CRISPR-Cas9 plasmids. This work was supported by grants from the National Natural Science Foundation of China (31788103, 31630052), the Chinese Academy of Sciences (XDB27010104, QYZDY-SSW-SMC023, 159231KYSB20200008), the Laboratory of Lingnan Modern Agriculture Project (NT2021002), CAS–Croucher Funding Scheme for Joint Laboratories and National Key Laboratory of Plant Molecular Genetics to H.-X.L.
Author information
Authors and Affiliations
Contributions
H.-X.L. conceived and supervised the project; H.-X.L. and Y.K. designed the experiments; Y.K. performed most of the experiments; X.-R.M., H.Z., J.G., W.-W.Y., J.-X.S. and H.-X.L. performed some of the experiments; Y.K. and H.-X.L. analysed data and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review information
Nature Plants thanks Scott Hayes, Julin Maloof and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
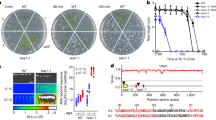
Extended Data Fig. 1 Thermotolerance phenotypes of CSSLs and various cultivars.
a, Phenotypic response to heat treatment of CSSL (HPS32) and huajingxian (HJX) as a negative control. Scale bars, 4 cm. b, The survival rates of CSSLs and the recurrent parent (HJX). Values are shown as mean ± s.d. (n = 3 samples, each sample represents 24 individual plants). P values according to two-tailed Student’s t-tests are shown. c, Analysis of the sequence identity of the HPS32 substitution segment across different cultivated accessions (chr3:16662761–17212811, 550 kb). Data were acquired from the genomes of 3010 Asian cultivated accessions (Rice SNP-Seek Database, https://snp-seek.irri.org/). We used the frequency of each SNP to evaluate the possible origination of the HPS32 target segment among different subpopulations. d, Distribution of the crucial SNP (C165A) in different subpopulations of rice. The allele frequencies in different subpopulations were calculated to make this histogram. e, Phenotypic response to heat treatment of 9 tropical japonica varieties (GP5, 7, 86, 112, 536, 629, 640, 641, and 642). Scale bars, 4 cm. f, Phenotypic response to heat treatment of six temperate japonica varieties (WYJ: Wuyunjing 7; KY131: Kongyu131; ZH11: Zhonghua11; Jiahua1; NIP: Nipponpare; and Xiushui09). Scale bars, 4 cm.
Extended Data Fig. 2 Molecular identification and phenotypic analysis of TT2 transgenic lines.
a, Schematic map of the sgRNA target sites in TT2, and a schematic showing vector construction and sequence alignment of the TT2-knockout transgenic lines with wild-type HJX. b, Relative expression level of TT2 in NIL-TT2HPS32 and pUbi:TT2HJX/NIL-TT2HPS32. P = 5.28 × 10−10 by one-way ANOVA (n = 6). c, Grain length in HJX, TT2KO/HJX, NIL-TT2HPS32, pTT2-gTT2HJX/NIL-TT2HPS32, and pUbi-TT2HJX/NIL-TT2HPS32. Scale bar, 1 cm. d, Survival rate of TT2 transgenic lines. The survival rate was determined after heat treatment (42 °C, RH > 90%) for 24 hr and subsequent recovery at 28 °C for 7 days. For box-and-whisker plots, the central line, box and whiskers indicate the median, IQR and 1.5 times the IQR, respectively. n = 6 samples, 24 individual plants per sample. TT2KO/HJX, P = 1.00 × 10−27; pUbi-TT2HJX/NIL-TT2HPS32, P = 1.09 × 10−24; pTT2-gTT2HJX/NIL-TT2HPS32, P = 2.58 × 10−32 by one-way ANOVA. The same letters indicate no significant difference at P > 0.05 as determined by Tukey’s HSD test (b, d).
Extended Data Fig. 3 Agronomic traits of NILs.
a, Pollen fertility of NILs under heat stress. Pollen grains were stained with KI-I2. Scale bars, 1 mm (left); 1000 μm (right). b, NILs at early filling stage. Scale bars, 10 cm. c, Grain length (n = 15), grain width (n = 15), grain number per panicle (n = 24), panicle number per plant (n = 30), panicle length (n = 30), and plant height (n = 48) of NILs. For box-and-whisker plots, the central line, box and whiskers indicate the median, IQR and 1.5 times the IQR, respectively. P values according to two-tailed Student’s t-tests are shown (c).
Extended Data Fig. 4 Transcriptomic analysis of DEGs between the two NILs.
a, Venn diagram and volcano plot of DEGs between the NILs under different treatments. Criteria for defining differentially expressed genes (DEGs) are |log2(Fold Change)| > 0 and P value < 0.05, n = 3 samples. H, heat treatment; C, control treatment. b, Genes clustered by expression level Cluster analysis for the two NILs under normal conditions or heat treatment. FKPM value was adjusted by log2(fpkm+1). The adjusted FKPM of every gene in a subcluster is shown as a gray line. A blue line represents the mean adjusted FKPM of the subcluster. c, Results of KEGG enrichment analysis of subcluster 2 and subcluster 3 are shown. Padj, adjusted P value; n = 3 samples. d, Heat map showing the expression patterns of wax-related DEGs in the two NILs under heat treatment. P < 0.05, n = 3 samples. Each sample represents 24 individual plants (a-d). P value was obtained by wald test using a model based on the negative binomial distribution (a,d), P value in KEGG obtained based on hypergeometric distribution model was adjusted to Padj using the Benjamini & Hochberg method (c).
Extended Data Fig. 5 Characteristics of OsWR2-knockout transgenic lines in NIL-TT2HPS32 backgrounds.
a, Sequence alignments of OsWR2 in wr2/NIL-TT2HPS32 with their wild-type counterparts. b, Toluidine blue (TB) staining assay for wr2/NIL-TT2HPS32 and NIL-TT2HPS32 after a 1-hr and 12-hr heat treatment. Heavier staining in leaves indicates higher permeability of the cuticle layer. Scale bars, 1 cm. Leaves are sampled from different individual plants. c, SEM images of cuticle wax crystal patterns on the surfaces of leaves in NIL-TT2HPS32 and wr2/NIL-TT2HPS32 plants under normal conditions. Images were obtained from three individual plants, respectively. Scale bars, 1 μm.
Extended Data Fig. 6 The homologs of the CaM-binding transcription factor SCT1.
a, The maximum-likelihood tree based on protein sequences of 42 SCT1 homologs. The amino acid sequences were obtained from a EnsemblPlants database search. The sequences were aligned using MEGA6.0. Scale bar represents 0.2 substitutions per site. Yellow, Teiticum aestivum; black, O. sativa; blue, Zea mays; red, Arabidopsis thaliana, green, Glycine max. b, Alignment of the DNA binding domain and CaMBD sequences of SCT1 homologs. The sequences were aligned using MEGA6.0. Hydrophobic amino acid residues corresponding to the CaM motif are marked with red arrows. c, Electrophoretic mobility shift assay (EMSA) of SCT2-NT binding to CG1-like cis-elements. The competition assay was performed with unlabeled wild-type oligonucleotides in 5-, 20-, and 50-fold excess of the wild-type probes. Experiments were independently conducted for at least twice (c).
Extended Data Fig. 7 Molecular identification and RNA-profiling of the sct1/sct2 mutant.
a, Sequence alignments of SCT1/SCT2 in sct1/sct2 with their wild-type counterparts. b, Results of KEGG enrichment analysis of DEGs which was more highly expressed in sct1/sct2 than in wild-type plants under normal conditions. c, The expression pattern of TT1 in sct1/sct2 and ZH11 under normal conditions and heat treatment. TT1 was used as a negative control. Values are shown as the mean ± s.d., Pinteraction = 0.690 by two-way ANOVA, n = 3 samples. d, Thirty one wax-related marker genes were differentially expressed in sct1/sct2 mutant plants as compared with ZH11 under normal conditions and heat treatment, based on RNA-seq profiling (n = 3 samples). Each sample represents 24 individual plants (b-d).
Extended Data Fig. 8 Calcium imaging based on NES-YC3.6 and aequorin assays.
a, The ratiometric image (cpVenus/ECFP) in living roots of NIL-TT2HJX and NIL-TT2HPS32 based on NES-YC3.6 before and after heat stimulation, scale bar, 20 μm. b, Images of heat-induced Ca2+ increases scaled by a pseudocolored reference in aequorin-expressing NIL-TT2HJX and NIL-TT2HPS32 seedlings treated with heat stress (left) and discharge buffer (right, recording total remaining luminescence).
Extended Data Fig. 9 CaM-SCT1 interaction is dependent on Ca2+.
a, Subcellular localization of SCT1, SCT2, and CaM in tobacco. Scale bars, 20 μm. b, Gel mobility shift assay with the CaMBD of SCT1. The amino acid sequences of the peptide correspond to amino acids 897–917 of SCT1. CaM was incubated with increasing amounts of the peptide (peptide: CaM molar ratios are indicated) in the presence of 0.1 mM CaCl2 or 2 mM EGTA. Samples were separated by non-denaturing PAGE and stained with Coomassie Brilliant Blue. In the mutant peptides, valine (Val900) and tryptophan (Trp907) were changed to arginine (Arg). c, Protein alignment of CaM1234 with CaM; four conserved Glu residues in the EF-hand motif were mutated to Arg. d, The TT1 mRNA levels under normal condition and heat treatment in coordination with normal, MgCl2, or LaCl3 conditions. P = 3.83 × 10−3 by one-way ANOVA (n = 3 to 4 biological replicates). e, Dual-luciferase assays performed upon blocking of Ca2+ channels by La3+ treatment in rice protoplasts. The expression of luciferase was driven by the 7xUAS reporter system. Mg2+ was used as a negative control. P = 2.31 × 10−10 by one-way ANOVA (n = 3 biological replicates). Values are shown as the mean ± s.d. One-way ANOVA being represented by Tukey’s HSD test using the same letters at P > 0.05 (d, e). Experiments were independently conducted for at least twice (a, b).
Extended Data Fig. 10 Calculation of KD value between SCT1 and CaM.
Three concentrations of CaM were used to calculate KD value between SCT1 and CaM fitted to one-to-one evaluation type at the presence of CaCl2 or EGTA. Three lines are fitted to one curve and the KD value was calculated. The signal after blocking the SCT1-CaMBD-immobilized chip was set to 0.
Supplementary information
Supplementary Information
Supplementary protocols.
Supplementary Tables 1–6
Detailed GSEA data, descriptions of wax/cutin-related genes, sequences of oligonucleotides (including primers, transgenic constructs, and so on), antibodies and strains.
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Unprocessed gels and statistical source data.
Source Data Fig. 5
Statistical source data.
Source Data Fig. 6
Unprocessed western blots and statistical source data.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 6
Unprocessed gels.
Source Data Extended Data Fig. 7
Statistical source data.
Source Data Extended Data Fig. 9
Unprocessed gels and statistical source data.
Source Data Extended Data Fig. 10
Statistical source data.
Rights and permissions
About this article
Cite this article
Kan, Y., Mu, XR., Zhang, H. et al. TT2 controls rice thermotolerance through SCT1-dependent alteration of wax biosynthesis. Nat. Plants 8, 53–67 (2022). https://doi.org/10.1038/s41477-021-01039-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41477-021-01039-0
This article is cited by
-
Agronomic potential of plant-specific Gγ proteins
Physiology and Molecular Biology of Plants (2024)
-
Utilization of natural alleles for heat adaptability QTLs at the flowering stage in rice
BMC Plant Biology (2023)
-
RGA1 Negatively Regulates Thermo-tolerance by Affecting Carbohydrate Metabolism and the Energy Supply in Rice
Rice (2023)
-
OsmiR159 Modulate BPH Resistance Through Regulating G-Protein γ Subunit GS3 Gene in Rice
Rice (2023)
-
Calcium signaling-mediated transcriptional reprogramming during abiotic stress response in plants
Theoretical and Applied Genetics (2023)