Abstract
Asymmetric cell division (ACD) requires protein polarization in the mother cell to produce daughter cells with distinct identities (cell-fate asymmetry). Here, we define a previously undocumented mechanism for establishing cell-fate asymmetry in Arabidopsis stomatal stem cells. In particular, we show that polarization of the protein phosphatase BSL1 promotes stomatal ACD by establishing kinase-based signalling asymmetry in the two daughter cells. BSL1 polarization in the stomatal ACD mother cell is triggered at the onset of mitosis. Polarized BSL1 is inherited by the differentiating daughter cell, where it suppresses cell division and promotes cell-fate determination. Plants lacking BSL proteins exhibit stomatal overproliferation, which demonstrates that the BSL family plays an essential role in stomatal development. Our findings establish that BSL1 polarization provides a spatiotemporal molecular switch that enables cell-fate asymmetry in stomatal ACD daughter cells. We propose that BSL1 polarization is triggered by an ACD checkpoint in the mother cell that monitors the establishment of division-plane asymmetry.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
All generated and analysed data from this study are included in the published article and its Supplementary Information. Source data are provided with this paper.
References
Knoblich, J. A. Mechanisms of asymmetric stem cell division. Cell 132, 583–597 (2008).
Abrash, E. B. & Bergmann, D. C. Asymmetric cell divisions: a view from plant development. Dev. Cell 16, 783–796 (2009).
Pierre-Jerome, E., Drapek, C. & Benfey, P. N. Regulation of division and differentiation of plant stem cells. Annu. Rev. Cell Dev. Biol. 34, 289–310 (2018).
Ruijtenberg, S. & van den Heuvel, S. Coordinating cell proliferation and differentiation: antagonism between cell cycle regulators and cell type-specific gene expression. Cell Cycle 15, 196–212 (2016).
Goldstein, B. & Macara, I. G. The PAR proteins: fundamental players in animal cell polarization. Dev. Cell 13, 609–622 (2007).
Sunchu, B. & Cabernard, C. Principles and mechanisms of asymmetric cell division. Development 147, dev167650 (2020).
Facette, M. R. & Smith, L. G. Division polarity in developing stomata. Curr. Opin. Plant Biol. 15, 585–592 (2012).
Pillitteri, L. J. & Torii, K. U. Mechanisms of stomatal development. Annu. Rev. Plant Biol. 63, 591–614 (2012).
Zoulias, N., Harrison, E. L., Casson, S. A. & Gray, J. E. Molecular control of stomatal development. Biochem. J. 475, 441–454 (2018).
Qi, X. & Torii, K. U. Hormonal and environmental signals guiding stomatal development. BMC Biol. 16, 21 (2018).
MacAlister, C. A., Ohashi-Ito, K. & Bergmann, D. C. Transcription factor control of asymmetric cell divisions that establish the stomatal lineage. Nature 445, 537–540 (2007).
Lau, O. S. et al. Direct roles of SPEECHLESS in the specification of stomatal self-renewing cells. Science 345, 1605–1609 (2014).
Dong, J., MacAlister, C. A. & Bergmann, D. C. BASL controls asymmetric cell division in Arabidopsis. Cell 137, 1320–1330 (2009).
Pillitteri, L. J., Peterson, K. M., Horst, R. J. & Torii, K. U. Molecular profiling of stomatal meristemoids reveals new component of asymmetric cell division and commonalities among stem cell populations in Arabidopsis. Plant Cell 23, 3260–3275 (2011).
Rowe, M. H., Dong, J., Weimer, A. K. & Bergmann, D. C. A plant-specific polarity module establishes cell fate asymmetry in the Arabidopsis stomatal lineage. Preprint at bioRxiv https://doi.org/10.1101/614636 (2019).
Guo, X. & Dong, J. To divide or differentiate: it is about scaffolding. Trends Plant Sci. 24, 481–484 (2019).
Bergmann, D. C., Lukowitz, W. & Somerville, C. R. Stomatal development and pattern controlled by a MAPKK kinase. Science 304, 1494–1497 (2004).
Zhang, Y., Wang, P., Shao, W., Zhu, J. K. & Dong, J. The BASL polarity protein controls a MAPK signaling feedback loop in asymmetric cell division. Dev. Cell 33, 136–149 (2015).
Houbaert, A. et al. POLAR-guided signalling complex assembly and localization drive asymmetric cell division. Nature 563, 574–578 (2018).
Lampard, G. R., Macalister, C. A. & Bergmann, D. C. Arabidopsis stomatal initiation is controlled by MAPK-mediated regulation of the bHLH SPEECHLESS. Science 322, 1113–1116 (2008).
Gudesblat, G. E. et al. SPEECHLESS integrates brassinosteroid and stomata signalling pathways. Nat. Cell Biol. 14, 548–554 (2012).
Kim, T. W., Michniewicz, M., Bergmann, D. C. & Wang, Z. Y. Brassinosteroid regulates stomatal development by GSK3-mediated inhibition of a MAPK pathway. Nature 482, 419–422 (2012).
Kim, T. W. et al. Brassinosteroid signal transduction from cell-surface receptor kinases to nuclear transcription factors. Nat. Cell Biol. 11, 1254–1260 (2009).
Mora-Garcia, S. et al. Nuclear protein phosphatases with Kelch-repeat domains modulate the response to brassinosteroids in Arabidopsis. Genes Dev. 18, 448–460 (2004).
Gutierrez, R., Lindeboom, J. J., Paredez, A. R., Emons, A. M. & Ehrhardt, D. W. Arabidopsis cortical microtubules position cellulose synthase delivery to the plasma membrane and interact with cellulose synthase trafficking compartments. Nat. Cell Biol. 11, 797–806 (2009).
Livanos, P. & Muller, S. Division plane establishment and cytokinesis. Annu. Rev. Plant Biol. 70, 239–267 (2019).
Nadeau, J. A. & Sack, F. D. Control of stomatal distribution on the Arabidopsis leaf surface. Science 296, 1697–1700 (2002).
Winter, D. et al. An ‘electronic fluorescent pictograph’ browser for exploring and analyzing large-scale biological data sets. PLoS ONE 2, e718 (2007).
Wang, H., Ngwenyama, N., Liu, Y., Walker, J. C. & Zhang, S. Stomatal development and patterning are regulated by environmentally responsive mitogen-activated protein kinases in Arabidopsis. Plant Cell 19, 63–73 (2007).
Shpak, E. D., McAbee, J. M., Pillitteri, L. J. & Torii, K. U. Stomatal patterning and differentiation by synergistic interactions of receptor kinases. Science 309, 290–293 (2005).
Meng, X. et al. Differential function of arabidopsis SERK family receptor-like kinases in stomatal patterning. Curr. Biol. 25, 2361–2372 (2015).
Shao, W. & Dong, J. Polarity in plant asymmetric cell division: division orientation and cell fate differentiation. Dev. Biol. 419, 121–131 (2016).
Han, S. K. & Torii, K. U. Lineage-specific stem cells, signals and asymmetries during stomatal development. Development 143, 1259–1270 (2016).
Gong, Y., Varnau, R., Wallner, E.-S., Bergmann, D. C. & Cheung, L. S. Quantitative and dynamic cell polarity tracking in plant cells. Preprint at bioRxiv https://doi.org/10.1101/2020.09.12.294942 (2020).
Zhang, Y., Guo, X. & Dong, J. Phosphorylation of the polarity protein BASL differentiates asymmetric cell fate through MAPKs and SPCH. Curr. Biol. 26, 2957–2965 (2016).
Park, C. H. et al. BSU1 family phosphatases mediate Flagellin–FLS2 signaling through a specific phosphocode. Preprint at bioRxiv https://doi.org/10.1101/685610 (2019).
Rasmussen, C. G., Humphries, J. A. & Smith, L. G. Determination of symmetric and asymmetric division planes in plant cells. Annu. Rev. Plant Biol. 62, 387–409 (2011).
Wang, H., Ouyang, Y., Somers, W. G., Chia, W. & Lu, B. Polo inhibits progenitor self-renewal and regulates Numb asymmetry by phosphorylating Pon. Nature 449, 96–100 (2007).
Witte, K., Strickland, D. & Glotzer, M. Cell cycle entry triggers a switch between two modes of Cdc42 activation during yeast polarization. eLife 6, e26722 (2017).
Moran, K. D. et al. Cell-cycle control of cell polarity in yeast. J. Cell Biol. 218, 171–189 (2019).
Reich, J. D. et al. Regulated activation of the PAR polarity network ensures a timely and specific response to spatial cues. Curr. Biol. 29, 1911–1923 (2019).
Van Damme, D. et al. Arabidopsis α Aurora kinases function in formative cell division plane orientation. Plant Cell 23, 4013–4024 (2011).
Vandepoele, K. et al. Genome-wide analysis of core cell cycle genes in Arabidopsis. Plant Cell 14, 903–916 (2002).
Boudolf, V. et al. B1-type cyclin-dependent kinases are essential for the formation of stomatal complexes in Arabidopsis thaliana. Plant Cell 16, 945–955 (2004).
Yang, K. et al. A conserved but plant-specific CDK-mediated regulation of DNA replication protein A2 in the precise control of stomatal terminal division. Proc. Natl Acad. Sci. USA 116, 18126–18131 (2019).
Spinner, L. et al. A protein phosphatase 2A complex spatially controls plant cell division. Nat. Commun. 4, 1863 (2013).
Schaefer, E. et al. The preprophase band of microtubules controls the robustness of division orientation in plants. Science 356, 186–189 (2017).
Lew, D. J. & Burke, D. J. The spindle assembly and spindle position checkpoints. Annu. Rev. Genet 37, 251–282 (2003).
Venkei, Z. G. & Yamashita, Y. M. The centrosome orientation checkpoint is germline stem cell specific and operates prior to the spindle assembly checkpoint in Drosophila testis. Development 142, 62–69 (2015).
Hartwell, L. H. & Weinert, T. A. Checkpoints: controls that ensure the order of cell cycle events. Science 246, 629–634 (1989).
Venkei, Z. G. & Yamashita, Y. M. Emerging mechanisms of asymmetric stem cell division. J. Cell Biol. 217, 3785–3795 (2018).
Lampard, G. R., Lukowitz, W., Ellis, B. E. & Bergmann, D. C. Novel and expanded roles for MAPK signaling in Arabidopsis stomatal cell fate revealed by cell type-specific manipulations. Plant Cell 21, 3506–3517 (2009).
Nakagawa, T. et al. Development of R4 gateway binary vectors (R4pGWB) enabling high-throughput promoter swapping for plant research. Biosci. Biotechnol. Biochem. 72, 624–629 (2008).
Kubo, M. et al. Transcription switches for protoxylem and metaxylem vessel formation. Genes Dev. 19, 1855–1860 (2005).
Craig, R., Cortens, J. P. & Beavis, R. C. Open source system for analyzing, validating, and storing protein identification data. J. Proteome Res. 3, 1234–1242 (2004).
Zhao, C. et al. MAP kinase cascades regulate the cold response by modulating ICE1 protein stability. Dev. Cell 43, 618–629 (2017).
Li, J. & Chory, J. A putative leucine-rich repeat receptor kinase involved in brassinosteroid signal transduction. Cell 90, 929–938 (1997).
Li, J. & Nam, K. H. Regulation of brassinosteroid signaling by a GSK3/SHAGGY-like kinase. Science 295, 1299–1301 (2002).
Yin, Y. et al. BES1 accumulates in the nucleus in response to brassinosteroids to regulate gene expression and promote stem elongation. Cell 109, 181–191 (2002).
Wang, Z. Y. et al. Nuclear-localized BZR1 mediates brassinosteroid-induced growth and feedback suppression of brassinosteroid biosynthesis. Dev. Cell 2, 505–513 (2002).
Acknowledgements
We thank the ABRC Stock Center for providing T-DNA insertional seeds, the Biological Mass Spectrometry Facility of Robert Wood Johnson Medical School for performing the MS analyses, and J. Winkelman for assistance with the data analyses. This research was supported by grants NIH GM109080/GM131827 and NSF 825885/1952823 to J.D., NIH GM066258 to Z.-Y.W., and NIH GM118059 to B.E.N.
Author information
Authors and Affiliations
Contributions
X.G. and J.D. designed the research. X.G. performed most of the experiments. C.H.P. assisted with the mutant analysis. X.G., C.H.P., Z.-Y.W., B.E.N. and J.D. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Plants thanks Laurie Smith and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Genetic Pathways Involving BSL Proteins in Arabidopsis.
a, Compositional and functional changes of the polarity complex before and after a stomatal ACD. In a young leaf, the initiation of the stomatal lineage cell (MMCs), is driven by the expression of the transcription factor SPEECHLESS (SPCH)11. The plant-specific protein BASL is polarized before, during and after a stomatal ACD13. Multiple components in the BASL polarity complex have been identified to regulate cellular events in stomatal ACD. The BRX proteins interact with BASL to attach the polarity complex to the plasma membrane15. The POLAR proteins associate with the polarity complex and require BASL to become polarized14. It is known that POLAR recruits the BIN2 GSK3-like kinases to the polarity complex in the MMC, where BIN2 phosphorylates and inhibits the MAPKKK YDA, leading to alleviated MAPK-mediated suppression of SPCH, thereby the MMC undergoes cell division19. It is also known that, BASL interacts with YDA to locally concentrate MAPK signaling, thus SPCH is suppressed in the large daughter cell SLGC that inherits the polarity complex, thereby the SLGC after an ACD undergoes differentiation to become a pavement cell18,35. Thus, a successful stomatal ACD necessitates the changes of the key regulators, that is BIN2 to be preferentially membrane-localized before ACD but nucleus-partitioned after ACD, whereas YDA to be suppressed before ACD but activated after ACD. Light blue, stomatal fate; dark blue, stomatal guard cells; pink, non-stomatal fate; light pink, pavement cell. Fading cells, PrCs not converted to MMC become non-stomatal pavement cells. Dotted rectangle, regions enlarged in bottom, containing protein components of the polarity complex. Bottom, enlarged view of polarity protein complexes required for the progenitor cell (MMC, light blue) and the daughter cell (SLGC, pink), respectively, in stomatal ACD. ?, unidentified regulator/s for POLAR to associate with the polarity complex. Fragmented molecules (SPCH and POLAR) indicate proteins undergo degradation. b, Schematics depicting simplified signal transduction pathways in stomatal development (left) and Brassinosteroid (BR) signaling (right). The models are mainly based on Arabidopsis research. In stomatal development, the receptor-like kinases/protein (TMM, TOO MANY MOUTHS; ERf, ERECTA family; SERKf, SOMATIC EMBRYOGENESIS RECEPTOR-LIKE KINASE family)27,30,31 transduce signals from the EPF peptide ligands (not specified here) to activate the YODA (YDA) MAPK signaling that suppresses the transcription factors (SPCH, SPEECHLESS; ICE1/SCRM, SCREAM) and stomatal development17,20,56. The BIN2 GSK-3 like kinases regulate stomatal development by inhibiting YDA and SPCH21,22. The polarity protein BASL interacts with YDA and forms a positive feedback regulation with the MAPK cascade to regulate stomatal ACD18. In this study, we identified the BSL protein phosphatases as BASL partners in the polarity complex. In BR signaling, the hormone (BR) is perceived by the receptor-like kinase (BRI1, BRASSINOSTEROID INSENSITIVE 1)57 that triggers cytoplasmic signal transduction mediated by the BSL protein phosphatases23 and the BIN2 GSK3-like kinases58 to ultimately active the expression of the transcription factors (BZR1, BES1)59,60 for plant responses. Both pathways involve BSL Ser/Thr protein phosphatases and the GSK3-like BIN2 kinase. Block lines indicate negative regulation and arrows indicate positive regulation. PM, plasma membrane; NE, nuclear envelope.
Extended Data Fig. 2 BSLf Physically Interact with BASL in Plants.
a-b, Co-IP assays using purified fusion proteins that are transiently overexpressed (driven by the ubiquitous 35S promoter) in N. benthamiana leaf cells. a, Results show physical association of FLAG-tagged BSL1/2/3 proteins with GFP-BASL. GFP-BASL (right) or GFP (left, control) was used as bait to bind to the GFP-Trap agarose. Immunoprecipitated proteins were detected by anti-FLAG. Data represent results of three biological repeats. b, Results show physical association of FLAG-tagged BSL1D584N (catalytically inactive BSL1 variant) proteins with GFP-BASL and BIN2-YFP. GFP alone (left, control), GFP-BASL (middle) or BIN2-YFP (right) was used as bait to bind to the GFP-Trap agarose. Immunoprecipitated proteins were detected by anti-FLAG. Data represent results of three biological repeats.
Extended Data Fig. 3 BSL1, BSL2 and BSL3 Co-localize with BASL in Arabidopsis.
a, qPCR data show relative expression levels of BSL1 (driven by the native promoter) in bsl1 (left) or basl-2 (right). Three independent transgenic lines were used. Total RNAs were extracted from 3-day-old seedlings. Gene expression levels were normalized by ACTIN2 and relative expression levels of BSL1 were compared with the values in the wild type. Data are presented as mean ± SD. n = three independent experiments. b, Co-expression of GFP-BASL (green) with BSL1-mRFP (magenta), both driven by the BASL promoter, in a true leaf of 5-dpg seedling. c, Representative images of co-localization of BSL1-mRFP, BSL2-mRFP and BSL3-mRFP (magenta) with GFP-BASL (green), all driven by the BASL promoter. Arrows mark protein polarization at the cell cortex. d, Expression patterns of endogenous promoter driven GFP-BASL (green) and BSL1-YFP (magenta) in progressive cell types during stomatal development. PrC (protodermal cells), early MMC (Meristemoid Mother Cell, small and rectangular), late MMC (asymmetrically expanded and triangle), M (Meristemoid), and SLGC (Stomatal Lineage Ground Cell). Protein polarizations were indicated by arrows. Scale bars in (b-d), 5 μm. e, qPCR data show relative expression levels of BASL and BSL in indicated genetic backgrounds. Total RNAs were extracted from 3-day-old seedlings. Gene expression levels were relative to ACTIN2. Data are presented as mean ± SD. n = three independent experiments. Primers used were listed in Supplementary Table 2.
Extended Data Fig. 4 Association of BSL1 with the Polarity Complex Promotes BIN2 Partitioning to the Nucleus.
a, qPCR data show relative expression levels of BIN2 (left) and BSL1/BSL1D584N (right) in the indicated genetic backgrounds. Total RNAs were extracted from 3-day-old seedlings. Gene expression levels were normalized by ACTIN2 and relative expression levels of BIN2 or BSL1 were compared with the values in the wild-type background. Data are presented as mean ± SD. n = three independent experiments. Primers used were listed in Supplementary Table 2. b, Representative confocal images show differential membrane/nuclear partitioning of BIN2 (yellow) in stomatal lineage cells from the wild type vs. in bsl-quad mutants vs. in BSL1 overexpression plants (driven by the TMM promoter). The expression of BIN2-YFP is driven by the native promoter. Right, florescence intensity profiling of the dotted lines drawn across the stomatal lineage cells shown on the left. Confocal images were taken with same settings and absolute fluorescence intensity values were obtained and profiled by Fiji. M, cell membrane; N, nucleus. Scale, 5 μm c, Co-expression of the TMM promoter driven BSL1-mRFP (red) with the endogenous promoter driven BIN2-YFP (yellow). The nuclear enrichment of BIN2 was verified by positive DAPI staining (cyan). Scale, 10 µm.
Extended Data Fig. 5 BSL1 Interferes the Interaction of BIN2-BASL in vitro.
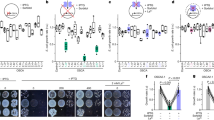
a, In vitro pull-down assays using recombinant proteins to test the BIN2-BASL interaction in the presence of an increasing amount of MBP-BSL1. His-BASL was used as bait and the amount of GST-BIN2 being pulled down reflects the interaction strength of BIN2-BASL. MBP was used as negative control. Histograms (below) show quantification of relative protein levels of BIN2 in the assay above. Results suggest the addition of MBP-BSL1 reduced the amount of BIN2 that interacted with BASL. Data are presented as mean ± SD. n = three independent experiments. Two-tailed Student’s t-tests. ** P < 0.005. b, Co-IP assays using purified fusion proteins produced by N. benthamiana leaves show physical association of BIN2-YFP with Myc-POLAR was not influenced by the presence of FLAG-tagged BSL1 or BSL1D584N. BIN2-YFP was used as bait. Immunoprecipitated proteins were detected by anti-Myc. Data represent results of experiments repeated three times. c, Results of yeast two-hybrid assays show no interactions between BSL1 and POLAR were identified. BD indicates Gal4 DNA-binding domain. AD indicates Gal4 activation domain. ‘Interaction tests’, assays performed using synthetic dropout medium (-Leu-Trp-His; 1 mM 3-AT added to suppress auto-activation); ‘growth controls’, assays performed using rich media (-Leu-Trp).
Extended Data Fig. 6 BSL1 Interferes the Interaction of BIN2-BASL in Planta.
a, BiFC assays show interactions between BIN2 and BASL (yellow) in N. benthamiana leaves. Half YFPs (nYFP and cYFP) were used as negative controls. Arrows indicate protein polarization in epidermal cells. b, BiFC interaction tests for BIN2-BASL (yellow) in the presence of BSL1-CFP (cyan, top) or BSL1D584N-CFP (cyan, bottom). Note, in the presence of BSL1, YFP signals were found diminished along the cell periphery but increased in the nucleus (arrowheads), suggesting BIN2-BASL interactions at the cell periphery were disturbed by the expression of wild-type BSL1 but not the phosphatase-dead BSL1D584N. Scale bars in (a-b), 50 μm. c, Quantification of YFP polarization at the cell periphery in the BIN2-BASL and BIN2-POLAR BiFC assays. The method for quantification of polarization in the BiFC assays was described in Fig. 1. *** P < 0.0001. n.s., not significant. d, BiFC assays show protein-protein interaction between BIN2 and POLAR (yellow). Half YFPs (nYFP or cYFP) were used as negative controls. e, BiFC assays show the interaction of BIN2-POLAR (yellow) was not changed by the expression of BSL1-CFP (cyan, top) or BSL1D584N-CFP (cyan, bottom). Representative individual cells were chosen from three independent experiments. Scale bars in (d-e), 50 μm. f, Quantification of membrane/nuclear (M/N) partition in the BIN2-BASL and BIN2-POLAR BiFC assays. Average fluorescence intensity values were taken from the cell periphery and from nucleus area for calculation. To generate c and f, confocal images were captured with same settings and absolute fluorescence intensity from z-stacked images were measured by Fiji. Box plot shows first and third quartiles, median (line) and mean (cross). n, number of cells. One-way ANOVA followed by Tukey’s post hoc test were used to compare with their respective control. *** P < 0.0001. n.s., not significant. g, Representative confocal images show differential membrane/nuclear partitioning of BIN2-YFP (driven by the endogenous promoter) in the Arabidopsis root elongation zone of the wild-type vs. in bsl-quad mutants. Scale bars, 10 μm. (z), images are z-stacked. Right, quantification of membrane/nuclear (M/N) partition of BIN2–YFP. Box plot shows first and third quartiles, median (line) and mean (cross). n, number of cells counted. Two-tailed Student’s t-tests were used to compare with in the wild type. *** P < 0.0001.
Extended Data Fig. 7 BSL1 Physically Interacts with YDA in vitro and in vivo.
a, In vitro pull-down assays using purified recombinant proteins show GST-BSL1 interaction with MBP-YDA. MBP-YDA used as bait. GST (negative control) or GST-BSL1 interaction were detected by anti-GST. b, Co-IP data show in vivo interaction between BSL1 and YDA. BSL1-FLAG was used as bait to detect the binding of YDAKI-Myc in 5-dpg Arabidopsis seedlings. The expression of BSL1-FLAG is driven by the TMM promoter and the expression of YDA is driven by the BASL promoter. YDAKI (Kinase Inactive YDA) was used to avoid the activity of YDA strongly suppressing plant growth. Data represent results of experiments repeated three times in a and b. c, BiFC assays show the interaction between BSL1 and YDAKI (kinase inactive YDA variant used to suppress active YDA-triggered cell death) occurs at the cell membrane in N. benthamiana leaf epidermis. Positive protein-protein interactions were visualized by YFP signal (yellow). Half YFPs (nYFP or cYFP) were uses as negative controls. Scale bars, 50 μm.
Extended Data Fig. 8 Characterization of the Loss-of-Function bsl Mutants.
a, Semi-quantitative RT–PCR analysis of single bsl T-DNA insertion lines using total RNAs isolated from Arabidopsis seedlings at 3-dpg. gDNA, genomic DNA from the wild type used as template. Primers used for RT–PCR were listed in Supplementary Table 2. Red color indicates the mutant lines used for generation of high order mutants in this study. b, Stomatal phenotype in 5-dpg adaxial cotyledon epidermis of wild type, single bsl T-DNA insertional mutants (top), and double mutants of bsl1;bsl2, bsl2;bsl3 and bsl1;bsl3 (bottom, cells are traced with blue for stomatal guard cells, dark pink for stomatal lineage cells, light pink for pavement cells). Note, mildly clustered guard cells appear in the double mutants. Scale bars, 40 μm. c, Quantification of Stomatal Index (# stomata relative to # total cells) of the designated genotypes in b. Box plot shows first and third quartiles, median (line) and mean (cross). n, number of cotyledons used for quantification of each genetic background. One-way ANOVA with Tukey’s post hoc test were used to compare with the wild type. n.s., not significant. * P < 0.05, ** P < 0.005. d, Elevated number of cells expressing the stomatal lineage identity marker SPCH in bsl-quad mutants. Left: Confocal images (z-projections) show the expression of SPCH-CFP (cyan) in the wild-type (top) and bsl-quad mutants (bottom) at different time point (after gemination) during stomatal development. Cell outlines were stained with PI (magenta). Each representative image was selected from 5 comparable cotyledons for each stage. Scale bars, 50 μm. Right, quantification of SPCH-CFP expression in wild-type and in bsl-quad mutant. Ratios of CFP-positive cells relative to the total cell numbers in a given areas were calculated. Box plot shows first and third quartiles, median (line) and mean (cross). n, total number of cells collected from > 20 cotyledons at 36- and 54-hpg. Data were analysed by two-tailed Student’s t-test. * P = 0.0169.
Extended Data Fig. 9 BSL Requires BIN2 and YDA to Regulate Stomatal Development in Arabidopsis.
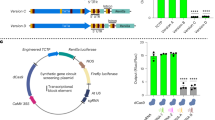
a, Overexpression phenotypes of the TMM promoter driven BSL1 (green) in 5-day adaxial cotyledon epidermis. Right, qPCR data show relative expression levels of BSL1 in two independent overexpression lines used in this study. Total RNAs were extracted from 3-day-old seedlings. Gene expression levels were normalized by ACTIN2 and relative expression levels of BSL1 were compared with the values in the wild type. Data are presented as mean ± SD. n = three independent experiments. Primers used were listed in Supplementary Table 2. b, Stomatal phenotypes in 5-day adaxial cotyledon epidermis overexpressing phosphatase-dead BSL1D584N (green) in wild-type (left) and overexpressing wild-type BSL1 (red, right) in the basl-2 mutant (right), both driven by the TMM promoter. Note the stomatal phenotype generated by BSL1D584N resembled that observed for bsl-quad mutants, suggesting a dominant negative effect of BSL1D584N. c, Overexpression of BSL1 (green) driven by the TMM promoter in the loss of function yda-3 mutant did not alter the stomatal phenotypes in yda-3. d, Overexpression of constitutively active YDA (YDACA, green) driven by the SPCH promoter in the wild-type (left) and in bsl-quad mutants (right). The activities of YDACA suppressed the loss-of-function phenotypes in bsl-quad. e, Overexpression of BSL1 (green) driven by the TMM promoter (BSL1 ++) in the loss-of-function mutants bin2-3;bil1;bil2 (bin). bin mutations appeared to be epistatic. Cell outlines were stained with Propidium Iodide (magenta). f, Stomatal phenotype in 5-dpg adaxial cotyledon epidermis of Ws-2 (background of bin2). Right, quantification of Stomatal Index (# stomata relative to # total cells) of the designated genotypes. Box plot shows first and third quartiles, median (line) and mean (cross). n, number of cotyledons used for quantification. One-way ANOVA with Tukey’s post hoc test were used to compare with the wild-type Col-0. n.s., not significant. *** P < 0.0001. Scale bars in (a-e), 50 μm.
Extended Data Fig. 10 The Membrane Receptors ERECTA, SERK, and TMM are not required for BSL to Regulate Stomatal Development in Arabidopsis.
a-c, Confocal images show stomatal development (black/white) and protein expression (green) in 5-day adaxial cotyledon epidermis of the designated genotypes. Overexpression of BSL1 (BSL1 ++, green) was driven by the TMM promoter in loss-of-function receptor mutants, that is er;erl1;erl2 (a), serk1;2;3;4 (b), and tmm (c). Cell outlines were highlighted by PI-staining (magenta). Representative images were selected from at least 10 individual cotyledons. Scale bars, 50 μm. d, Quantification of Stomatal Lineage Index (SLI) of the genotypes shown in (a-c). Box plot shows first and third quartiles, median (line) and mean (cross). More than five cotyledons were scored for each genotype. Two-tailed Student’s t-test were used to compare with their respective genetic backgrounds. ** P < 0.005, *** P < 0.0001.
Supplementary information
Supplementary Information
Supplementary Figs. 1 and 2 and Supplementary Table 2.
Supplementary Table 1
Co-IP and MS identification of BSL proteins as BASL-interacting proteins in Arabidopsis.
Source data
Source Data Fig. 4
Unprocessed western blots.
Source Data Fig. 5
Unprocessed western blots.
Source Data Extended Data Fig. 2
Unprocessed western blots.
Source Data Extended Data Fig. 5
Unprocessed western blots.
Source Data Extended Data Fig. 7
Unprocessed western blots.
Rights and permissions
About this article
Cite this article
Guo, X., Park, C.H., Wang, ZY. et al. A spatiotemporal molecular switch governs plant asymmetric cell division. Nat. Plants 7, 667–680 (2021). https://doi.org/10.1038/s41477-021-00906-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41477-021-00906-0
This article is cited by
-
Deconvoluting signals downstream of growth and immune receptor kinases by phosphocodes of the BSU1 family phosphatases
Nature Plants (2022)
-
Dichotomy of the BSL phosphatase signaling spatially regulates MAPK components in stomatal fate determination
Nature Communications (2022)
-
Connected function of PRAF/RLD and GNOM in membrane trafficking controls intrinsic cell polarity in plants
Nature Communications (2022)