Abstract
Recent advances in the field of landscape genetics provide ways to jointly analyze the role of present-day climate and landscape configuration in current biodiversity patterns. Expanding this framework into a phylogeographic study, we incorporate information on historical climatic shifts, tied to descriptions of the local topography and river configuration, to explore the processes that underlie genetic diversity patterns in the Atlantic Forest hotspot. We study two montane, stream-associated species of glassfrogs: Vitreorana eurygnatha and V. uranoscopa. By integrating species distribution modeling with geographic information systems and molecular data, we find that regional patterns of molecular diversity are jointly explained by geographic distance, historical (last 120 ky) climatic stability, and (in one species) river configuration. Mitochondrial DNA genealogies recover significant regional structure in both species, matching previous classifications of the northern and southern forests in the Atlantic Forest, and are consistent with patterns reported in other taxa. Yet, these spatial patterns of genetic diversity are only partially supported by nuclear data. Contrary to data from lowland taxa, historical climate projections suggest that these montane species were able to persist in the southern Atlantic Forest during glacial periods, particularly during the Last Glacial Maximum. These results support generally differential responses to climatic cycling by northern (lowland) and southern (montane) Atlantic Forest species, triggered by the joint impact of regional landscape configuration and climate change.
Similar content being viewed by others
Introduction
Climate and landscape configuration—and their changes over time—jointly impact how organism occupy and move through geographic space (Knowles 2001; Hewitt 2004). By uncovering the spatial patterns of phylogeographic structure emerging from these processes, and asking how they correlate with descriptors of topography and the environment, recent studies have explored the joint roles of physical and climatic barriers on genetic structure (Goldberg and Waits 2010; Wang et al. 2013; Gutiérrez-Rodríguez et al. 2017; Oliveira et al. 2018). This approach can be especially insightful by elucidating the drivers of lineage diversification, and hence guiding conservation, in megadiverse, threatened regions of the world.
Here, we apply it to one such biodiversity hotspot, the Atlantic Forest of Brazil. Topographically complex and environmentally heterogeneous, the Atlantic Forest harbors one of the highest levels of biodiversity and endemism in the world (Ribeiro et al. 2009). This forest domain is characterized by strong seasonality, sharp environmental gradients, and orographic-driven rainfall because of the easterly winds from the tropical Atlantic (Fundação Instituto Brasileiro de Geografia e Estatística 1993). Both geographical and environmental features are expected to impact patterns of genetic structure in Atlantic Forest taxa. On one hand, river systems, mountains, and tectonic faults have all been shown to coincide with phylogeographic breaks in terrestrial species, including pitvipers (Grazziotin et al. 2006), lizards (Pellegrino et al. 2005), birds (Cabanne et al. 2008; Amaral et al. 2013), mammals (Costa 2003), and anurans (Brunes et al. 2010; Thomé et al. 2010). On the other hand, paleodistribution models, coupled with genetic data, suggest that current patterns of intraspecific genetic variation in Atlantic Forest species may be explained by demographic shifts in response to Late Quaternary climate change (Cabanne et al. 2008; Carnaval and Moritz 2008; Carnaval et al. 2009; Thomé et al. 2010). Paleoclimate models suggest that presently lowland forest-dependent species had their ranges reduced to refugial areas under climatic conditions of the Last Glacial Maximum (LGM; Carnaval et al. 2009). In contrast, paleoclimate, molecular, and pollen data suggest that the montane forests, as well as their associated taxa, persisted or expanded during the LGM (Carnaval et al. 2009; Amaro et al. 2012; Leite et al. 2016).
In this paper, we seek to investigate the joint contributions of these elements—namely geographical distance, landscape features, and climate—on the genetic structure of the Atlantic forest biota. For that, we apply a multiple matrix regression approach to DNA polymorphism data from two small-sized species of stream-breeding glassfrogs that partially co-occur in its montane environments: Vitreorana eurygnatha and V. uranoscopa (Guayasamin et al. 2009; Fig. 1). V. eurygnatha has been collected in the vicinity of narrow streams and rivulets, up to 1700 m above sea level, while Vitreorana uranoscopa occurs in similar habitats but also near larger rivers (Heyer 1985), and reaches altitudes up to 1200 m. Given their association with montane regions and rivers, and in the face of the limited dispersal capacity and strong habitat dependence of amphibians in general (Pounds and Crump 1994; Pounds et al. 1999; Gardner 2001), we expect these species to exhibit strong phylogeographic structure in response to landscape and environmental conditions.
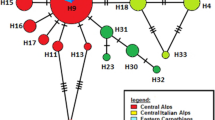
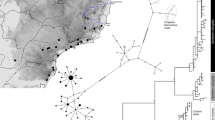
a Maximum clade credibility mtDNA ultrametric tree obtained in Beast v.1.8. Symbols in the nodes represent posterior support (+>0.7 and *>0.9). The tree was rooted using Hyalinobatrachium taylori as an outgroup (not depicted in the figure). Marked clades correspond to groups as defined by the GMYC analysis. b, c Maps of collected samples and corresponding groups according to the GMYC analysis (black dots denote non-clustered individuals) for both V. eurygnatha (b) and V. uranoscopa (c). Symbols for each species are independent. Maps include rivers which are important drivers of genetic differentiation in these groups, and the Brazilian states where the studied species were sampled (Rio Grande do Sul, Santa Catarina, Paraná, São Paulo, Minas Gerais, Rio de Janeiro, Espírito Santo, and Bahia). Shadowed areas on the maps correspond to forest regions defined by Carnaval et al 2014: southern (largely montane) Atlantic Forest spaces in red, and northern (largely lowland) Atlantic Forest in gray. d Coalescent-based species tree based on four nuclear genes (CMYC, POMC, BDNF and RAG) in both species, reconstructed using *BEAST. Numbers represent posterior support for nodes when >60%
For that, we characterize not only contemporary climates and present-day barriers to gene flow (e.g., Nielson et al. 2001; Pellegrino et al. 2005; Wang 2013; Manthey and Moyle 2015), but also Late Quaternary climatic landscapes in the Atlantic Forest. Because abundant data point to the importance of past environmental conditions on the generation, maintenance, and erosion of lineage diversity (Hewitt 2000; Knowles 2000; Carnaval et al. 2009), we build on previous matrix regression studies (Wang et al. 2013) and explicitly include the distribution of historical climates (over the past 120 ky) in our analysis of potential drivers of phylogeographic structure.
We expect multiple processes to have contributed to genetic structure within Vitreorana eurygnatha and V. uranoscopa. Given these species’ present-day association with cool, montane environments, we hypothesize that both former and present-day climates had and have an important role defining the geographic areas that allowed for long-term persistence and accumulation of lineages. Because these are small and stream-associated frogs, we also expect local patterns of gene flow to be correlated with geographic distance and mediated by the spatial configuration of river basins. To verify how rivers may impact gene flow in these taxa, we explore their role not only as corridors (to the aquatic larvae of the species), but also as potential barriers (to their terrestrial adults). Beyond isolation by distance, we also hypothesize that topography itself has had an impact on the distribution of genetic diversity within each species. We expect so because of the topographical complexity of the Atlantic Forest region, and the species’ association with montane environments.
To ask whether we can find evidence of the impact of these multiple forces on local genetic structure, we apply a multiple matrix regression with randomization approach to verify if and to what extent these elements explain the patterns of genetic variation observed within these species today. To describe phylogeographic structure, we present new mitochondrial and nuclear DNA sequences from Vitreorana tissues across the known range of these species (IUCN, 2017), which are here analyzed under coalescent and Bayesian approaches. To evaluate the extent to which geography, landscape configuration, and historical climate change explain intraspecific genetic patterns, we then generate and employ resistance layers to map how distance, relief, rivers, long-term climatic stability (over the past 120 ky), and climatic extremes over this period each may have impacted gene flow throughout the forest.
Materials and methods
DNA sampling, extraction and sequencing
Long-term collection efforts allowed us to gather 108 tissue samples covering most of the known ranges of our target taxa. This includes 34 samples of V. eurygnatha (17 localities; Fig. 1b) and 74 of V. uranoscopa (43 localities, Fig. 1c). We also gathered two samples of Hyalinobatrachium taylori, a genus recovered as an outgroup of Vitreorana (Castroviejo-Fisher et al. 2014), to root our phylogenetic trees. Collection sites were either assigned to geographic coordinates in the field, using a GPS, or geo-referenced using collection notes (Supplementary Table 1). Vouchers are housed in the following Brazilian institutions: Célio F. B. Haddad amphibian collection in the Departamento de Zoologia, I.B., Universidade Estadual Paulista, Rio Claro, SP (CFBH); Museu de Ciências e Tecnologia da PUCRS (MCP), Rio Grande do Sul; Museu Nacional Rio de Janeiro (MNRJ); and Tissue Collection of the Herpetology Lab, Instituto de Biociências, Universidade de São Paulo (MTR).
Genomic DNA was extracted from ethanol-preserved liver or muscle samples, with a high salt extraction method (Miller et al. 1988). We used Polymerase Chain Reaction (PCR) to amplify two mitochondrial and four nuclear gene fragments. The mitochondrial DNA fragments consisted of NADH Dehydrogenase Subunit 1 (ND1, 1014bp) and cytochrome c oxidase subunit I (COI, 608 bp). The nuclear fragments included the proto-oncogene cellular maelocytomatosis (CMYC, 415 bp), the proopiomelanocortin A gene (POMC, 589 bp), the recombination activating gene (RAG, 444 bp), and the brain derived neurotrophic factor (BDNF, 615 bp). Gene-specific primer sequences and amplification programs (Supplementary Tables 2, 3) were adapted from the literature (van der Meijden et al. 2007; Guayasamin et al. 2008; Lyra et al. 2017).
DNA concentration was determined with Nanodrop (Thermo Scientific), and aliquots were diluted to 100 ng/ml for amplification. Polymerase Chain Reactions (PCRs, 12.5 µl total volume) contained 2.5 µl of 10x reaction buffer, 1–3 mM MgCl2, 0.25 mM mixed dNTPs, 0.5 µM of each primer, and 0.0625 of Promega Hot Start Taq. PCR products were visualized in agarose gels, and purified with ExoSap, following the manufacturer’s protocol (ExoSap-it, GE Healthcare). Samples with unsuccessful PCRs were amplified with Ready-to-go RT-PCR Beads (illustra, GE Healthcare, Pittsburgh), following manufacturer’s protocol. Sequencing reactions and sequencing were outsourced to Macrogen Corporation (www.macrogen.com) and GENEWIZ Inc. (www.genewiz.com).
Chromatograms were visually inspected for errors, and contiguous sequences were assembled in Sequencher v4.1 (Gene Codes Corp. 2000, http://www.genecodes.com/) and Geneious v5.4 (Biomatters, http://www.geneious.com). All sequences were aligned using the MUSCLE algorithm in Geneious, under default settings, and checked by eye.
Phylogeographic analyses
To compare patterns of genetic structure, and to delineate target units for the study of diversity correlates, we first estimated a mitochondrial (mtDNA) gene genealogy. We combined the mitochondrial sequences of both species in a single analysis, given that they had been already identified as sister species (Castroviejo-Fisher et al. 2014), and inferred a Bayesian tree in BEAST v 1.8 (Drummond and Rambaut 2007). First, we selected the best partition scheme (including genes and codon positions) for our dataset. We then identified the model of nucleotide evolution best fit to each partition, using the BIC criteria as implemented in PartitionFinder V1.1.1 (Table S4, Lanfear et al. 2012). Finally, we used a strict clock with mean substitution rate fixed to 1, and ran the analysis for 100,000,000 generations, logging every 10,000 generations. The use of a strict clock is appropriate in this case given the recent divergence times involved, and the close relationship between the species. Under these conditions, molecular rate variation is expected to be low across lineages, justifying the choice of clock (Weir and Schluter 2008; Brown and Yang 2011). Chain convergence was evaluated with Tracer v1.6 (Rambaut et al. 2014), both visually and numerically, ensuring a minimum ESS value of 200. A maximum clade credibility tree was estimated with Tree Annotator, applying a burnin of 10%.
To objectively identify regional mtDNA clusters and singletons (i.e., sequences not belonging to any cluster), we used the resulting mtDNA Bayesian tree for each species in a single threshold General Mixed Yule-Coalescent approach (GMYC, Pons et al. 2006). We implemented GMYC in the SPLITS package for R (available from http://r-forge.r-project.org/projects/split). To verify whether the clusters identified through the mtDNA analysis were supported by the nuclear dataset, we used two methodologies: a coalescent-based species tree as implemented in *BEAST (Heled and Drummond 2009), and a Bayesian clustering method as implemented in Structure V2.3.4 (Pritchard et al. 2000) and Bayesian Analysis of Population Structure, BAPS (Corander et al. 2008). Details of both analyses are outlined below.
To use a *BEAST coalescent-based species tree to evaluate whether the nuclear data supported the mitochondrial clusters identified by GMYC (Heled and Drummond 2009), we first selected the model of nucleotide evolution that best fitted each nuclear gene, using the BIC criteria as implemented in PartitionFinder V1.1.1 (Table S4, Lanfear et al. 2012). Codon partitions were not included in this analysis. Then, we used the mtDNA clusters and singletons, as determined by the GMYC analysis, as taxon sets—with the exception of V. eurygnatha mtDNA cluster 2 (Fig. 1 yellow triangle) and singleton 17, and V. uranoscopa singleton 22, which did not have a minimum of one sequence per gene (e.g., cluster 2 has two samples but we were unable to amplify sequences of BDNF for either one). Files were prepared as per the *BEAST template for BEAUti v2.4.4 (Heled and Drummond 2009), and the species tree was reconstructed in Beast v2.4.4, with 50,000,000 generations, by sampling every 25,000 generations. Chain convergence was evaluated in Tracer v1.6 (Rambaut et al. 2014), and the tree was plotted with DensiTree v2.2.5 (Bouckaert and Heled 2014), applying a 10% burn-in.
To evaluate the clustering of lineages based on the nuclear data with Structure V2.3.4, we first performed a statistical haplotype reconstruction of the nuclear fragments (CMYC, RAG, BDNF, and POMC), under a Bayesian framework, in the program PHASE v2.1.1 (Stephens et al. 2001; Stephens and Scheet 2005). For that, we used SeqPHASE (Flot, 2010) to generate PHASE input files for all loci. We performed five independent runs of PHASE and considered the gametic phases resolved whenever the posterior probabilities were equal or higher than 0.9. For the Structure analyses, we used an ancestry model that allowed for admixture and a correlated allele frequency model. By treating every single nucleotide polymorphisms (SNPs) in the concatenated nuclear alignment as separate locus, we used the distance between adjacent SNPs from a single gene to create a “map distance” between loci. Because Structure can be biased by the higher levels of hierarchy in a dataset, we ran the analysis separately for each species, varying K between one and the total number of mtDNA clusters and singletons as determined by GMYC (i.e., ranging between 1 and 13 for V. eurygnatha, and 1 and 14 for V. uranoscopa) (Kalinowski, 2011; Puechmaille, 2016). Analyses were repeated ten times for every K value, including an initial set of 100,000 generations that were discarded as burn-in, followed by 100,000 generations. We calculated averages of cluster membership coefficients from all runs for each K, with the Software CLUMPP v.1.1.2 (Jakobsson and Rosenberg, 2007), and plotted the data in R. For all analyses, we identified the best K through the Evanno method (Evanno et al. 2005), as implemented in Structure Harvester (Earl and VonHoldt, 2012). Because Structure is most commonly used for marker data (instead of sequence information; but see Falush et al. 2003), we ran BAPS (Corander et al. 2008) considering each gene as a single locus. As in the structure analysis, we varied K between one and the number of mtDNA clusters, and then applied the clustering-with-linked-loci option. For each species, the analyses for the best K (K = 3) and K = total number of mtDNA clusters (that is, K = 13 or K = 14) were repeated five times each.
To calculate the dependent matrix of genetic distances for the analyses of matrix correlation (see below), we calculated the between-group mean distance between each pair of mtDNA clusters or singletons in Mega 7 (Kumar et al. 2016), using the Tamura-Nei model of substitution.
Landscape resistance layers
To assess the effect of geography, landscape configuration, and historical climatic shifts on the intraspecific levels of divergence in V. eurygnatha and V. uranoscopa, we first created seven resistance layers. From them, we calculated least-cost paths to gene flow. Each layer represented one element that potentially impacts genetic structure within these glassfrogs. They are: geographic distance (layer 1), spatial distance given the topography (layer 2), river as barriers (layer 3), rivers as corridors (layer 4), climate stability over the last 120 ky (layer 5), and climatic suitability at both the most suitable (layer 6) and least suitable (layer 7) climate period experienced over the past 120 kya.
To construct layer 1 (IBD), we created a resistance layer that solely measures isolation-by-distance based on simple geographical distances between points. To construct layer 2, we created a topographic resistance surface, using a digital elevation model of the region (Jarvis et al. 2008), to estimate the physical distance (including changes in altitude) to traverse through each pixel.
To construct layer 3, which maps the extent by which river may acts as barriers to dispersal, we created a river-based resistance layer. To build layer 4, a surface depicting how rivers may act as corridors. For both, we used a hydrographic information GIS layer from the HydroSHEDS project, deriving hydrographic information from topographic data (Lehner et al. 2008). For layer 3, we considered all rivers as equal barriers to gene flow, placing a single value of 100 on each one, whereas all other cells were given a value of 1. For layer 4 we considered all rivers as equal corridors to gene flow, placing a single value of 1 on each one, whereas all other cells were given a value of 100. A more complex friction layer created under the assumption of differential river permeability (in which river size was taken into consideration according to the Strahler number; Strahler 1957) was highly correlated with the one which assumed rivers as equal barriers to gene flow. For simplicity purposes, we opted to utilize the equal permeability layer.
Building resistance layers from past climates
To evaluate the extent by which historical climate and climatic shifts may have impacted the distribution of haplotypes over time, we combined eight bioclimatic descriptors related to temperature and precipitation and known occurrence data to build distribution models (SDM; Elith and Leathwick 2009) for each species. Environmental information used to build the SDMs consisted of bioclimatic variables derived through the interpolation of weather station data—including temperature, precipitation, and seasonality from the Worldclim database (Hijmans et al. 2005). To be able to generate projections over regular intervals of time, we used the same eight variables for which multiple past climate reconstructions are available, at 4 kya intervals, through the Hadley Center Climate Model (HadCM3, Singarayer and Valdes 2010). Those include four variables related to temperature (Bio 1: Annual mean temperature, Bio 4: Temperature Seasonality, Bio 10: Mean temperature of the warmest quarter, Bio 11: Mean temperature of the coldest quarter), and four variables related to precipitation (Bio 12: Annual precipitation, Bio 15: Precipitation seasonality, Bio 16: Precipitation of wettest quarter, Bio 17: Precipitation of driest quarter). Present-day climate models were trained with 1 km resolution data and projected to snapshot simulations covering the last 120 ky, at 4 ky intervals, using HadCM3 at 5 km resolution (Carnaval et al. 2014). To minimize the impact of sampling bias on the SMDs, we used the spThin package in R (Aiello-Lammens et al. 2015) to identify all occurrence records with a linear distance of less than 5 km to each other, using just one for model building. This resulted in a database of eight occurrence records for V. eurygnatha, and 17 for V. uranoscopa (Table S1). We restricted model background selection to an ecologically relevant dispersal limit consisting of a minimum-convex polygon based on known occurrences buffered by 100 km. Models were built using Maxent version 3.3.3k (Phillips et al. 2006; Phillips and Dudík 2008). Maxent parameters were tuned with the ENMeval package for R (Muscarella et al. 2014), experimenting several combinations of three feature classes (“L”, “LQ”, “H”, “LQH”, “LQHP”) and different regularization multipliers (0.5–4 with 0.5 increases). For each species, we selected the model with the lowest AICc as the one with the best combination of parameters. All models and projections were converted from continuous to binary predictions using the 10th percentile training presence threshold (Pearson et al. 2007), which indicates the lowest value of prediction for 90% of the known presence localities used in training the model.
With the optimized SDMs, we first estimated a species-specific climatic stability map over the past 120 ky; from it, we created a historical climatic resistance layer (layer 5). The latter represents how difficult or easy it has been for each species, historically, to disperse through the Atlantic Forest—based on climate. This estimate takes into account the climatic niche occupied by the species today, and the environmental shifts inferred as per the paleoclimatic simulations employed. To create the historical climatic resistance layer, we first prepared a climate stability map for each species by summing all binary SDM projections, from present time to 120 kya, at 4 ky intervals. This stability map was then converted into a resistance layer by inverting the pre-grid cell stability values in SDMtoolbox 1.1b (Brown 2014). With this step, areas of high stability became areas of low friction.
Further exploring the binary SDM results, we also selected, for each species, the two periods within the last 120 ky that allowed for their ranges to be the broadest and the most contracted, respectively. Using SDMs from these most impacting times, we estimated two landscape resistance layers per species (layers 6 and 7) that represent, respectively, how difficult and how easy it has been for each taxon to disperse during those periods.
Multiple matrix regressions with randomization
To measure how much geography, landscape configuration, and historical climate explain the genetic divergence presently observed within the target species, we created mixed-models using multiple-matrix regressions with randomization (MMRR), through the R function “MMRR” (Wang 2013) available from the Dryad Data Repository (Dryad https://doi.org/10.5061/dryad.kt71r). This MMRR treats the amount of genetic divergence observed between each pair of mtDNA clusters and singletons, as defined by the GMYC analysis, as a dependent (response) matrix. Seven matrices, each describing an aspect of the environment (geography, topography, rivers as barriers and rivers as corridors, climate stability, and climatic extremes), were input as the independent (predictor) matrices.
To generate an independent matrix to represent each landscape resistance layer, we calculated a cost-distance nxn matrix for resistance layers 1–7, where n was the number of mtDNA clusters and singletons identified by the GMYC analysis (13 for V. eurygnatha and 14 for V. uranoscopa). For each layer, we measured the dispersal cost between each pair of mtDNA clusters to identify the most efficient route, or the least-cost-path (LCP), connecting them. Least-cost distances were calculated between all pairs of clusters, for each species, using the pairwise distance matrix function in SDMtoolbox v1.1a (Brown 2014). Whenever a cluster was represented in more than one locality, the centroid of the distribution of the cluster was used as a focal site. This resulted in seven independent distance matrices per species, each representing the least-cost distances connecting two mtDNA clusters, given the Atlantic Forest geography (matrix 1, which is the IBD matrix), landscape (including rivers, matrices 2, 3, 4), and historical climates (matrices 5, 6, 7).
An MMRR is a multiple regression of distance matrices, where significance is tested through random permutations of the rows and columns of the dependent matrix, to assess the regression and its coefficients (Manly 1991; Legendre et al. 1994). This method maintains the non-independence of the matrix elements (Manly 1986, 1991; Smouse et al. 1986; Legendre et al. 1994) and, because of this, does not suffer from type I errors associated with randomizing matrix values (Harmon and Glor 2010; Guillot and Rousset 2013; Wang 2013). To properly run the MMRR, we first measured the amount of correlation between all independent matrices. Whenever least-cost distance matrices 2, 3, 4, 5, 6, or 7 were correlated to matrix 1 (that is, the IBD matrix), they were regressed against the latter and only the residuals were used for further analyses. When other variables were correlated with each other, we deleted one variable from the model. For consistency purposes, we opted to prefer climatic stability (matrix 5) over climatic extremes (matrices 6 and 7). The uncorrelated matrices that resulted from this verification step (either residuals or original), along with the IBD least-cost-path matrix (matrix 1), were simultaneously loaded as explanatory variables in a species-specific MMRR to generate a full model.
The MMRR analyses were run iteratively, starting with all matrices as predictor variables and then excluding all that did not contribute significantly to the model (α = 0.05). Exclusion steps happened until a final model was reached, including only those explanatory variables that contributed significantly to the results. Because both species have a highly differentiated and largely isolated lineage in the northernmost part of their distribution (the northern tip of the state of Minas Gerais, in the case of V. eurygnatha, and the northeastern state of Bahia, for V. uranoscopa; Fig. 1), we also ran an MMRR, for each species, in which we excluded these highly differentiated and isolated mtDNA clusters. With this, we avoided the potential overwhelming effect of geographical distance, given the geographical distribution of the samples, on the MMRR analysis.
Results
Phylogenetic and population genetic analyses
Amplification and sequencing success varied across markers (RAG n = 108; POMC n = 108; CMYC n = 107; BDNF n = 80; COI n = 106; ND1 n = 103; Table S1). The results indicate that both species have spatially structured mitochondrial clades along the latitudinal range of the forest. Vitreorana eurygnatha has a smaller range, and its southernmost mtDNA clusters are distributed within the range of the central clades of V. uranoscopa. A GMYC analysis recognizes four unique mtDNA clusters within V. eurygnatha (symbols in Fig. 1a, b; nine individuals (singletons) were left without a cluster), which are here described. A southern mtDNA clade is distributed between the Paranapanema and Tietê rivers in the state of São Paulo (samples 17 and 24, Fig. 1a, b). A central mtDNA clade includes representatives in the highlands of northern São Paulo, Rio de Janeiro and Minas Gerais states (Fig. 1a, b; green squares, yellow triangles, singleton 25). Further north, a well-supported mtDNA clade includes samples from the Caparaó mountain range near the border of the states of Espírito Santo and Minas Gerais, as well as an individual further north in Minas Gerais, by the Jequitinhonha river (Fig. 1a, b; red circle, singletons 18 and 21). A fourth, northernmost clade includes samples from the Caparaó mountain range, inland and northern Minas Gerais (Jequitinhonha, the Serra do Cipó National Park and Cardeal Mota) and the coastal mountains of Espírito Santo (Santa Teresa; Fig. 1a, b; blue pentagon, singletons 19, 20, 23, and 26).
In V. uranoscopa, the GMYC analysis also recovers a well-supported mtDNA southern clade distributed from the Uruguai river, in the south, to the Paranapanema river. It includes representatives of two major lineages. One of them, which is well-supported in the mtDNA Bayesian analysis, includes specimens restricted to the state of São Paulo (Fig. 1a, c, yellow square, inverted purple triangle, red pentagon). The second southernmost clade includes samples from the south of the State of São Paulo to the state of Santa Catarina (Fig. 1a, c, green circle, white inverted triangle, purple cross). The northern clade recovered by the mtDNA phylogenetic reconstruction is split in two subclades. A northernmost, geographically isolated, and well-supported mtDNA subclade is restricted to the region between the Pardo and Paraguaçu rivers, in Bahia (nearest samples located more than 500 km away; Fig. 1a, c, yellow pentagon). The other mtDNA clade, with lower support, spans the central portion of the forest, including the coastal regions from northern São Paulo to the state of Espírito Santo, just south of the Doce river. The GMYC analysis detected high levels of genetic structure within this dataset, recognizing 12 mtDNA clusters within the clades described above (two individuals, numbered 22 and 27, were not clustered).
The use of nuclear data in a species-tree analysis of V. eurygnatha recovers groupings that largely reflect the southern, central, and northern clades detected by the mtDNA—only analyses (although with lower support for the southern and central groups). One V. eurygnatha clade (posterior = 100) includes individuals collected near the Jequitinhonha river in northern Minas Gerais, several of which (all but sequence 21) correspond to the northernmost mtDNA cluster (Fig. 1b, d, blue pentagon). Another clade (posterior = 76) contains individuals from one mtDNA cluster from the central region and two singletons from southern Minas Gerais (Fig. 1b, green squares, singletons 23 and 26). A third clade (posterior = 88) contains individuals from one mtDNA cluster and two singletons from Minas Gerais (Fig. 1b, d, red circle, singletons 18 and 19), all south of the Doce river and north of the Grande river. Clade relationships recovered by the species tree analysis differ, however, from those inferred by the mtDNA data (Fig. 1a). The incorporation of nuclear data through a species tree analysis (Fig. 1d) also finds lower overall support for the monophyly of V. eurygnatha (posterior = 48).
In V. uranoscopa, the analysis of the nuclear data provides different insights relative to the mtDNA-only data (Fig. 1). Although the northern individuals are placed in a clade sister to the remaining samples, only the latter is well-supported. Moreover, the analysis of the nuclear data fails to find support for the clades identified by the mtDNA-only Bayesian inference. All posteriors support values within the nuclear data analysis of V. uranoscopa were lower than 50.
Differently from the large number of clusters identified by the mtDNA-based GYMC analysis, a Structure analysis of the nuclear DNA found support for only two groups within each species (that is, the minimum possible number of clusters that can be determined with this algorithm; Fig. S1 and Fig. S2). The BAPS analyses identified, however, three groupings within each taxon. In V. eurygnatha, both assignment analyses recognize the uniqueness of the northernmost Minas Gerais sites (blue pentagon and sequence 21 in Fig. 1a, b and Fig. S1). The Structure analysis assigned samples from all remaining localities within a single population, whereas, in BAPS, a few singletons were grouped in a third population. In V. uranoscopa, Structure assigned all individuals from the northernmost and southernmost sites into one population (Fig. S1c, green), whereas BAPS distributed them into two groups. Central sites were assigned to a unique population in both analyses (Fig. S1b, blue).
When we force K to match the number of clusters and singletons identified by the GMYC analysis, the V. eurygnatha Structure outputs allows us to detect patterns that match the four mitochondrial clusters identified with the mtDNA data—a pattern not recovered by the BAPS analysis. However, this is not observed in V. uranoscopa. Only one of the 11 mitochondrial clusters detected in V. uranoscopa is identified as genetically unique in the structure analysis (Bahia samples: yellow pentagon; Fig. S1 c,d, structure plots K = 13 and 14, respectively). When using BAPS, the northernmost population can still be detected, and the southern and central clades are still apparent—with several single samples assigned to unique populations.
Climate modeling, stability, and historical demography
Present-day SDMs suggest a broader distribution for V. eurygnatha compared to V. uranoscopa (Fig. 2a, b). Climate-based stability maps of both species indicate the persistence of suitable climate across time along most of their present range (Fig. 2c, d). Both V. eurygnatha and V. uranoscopa appear to have had large and continuous stable areas in the southern half of the Atlantic Forest, with some smaller and discontinuous stable areas to the north (Fig. 2c, d).
Left. Species distribution models for Vitreorana eurygnatha (a) and V. uranoscopa (b). Darker colors represent higher habitat suitability, with black depicting suitability = 1 in Maxent’s logistic output. Right. Stability maps for V. eurygnatha (c) and V.uranoscopa (d). Stability maps were obtained by adding all model projections to the past up to 120 kya every 4000 years. Darker colors represent higher stability, with black depicting presence in all 30 layers
Range projections under former climates suggest that the distribution of suitable climates for both species of Vitreorana expanded and contracted multiple times over the past 120 ky. For both species, the most contracted potential distribution was projected at 120 kya. Although similar to the present-day extent of the species, their inferred ranges during the last interglacial (120 kya) expanded slightly towards the coast and the south (Fig. 3a, b). The models suggest that the distributions of suitable habitat remained small from 120 kya until around 100 kya, and were reduced again starting ca. 12 kya, for both species. On the other hand, an expansion of suitable habitats was inferred to have happened, for both species, at 80 Kya, at 60–68, 32, and 21 kya. The broadest potential distribution was predicted not to have happened at the Last Glacial Maximum (LGM, ~21 kya), but, instead, at 32 kya. For both species, the inferred distribution at the LGM does not recover the same level of connectivity between the south and north of the forest that is inferred at 32 kya (Fig. 3a, b).
Species distribution model projections into the past, for time periods inferred to have resulted in the most contracted distribution (120 kya) in a Vitreorana eurygnatha and b V. uranoscopa, and time inferred to hae resulted in the broadest distribution (32 kya) of c V. eurygnatha and d V. uranoscopa. Darker colors represent higher habitat suitability, with black depicting suitability = 1 in Maxent’s logistic output
Multiple matrix regressions with randomization
Least-cost distance matrices were correlated with the IBD matrix in both Vitreorana species, so we used the residuals for further analyses. Climate suitability at 32 kya was highly correlated with climate stability, in both taxa. Because of this correlation, we only considered climatic stability in all further analyses. In V. uranoscopa, suitability at 120 kya was also highly correlated with climate stability, and hence eliminated from downstream analyses.
In both species, the observed genetic structure was best explained by a combination of variables. In V. eurygnatha, the model that best explained genetic patterns included geographic distance and climatic stability as explanatory variables (R2 = 0.24, p-value = 0.0003). This result held true even after removing the highly differentiated and isolated northernmost samples (R2 = 0.26, p-value = 0.0003). In V. uranoscopa, the model that best explained structure across all sampled sites had geographic distance as a single best explanatory variable (R2 = 0.32, p-value = 0.001). However, after removal of the highly differentiated northernmost samples, the best model changed to include geographic distance, stability and rivers as barriers (R2 = 0.36, p-value = 0.0009). If we had only used distance to predict this new set of samples, we would have found a significant but poorer predictor of genetic diversity (R2 = 0.23, p-value = 0.0014). If only rivers were used as predictors, R2 would have been smaller (0.13). If only stability had been used, R2 would have been smaller still (0.01).
Discussion
Multiple studies have discussed the potential influence of landscape and climate changes on the spatial distribution of genetic lineages in the Atlantic forest (e.g., Pellegrino et al. 2005; Cabanne et al. 2007; Thomé et al. 2010; Zamborlini Saiter et al. 2016). Yet, none has, to date, simultaneously evaluated the relative importance of geographical distance, landscape configuration (topography, rivers), and historical climate change (climatic stability, former environmental suitability) as predictors of diversification patterns. While doing so for Vitreorana eurygnatha and V. uranoscopa, we found that genetic patterns are most strongly correlated with geographic distance—despite the topographical complexity and the historical climate dynamics of this area. We also find evidence for the role of historical climatic stability in predicting the genetic structure of both species. In one species—V. uranoscopa, which is found along larger streams (Heyer 1985)—river barriers also help to explain the distribution of diversity. Yet, their contribution seems relatively small.
These findings make sense in the light of previously published data for Neotropical species. For instance, the structuring role of geographic distance (or IBD) has been demonstrated in frog species of the Northern Andes (Guarnizo et al. 2015) and in plants in Central America (Ortego et al. 2015). In the Atlantic Forest, the importance of IBD in explaining genetic structure has been shown in rodents (Colombi et al. 2010) and birds (Cabanne et al. 2007).
Rivers have also been frequently associated with distribution breaks in the Atlantic Forest and other Neotropical systems. Originally postulated to explain the distributions of monkeys in the Amazon (Wallace 1852), the riverine hypothesis received variable support from studies of other taxa (Gascon et al. 2000; Bates et al. 2004; Ribas et al. 2012). In the Brazilian Atlantic Forest, congruence between the placement of phylogeographic breaks and that of rivers has been noted in lizards (Pellegrino et al. 2005), frogs (Thomé et al. 2010) and birds (Cabanne et al. 2007). Tests of this hypothesis are rare, and it has been suggested that this spatial congruence with river courses may in fact result from other associated geological processes that acted as barriers (e.g., tectonic faults, Thomé et al. 2014; Thomaz et al. 2015), or from sparse sampling. In a very densely sampled study of tree species in the central corridor of the Atlantic Forest, for instance, it has been demonstrated that turnover patterns in species composition reflect environmental gradients, not river location (Zamborlini Saiter et al. 2016). In our analysis, rivers are correlated with lineage structure in only one of the studied species, Vitreorana uranoscopa. We were unable to find support for the hypothesis that they promote connectivity within basin and keep populations differentiated across basins. Instead, we found support for the idea that they correspond to areas of interrupted gene flow. The largest phylogeographic breaks identified within both species are broadly coincident with the location of two major rivers and, detected both in the mitochondrial and nuclear datasets. The break between the northern and central clades in both Vitreorana species coincide with the Jequitinhonha and Doce rivers, with the northern clades limited to the south by the Jequitinhonha, and the southern clades limited to the north by the Doce. The importance of these two rivers has been stated in the literature; the Doce river has been identified as a contact zone in multiple groups (Costa et al. 2000; De Mello Martins 2011) and as a contact zone between the two types of forest (Carnaval et al. 2014), and the Jequitinhonha river has been highlighted as coincident with genetic breaks in lizards (Pellegrino et al. 2005; Rodrigues et al. 2014).
In addition, both V. eurygnatha and V. uranoscopa share a phylogeographic break in the south of the state of São Paulo, as several other frog species (Fitzpatrick et al. 2009; Brunes et al. 2010; Amaro et al. 2012), birds (Cabanne et al. 2008), and snakes (Grazziotin et al. 2006). This break, however, does not seem to coincide with a major river, topographic, or climatic shifts in the area. Instead, it is largely (but coarsely) congruent with a NW-SE fault of the Southern Brazil Continental Rift (Amaro et al. 2012). Movement along this fault has been shown to have influenced the configuration of hydrographic basins in the Quaternary (Ribeiro 2006; Riccomini et al. 2010), and may have promoted the structuring of genetic variation in these stream-associated frogs.
Based on previous studies of montane or sub-tropical Atlantic Forest species, we expected that past climates had played an important role in structuring genetic variation in Vitreorana species (Carnaval et al. 2009; Amaro et al. 2012; Leite et al. 2016). We found support for this hypothesis: the data show that climatic stability in the past 120 kya is an important predictor of genetic variation. Moreover, for both taxa, paleomodeling provides evidence of multiple range expansions and contractions over the past 120 kya, with stable areas including most of their presently restricted range (Fig. 2). These data suggests that both montane species have been limited to present (refugial) areas several times in the past, yet likely able to explore currently unsuitable (mostly lowland) regions and hence to expand during colder periods, such as 32 kya. While this demographic syndrome may not appear intuitive for a tropical species, it is consistent with the fact that these taxa are currently restricted to sub-tropical or tropical montane areas.
Demographic expansions during the LGM are reflected in the inferred history of this and other mostly southern or montane species (Leite et al. 2016). Previous studies of lowland species from the northern forest suggested pervasive climatic instability in the Quaternary and persistence of species in wet forest stable refugia during presumably colder periods (Carnaval et al. 2009; De Mello Martins 2011). Contrary to those findings, the SDMs here presented suggest that glassfrogs encountered fairly stable habitats throughout the late Pleistocene in the south of the Atlantic Forest, with some smaller stable discontinuous areas in the north. This supports the view that the Atlantic Forest does not have a unique history, and that the northern (more tropical) and southern (more sub-tropical) components of the forest have responded differentially—and even in opposite ways—to past climatic cycles. We argue that Atlantic Forest species of distinct life habitat associations have responded differentially to the many climatic changes impacting this region over the Pliocene and Pleistocene (D’Horta et al. 2011; Carnaval et al. 2014; Cabanne et al. 2016; Raposo do Amaral et al. 2016). Vitreorana hence appears to function as a model for several other montane and sub-tropical species distributed in the southern portion of the forest.
Data archiving
DNA sequences for new mitochondrial DNA (COI and ND1) and nuclear DNA (RAG, BDNF, POMC, and CMYC) were deposited in GenBank under accession numbers MH987782–MH988395. See Table S1 for a detailed list. Voucher information and localities used for constructing environmental niche models are provided in Table S1. Climatic layers used for niche modeling are available from worldclim.org.
References
Aiello-Lammens ME, Boria RA, Radosavljevic A, Vilela B, Anderson RP (2015) spThin: an R package for spatial thinning of species occurrence records for use in ecological niche models. Ecography 38:1–5
Amaral F, Albers PK, Edwards SV, Miyaki CY (2013) Multilocus tests of Pleistocene refugia and ancient divergence in a pair of Atlantic Forest antbirds (Myrmeciza). Mol Ecol 22:3996–4013
Amaro RC, Rodrigues MT, Yonenaga-Yassuda Y, Carnaval AC (2012) Demographic processes in the montane Atlantic rainforest: molecular and cytogenetic evidence from the endemic frog Proceratophrys boiei. Mol Phylogenet Evol 62:880–888
Bates JM, Haffer J, Grismer E (2004) Avian mitochondrial DNA sequence divergence across a headwater stream of the Rio Tapajós, a major Amazonian river. J Ornithol 145:199–205
Bouckaert RR, Heled J (2014) DensiTree 2: Seeing trees through the forest. bioRxiv: 1–11.
Brown JL (2014) SDMtoolbox: a python-based GIS toolkit for landscape genetic, biogeographic and species distribution model analyses. Methods Ecol Evol 5:694–700
Brown RP, Yang Z (2011) Rate variation and estimation of divergence times using strict and relaxed clocks. BMC Evol Biol 11:271
Brunes TO, Sequeira F, Haddad CFB, Alexandrino J (2010) Gene and species trees of a Neotropical group of treefrogs: genetic diversification in the Brazilian Atlantic Forest and the origin of a polyploid species. Mol Phylogenet Evol 57:1120–1133
Cabanne GS, Calderón L, Trujillo-Arias N, Flores P, Pessoa RO, D’Horta F et al. (2016) Pleistocene climate changes affected species ranges and evolutionary processes in the Atlantic Forest. Biol J Linn Soc 119:856–872
Cabanne GS, D’Horta FM, Sari EHR, Santos FR, Miyaki CY (2008) Nuclear and mitochondrial phylogeography of the Atlantic forest endemic Xiphorhynchus fuscus (Aves: Dendrocolaptidae): biogeography and systematics implications. Mol Phylogenet Evol 49:760–773
Cabanne GS, Santos FR, Miyaki CY (2007) Phylogeography of Xiphorhynchus fuscus (Passeriformes, Dendrocolaptidae): vicariance and recent demographic expansion in southern Atlantic forest. Biol J Linn Soc 91:73–84
Carnaval AC, Hickerson MJ, Haddad CFB, Rodrigues MT, Moritz C (2009) Stability predicts genetic diversity in the Brazilian Atlantic forest hotspot. Science 323:785–789
Carnaval AC, Moritz C (2008) Historical climate modelling predicts patterns of current biodiversity in the Brazilian Atlantic forest. J Biogeogr 35:1187–1201
Carnaval AC, Waltari E, Rodrigues MT, Rosauer DF, VanDerWal J, Damasceno R et al. (2014) Prediction of phylogeographic endemism in an environmentally complex biome. Proc R Soc B Biol Sci 281:1461
Castroviejo-Fisher S, Guayasamin JM, Gonzalez-Voyer A, Vilà C (2014) Neotropical diversification seen through glassfrogs. J Biogeogr 41:66–80
Colombi VH, Lopes SR, Fagundes V (2010) Testing the Rio Doce as a riverine barrier in shaping the atlantic rainforest population divergence in the rodent Akodon cursor. Genet Mol Biol 33:785–789
Corander J, Marttinen P, Sirén J, Tang J (2008) Enhanced Bayesian modelling in BAPS software for learning genetic structures of populations. BMC Bioinform 9:539
Costa LP (2003) The historical bridge between the Amazon and the Atlantic Forest of Brazil: a study of molecular phylogeography with small mammals. J Biogeogr 30:71–86
Costa L, R Leite YL, Da Fonseca GA, Da Fonseca MT (2000) Biogeography of South American forest mammals: endemism and diversity in the Atlantic Forest. Biotropica 32:872–881
De Mello Martins F (2011) Historical biogeography of the Brazilian Atlantic forest and the Carnaval-Moritz model of Pleistocene refugia: what do phylogeographical studies tell us? Biol J Linn Soc 104:499–509
D’Horta FM, Cabanne GS, Meyer D, Miyaki CY (2011) The genetic effects of Late Quaternary climatic changes over a tropical latitudinal gradient: diversification of an Atlantic Forest passerine. Mol Ecol 20:1923–1935
Drummond AJ, Rambaut A (2007) BEAST: Bayesian evolutionary analysis by sampling trees. BMC Evol Biol 7:214
Earl DA, VonHoldt BM (2012) Structure harvester: a website and program for visualizing structure output and implementing the Evanno method. Conserv Genet Resour 4:359–361
Elith J, Leathwick JR (2009) Species distribution models: ecological explanation and prediction across space and time. Annu Rev Ecol Evol Syst 40:677–697
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14:2611–2620
Falush D, Stephens M, Pritchard JK (2003) Inference of population structure using multilocus genotype data: Linked loci and correlated allele frequencies. Genetics 164:1567–1587
Fitzpatrick SW, Brasileiro CA, Haddad CFB, Zamudio KR (2009) Geographical variation in genetic structure of an Atlantic Coastal Forest frog reveals regional differences in habitat stability. Mol Ecol 18:2877–2896
Flot JF (2010) Seqphase: a web tool for interconverting phase input/output files and fasta sequence alignments. Mol Ecol Resour 10:162–166
Gardner TA (2001) Declining amphibian populations: a global phenomenon in conversation biology. Anim Biodivers Conserv 24:25–44
Gascon C, Malcolm JR, Patton JL, da Silva MNF, Bogart JP, Lougheed SC et al. (2000) Riverine barriers and the geographic distribution of Amazonian species. Proc Natl Acad Sci USA 97:13672–13677
Goldberg CS, Waits LP (2010) Comparative landscape genetics of two pond-breeding amphibian species in a highly modified agricultural landscape. Mol Ecol 19:3650–3663
Grazziotin FG, Monzel M, Echeverrigaray S, Bonatto SL (2006) Phylogeography of the Bothrops jararaca complex (Serpentes: Viperidae): past fragmentation and island colonization in the Brazilian Atlantic Forest. Mol Ecol 15:3969–3982
Guarnizo CE, Paz A, Muñoz-Ortiz A, Flechas SV, Méndez-Narváez J, Crawford AJ (2015) DNA barcoding survey of anurans across the Eastern Cordillera of Colombia and the impact of the Andes on cryptic diversity. PLoS ONE 10:e0127312
Guayasamin JM, Castroviejo-Fisher S, Ayarzagüena J, Trueb L, Vilà C (2008) Phylogenetic relationships of glassfrogs (Centrolenidae) based on mitochondrial and nuclear genes. Mol Phylogenet Evol 48:574–595
Guayasamin JM, Castroviejo-Fisher S, Trueb L, Ayarzagüena J, Rada M, Vilà C (2009) Phylogenetic systematics of Glassfrogs (Amphibia: Centrolenidae) and their sister taxon Allophryne ruthveni. Zootaxa 2100:1–97
Guillot G, Rousset F (2013) Dismantling the mantel tests. Methods Ecol Evol 4:336–443
Gutiérrez-Rodríguez J, Gonçalves J, Civantos E, Martínez-Solano I (2017) Comparative landscape genetics of pond-breeding amphibians in Mediterranean temporal wetlands: the positive role of structural heterogeneity in promoting gene flow. Mol Ecol 26:5407–5420
Harmon LJ, Glor RE (2010) Poor statistical performance of the Mantel test in phylogenetic comparative analyses. Evolution 64:2173–2178
Heled J, Drummond AJ (2009) Bayesian inference of species trees from multilocus data. Mol Biol Evol 27:570–580
Hewitt GM (2000) The genetic legacy of the quarternary ice ages. Nature 405:907–913
Hewitt GM (2004) The structure of biodiversity—insights from molecular phylogeography. Front Zool 1:1–16
Heyer WR (1985) Taxonomic and natural history notes on frogs of the genus Centrolenella (Amphibia: Centrolenidae) from southeastern Brazil and adjacent Argentina. Pap Avulsos Zool 36:1–21
Hijmans RJ, Cameron SE, Parra JL, Jones PG, Jarvis A (2005) Very high resolution interpolated climate surfaces for global land areas. Int J Climatol 25:1965–1978
IUCN (2017) IUCN red list of threatened species. Version 2017.3. http://www.iucnredlist.org
Jakobsson M, Rosenberg NA (2007) CLUMPP: a cluster matching and permutation program for dealing with label switching and multimodality in analysis of population structure. Bioinformatics 23:1801–1806
Jarvis A, Reuter HI, Nelson A, Guevara E (2008) Hole-filled SRTM for the globe Version 4. CGIAR-CSI SRTM 90mDatabase http://srtm.csi.cgiar.org
Kalinowski ST (2011) The computer program structure does not reliably identify the main genetic clusters within species: simulations and implications for human population structure. Heredity 106:625–632
Knowles LL (2000) Tests of Pleistocene speciation in montane grasshoppers (genus Melanoplus) from the sky islands of western North America. Evolution 54:1337–1348
Knowles LL (2001) Did the Pleistocene glaciations promote divergence? Tests of explicit refugial models in montane grasshopprers. Mol Ecol 10:691–701
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33:1870–1874
Lanfear R, Calcott B, Ho SYW, Guindon S (2012) PartitionFinder: combined selection of partitioning schemes and substitution models for phylogenetic analyses. Mol Biol Evol 29:1695–1701
Legendre P, Lapointe FJ, Casgrain P (1994) Modeling brain evolution from behavior—a permutational regression approach. Evolution 48:1487–1499
Lehner B, Verdin K, Jarvis A (2008) New global hydrography derived from spaceborne elevation data. Eos 89:93–94
Leite YLR, Costa LP, Loss AC, Rocha RG, Batalha-Filho H, Bastos AC et al. (2016) Neotropical forest expansion during the last glacial period challenges refuge hypothesis. Proc Natl Acad Sci 113:1008–1013
Lyra ML, Haddad CFB, Azeredo-Espin AML (2017) Meeting the challenge of DNA barcoding amphibians from Neotropics: Polymerase chain reaction optimization and new COI primers. Mol Ecol Resour 17:966–980
Manly BFJ (1986) Randomization and regression methods for testing for associations with geographical, environmental and biological distances between populations. Res Popul Ecol 28:201–218
Manly BFJ (1991). Randomization and Monte Carlo Methods in Biology. Chapman & Hall/CRC, New York
Manthey JD, Moyle RG (2015) Isolation by environment in white-breasted nuthatches (Sitta carolinensis) of the Madrean Archipelago sky islands: a landscape genomics approach. Mol Ecol 24:3628–3638
Miller S, Dykes D, Polesky H (1988) A simple salting out procedure for extracting DNA from human nucleated cells. Nucleic Acids Res 16:1135–1141
Muscarella R, Galante PJ, Soley-Guardia M, Boria RA, Kass JM, Uriarte M et al. (2014) ENMeval: an R package for conducting spatially independent evaluations and estimating optimal model complexity for Maxent ecological niche models. Methods Ecol Evol 5:1198–1205
Nielson M, Lohman K, Sullivan J (2001) Phylogeography of the tailed frog (Ascaphus Truei): implications for the biogeography of the Pacific Northwest. Evolution 55:147–160
Oliveira EF, Martinez PA, São-Pedro VA, Gehara M, Burbrink FT, Mesquita DO et al. (2018) Climatic suitability, isolation by distance and river resistance explain genetic variation in a Brazilian whiptail lizard. Heredity 120:251–265
Ortego J, Bonal R, Muñoz A, Espelta JM (2015) Living on the edge: the role of geography and environment in structuring genetic variation in the southernmost populations of a tropical oak. Plant Biol 17:676–683
Pearson RG, Raxworthy CJ, Nakamura M, Peterson AT (2007) Predicting species distributions from small numbers of occurrence records: a test case using cryptic geckos in Madagascar. J Biogeogr 34:102–117
Pellegrino KCM, Rodrigues M, Waite AN, Morando M, Yassuda YY, Sites JWJ (2005) Phylogeography and species limits in the Gymnodactylus darwinii complex (Gekkonidae, Squamata): genetic structure coincides with river systems in the Brazilian Atlantic Forest. Biol J Linn Soc 85:13–26
Phillips SJ, Anderson RP, Schapire RE (2006) Maximum entropy modeling of species geographic distributions. Ecol Modell 190:231–259
Phillips SJ, Dudík M (2008) Modeling of species distributions with Maxent: new extensions and a comprehensive evaluation. Ecography 31:161–175
Pons J, Barraclough TG, Gomez-Zurita J, Cardoso A, Duran DP, Hazell S et al. (2006) Sequence-based species delimitation for the DNA taxonomy of undescribed insects. Syst Biol 55:595–609
Pounds JA, Crump ML (1994) Amphibian declines and climate disturbance: the case of the golden toad and the harlequin frog. Conserv Biol 8:72–85
Pounds JA, Fogden MPL, Campbell JH (1999) Biological response to climate change on a tropical mountain. Nature 398:611–615
Pritchard JK, Stephans M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Puechmaille SJ (2016) The program structure does not reliably recover the correct population structure when sampling is uneven: subsampling and new estimators alleviate the problem. Mol Ecol Resour 16:608–627
Rambaut A, Suchard MA, Xie D, Drummond AJ (2014) Tracer v1.6. http://beast.bio.ed.ac.uk/Tracer
Raposo do Amaral F, Edwards SV, Pie MR, Jennings WB, Svensson-Coelho M, d’Horta FM et al. (2016) The “Atlantis Forest hypothesis” does not explain Atlantic Forest phylogeography. Proc Natl Acad Sci 113:201602213
Ribas CC, Aleixo A, Nogueira ACR, Miyaki CY, Cracraft J (2012) A palaeobiogeographic model for biotic diversification within Amazonia over the past three million years. Proc R Soc B Biol Sci 279:681–9
Ribeiro AC (2006) Tectonic history and the biogeography of the freshwater fishes from the coastal drainages of eastern Brazil: An example of faunal evolution associated with a divergent continental margin. Neotrop Ichthyol 4:225–246
Ribeiro MC, Metzger JP, Martensen AC, Ponzoni FJ, Hirota MM (2009) The Brazilian Atlantic Forest: how much is left, and how is the remaining forest distributed? Implications for conservation. Biol Conserv 142:1141–1153
Riccomini C, Grohmann CH, Sant’Anna LG, Silvio T, Hiruma (2010) A captura das cabeceiras do Rio Tietê pelo Rio Paraíba do Sul. In: Modenesi-Gauttieri MC, Bartorelli A, Carneiro VM-N, Ré CD, Lisboa MBDAL (eds) A Obra de Aziz Nacib Ab’ Sáber. Beca, São Paulo, p 157–169
Rodrigues MT, Bertolotto CEV, Amaro RC, Yonenaga-Yassuda Y, Freire EMX, Pellegrino KCM (2014) Molecular phylogeny, species limits, and biogeography of the Brazilian endemic lizard genus Enyalius (Squamata: Leiosauridae): an example of the historical relationship between Atlantic Forests and Amazonia. Mol Phylogenet Evol 81:137–46
Singarayer JS, Valdes PJ (2010) High-latitude climate sensitivity to ice-sheet forcing over the last 120 kyr. Quat Sci Rev 29:43–55
Smouse PE, Long JC, Sokal RR (1986) Multiple regression and correlation extensions of the Mantel test of matrix correspondence. Syst Zool 35:627–632
Stephens M, Scheet P (2005) Accounting for decay of linkage disequilibrium in haplotype inference and missing-data imputation. Am J Hum Genet 76:449–462
Stephens M, Smith NJ, Donnelly P (2001) A new statistical method for haplotype reconstruction from population data. Am J Hum Genet 68:978–989
Thomaz AT, Malabarba LR, Bonatto SL, Knowles LL (2015) Testing the effect of palaeodrainages versus habitat stability on genetic divergence in riverine systems: study of a Neotropical fish of the Brazilian coastal Atlantic Forest. J Biogeogr 42:2389–2401
Thomé MTC, Zamudio KR, Giovanelli JGR, Haddad CFB, Baldissera FA, Alexandrino J (2010) Phylogeography of endemic toads and post-Pliocene persistence of the Brazilian Atlantic Forest. Mol Phylogenet Evol 55:1018–1031
Thomé MTC, Zamudio KR, Haddad CFB, Alexandrino J (2014) Barriers, rather than refugia, underlie the origin of diversity in toads endemic to the Brazilian Atlantic Forest. Mol Ecol 23:6152–6164
van der Meijden A, Vences M, Hoegg S, Boistel R, Channing A, Meyer A (2007) Nuclear gene phylogeny of narrow-mouthed toads (Family: Microhylidae) and a discussion of competing hypotheses concerning their biogeographical origins. Mol Phylogenet Evol 44:1017–1030
Wallace AR (1852) On the monkeys of the Amazon. Proc Zool Soc Lond 20:107–110
Wang IJ (2013) Examining the full effects of landscape heterogeneity on spatial genetic variation: a multiple matrix regression approach for quantifying geographic and ecological isolation. Evolution 67:3403–3411
Wang IJ, Glor RE, Losos JB (2013) Quantifying the roles of ecology and geography in spatial genetic divergence. Ecol Lett 16:175–182
Weir JT, Schluter D (2008) Calibrating the avian molecular clock. Mol Ecol 17:2321–2328
Zamborlini Saiter F, Brown JL, Thomas WW, de Oliveira-Filho AT, Carnaval AC (2016) Environmental correlates of floristic regions and plant turnover in the Atlantic Forest hotspot. J Biogeogr 43:2322–2331
Acknowledgements
Work by AP is co-funded by a Fulbright-Colciencias fellowship and FAPESP (BIOTA, 2013/50297-0), NSF (DEB 1343578), and NASA, through the Dimensions of Biodiversity Program. This study also profited from interactions enabled by the NESCent Montane Biodiversity Working Group and used data generated through National Science Foundation awards to A. Carnaval (DEB-1035184, DEB-1120487). CFBH is grateful to São Paulo Research Foundation (FAPESP) grants #2013/502297-0, 2013/50741-7, #2014/50342-8, and Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq) for a research fellowship. MTR wishes to thank MTR FAPESP (grants # 2011/50146-6 and 2003/10335-8) and CNPq. MTR is grateful to ICMBIO for granting collection permits. For help during fieldwork we thank R. Recoder, F. Dal Vechio, M. Teixeira Jr., J. Cassimiro, M. Sena, V. Verdade, F. Curcio, D. Pavan. The authors also thank two anonymous reviewers and the subject editor of this paper, for their contributions to the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Rights and permissions
About this article
Cite this article
Paz, A., Spanos, Z., Brown, J.L. et al. Phylogeography of Atlantic Forest glassfrogs (Vitreorana): when geography, climate dynamics and rivers matter. Heredity 122, 545–557 (2019). https://doi.org/10.1038/s41437-018-0155-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41437-018-0155-1
This article is cited by
-
Influence of anthropogenic pressure on the genetic diversity and chromosomal instability of an endangered forest-specialist anuran
Hydrobiologia (2022)
-
From micro- to macroevolution: insights from a Neotropical bromeliad with high population genetic structure adapted to rock outcrops
Heredity (2020)
-
Salinity tolerance explains the contrasting phylogeographic patterns of two swimming crabs species along the tropical western Atlantic
Evolutionary Ecology (2020)