Abstract
Purpose
Family-based cascade screening from index probands is considered an effective way of identifying undiagnosed individuals with familial hypercholesterolemia (FH). The role of genetic testing of the proband in the success of cascade screening for FH is unknown.
Methods
We randomized 240 individuals with a clinical diagnosis of FH to genetic testing for FH (n = 160) or usual care with lipid testing alone (n = 80). The primary study endpoint was the proportion of probands with at least one relative enrolled in the study within one year after the notification of results.
Results
Proband median age was 59 (47–67) and 71% were female. Only 28 (12%) probands succeeded in enrolling a relative. While the genetic testing group had a higher proportion of probands with relatives enrolled (13.1%) compared with the usual care group (8.8%), this difference was not significant (p = 0.40). In subgroup analyses, enrollment of a relative was higher in the pathogenic variant group (22.7%) compared to the no pathogenic variant (9.5%) and usual care groups (8.8%) (p = 0.04).
Conclusion
We observed a low rate of family participation in cascade screening despite repeated recommendations to probands. Compared to usual care, genetic testing did not improve family participation in cascade screening for FH.
Clinical trial number
NCT04526457
Similar content being viewed by others
INTRODUCTION
Familial hypercholesterolemia (FH) is a heritable autosomal dominant condition of elevated low-density lipoprotein cholesterol (LDL-C) and premature cardiovascular disease. With an estimated prevalence of ~1:250 in the general population, it is among the most common monogenic inherited causes of coronary artery disease (CAD).1,2 Despite the availability of highly effective therapies, FH accounts for 1–8% of nonpremature3,4 and 1–20% of premature CAD cases5,6 worldwide and remains markedly underdiagnosed and undertreated.7 FH meets the World Health Organization criteria for population screening.8 In 2012, the Centers for Disease Control and Prevention (CDC) Office of Public Health Genomics (OPHG) designated FH one of only three adult tier 1 conditions for which family-based cascade screening using DNA testing is recommended.9 Cascade screening involves starting with a proband diagnosed with an autosomal dominant genetic condition and performing systematic tracing and testing of immediate and extended family members to identify additional cases. Cascade screening in FH has the potential to be exceptionally effective, but is very poorly executed, particularly in the United States.10
Recently, consensus guidelines for genetic testing for FH were proposed11 and one rationale for more active consideration of genetic testing was that it could potentially facilitate cascade screening. Evidence from national FH screening programs in Europe has shown that a comprehensive approach to FH inclusive of genetic testing and cascade screening improves the identification of FH, treatment uptake, adherence, and cardiovascular morbidity12,13,14 while being highly cost-effective.15 However, the role of genetic testing in cascade screening for FH has never been formally studied. As there is currently no standardized approach to cascade screening for FH in the United States, the US health-care system provides an optimal setting to address questions regarding interventions that may improve cascade screening for FH. Developments that contribute to the timeliness of this work include the approval of novel classes of LDL-lowering therapies for patients with FH; the emergence of robust electronic health records (EHRs) to facilitate case identification; the existence of a national FH registry, Cascade Screening for Awareness and Detection of FH (CASCADE FH);16 and rapidly declining costs of DNA sequencing. We hypothesized that genetic testing of the proband would enhance the efficacy of cascade screening for FH, and performed a randomized controlled trial to compare the effect of genetic testing to usual care on family participation in cascade screening for FH.
MATERIALS AND METHODS
Study design and subject recruitment
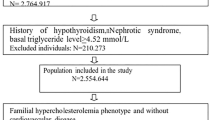
Our trial was titled “Is Family screening Improved by Genetic Testing in FH” (I FIGhT FH). It was conducted within the University of Pennsylvania Health System (UPHS) from November 2014 to April 2017. An overview of the study design is shown in Figure S1. To be eligible for the study, patients had to be 18 years or older, have at least one documented LDL-C ≥ 220 mg/dl in the preceding 5 years without evidence of a secondary cause of hypercholesterolemia. The threshold of 220 mg/dL was chosen as it has been used to identify adults (age ≥ 20) in the general population very likely to have FH by the US Make Early Diagnosis to Prevent Early Deaths (MEDPED) criteria, well above the threshold of ≥190 mg/dL used in other diagnostic criteria for FH.17 Probands were identified by screening the UPHS outpatient clinic patient databases using an EHR query (Fig. 1). A few probands meeting the eligibility criteria were identified from referral by a lipid specialist. Patients were excluded if they had a history of severe hypertriglyceridemia (triglycerides > 400 mg/dl or fibrate use), diagnoses associated with secondary hypercholesterolemia (history of obstructive liver disease, primary biliary cirrhosis, direct bilirubin >1.5 mg/dl, nephrotic syndrome, hypothyroidism, or thyroid stimulating hormone >10 mIU/L), previous genetic testing for FH, fewer than two contactable biological relatives, or were unable to provide informed consent.
Proband procedures
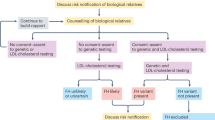
At the baseline visit, probands were asked to complete a health questionnaire, provide a fasting blood sample, and undergo a targeted physical examination for signs of FH (xanthomas and corneal arcus). They were then randomized 2:1 to genetic testing plus lipid testing (“genetic testing”) or lipid testing alone (“usual care”). Randomization was achieved using a random number generator in Microsoft Excel to produce numbers between 0 and 1. Approximately six weeks after enrollment, all probands were called by a certified genetic counselor to discuss their lipid and genetic testing (genetic testing arm) results, review their family history, and discuss the approach to cascade screening (Figure S1). Probands were asked to request that relatives contact the study team or to solicit permission from relatives for the study team to contact them directly about being screened.
Study questionnaires
Probands were asked to complete questionnaires at baseline, week 26 and week 58. These questionnaires were administered by paper or web-based survey (baseline and week 26) or by telephone interview (week 58). See Supplemental Methods for further details. The week 26 survey also provided an opportunity for the study team to encourage family member recruitment.
Family member procedures
First, second- and third-degree relatives that contacted the study team were able to enroll for up to one year after the proband’s genetic counseling call. Relatives had to be 10 years or older, able to give informed consent or assent to participate in the study and willing/able to participate in the study. After initial contact, relatives were invited for a study visit or sent a testing kit so that blood could be drawn at a local site. Family members of probands in whom a causative FH pathogenic variant had been identified underwent both genetic and lipid testing. Family members of probands in whom either a pathogenic variant had not been identified or who did not undergo genetic testing underwent only lipid testing. In these cases, a clinical diagnosis of FH was ascertained using the Make Early Diagnosis To Prevent Early Deaths (MEDPED) criteria. Within 6 weeks of enrollment, genetic and lipid test results were communicated to relatives by a genetic counselor or a member of the study team.
Genetic analysis
DNA was isolated from whole blood as previously described.18 Participant’s DNA was sequenced using SEQPRO LIPO RS (Progenika Biopharma, Derio, Spain), a next-generation sequencing kit designed to detect variants in LDLR, APOB, PCSK9, and LDLRAP1.19 Exons and exon–intron boundaries of LDLR (18 exons), PCSK9 (12 exons), and LDLRAP1 (9 exons) were pyrosequenced (454 Life Science, Roche). Regions in exons 26 and 29 (between the nucleotides 10,416 and 10,779 for exon 26 and between nucleotides 12,987 and 13,221 for exon 29) of the APOB gene were pyrosequenced. Multiplex ligation-dependent probe amplification (MLPA) was performed on specimens in which ambiguous results were obtained to detect large deletions or duplications. Variants identified were labeled by Progenika as pathogenic, probable or possibly pathogenic, or of unknown significance or hypocholesterolemic based on functional data within proprietary and literature-based reference databases.20 Variants established as nonpathogenic by Progenika were not reported—this included APOB and PCSK9 variants of unknown pathogenicity.20 Independently, the study team assessed for pathogenicity all LDLR variants labeled as of unknown significance by Progenika (Supplemental Methods).
Outcomes
The primary study endpoint was the proportion of probands with at least one relative enrolled within one year of the call from the genetic counselor in the genetic testing group compared to the usual care group. Enrollment was defined by the return of a completed test kit during the study period. Secondary endpoints included (1) the number of relatives enrolled within 52 weeks of the genetic counseling call and (2) the number of enrolled relatives diagnosed with FH, expressed as the new case per index case ratio (NCIC, relatives diagnosed with FH/total number of index cases) as previously described.21 Exploratory subgroup analyses were conducted stratifying the cohort by randomization/genetic test result (pathogenic variant, no pathogenic variant, and usual care). Further exploratory analyses compared probands’ perceptions about their high cholesterol diagnosis, its heritability, control and familial risk at baseline and at 26 weeks after enrollment.
Statistical analyses
There were no previous studies on which to base a sample size calculation. Baseline characteristics are expressed as frequencies and percentages for categorical variables and median (interquartile range) for continuous variables unless otherwise stated. For the study primary and secondary outcomes, categorical and continuous variables were compared by Z-test and likelihood ratio based on logistic and linear regression models, respectively. Otherwise, categorical variables were compared using Mann–Whitney tests for the comparison of two group and Kruskal–Wallis test for comparisons of three or more groups. Significance level was set at a p value of <0.05, unless otherwise specified. Statistical analyses were performed using GraphPad Prism version 7.00 for Mac (GraphPad Software, La Jolla, CA, USA, www.graphpad.com).
RESULTS
Proband characteristics and genetic test results
Of the 240 probands enrolled in the study, 160 were randomized to receive genetic testing and 80 to usual care (Fig. 1). Baseline clinical and demographic characteristics of the participants are presented in Table 1. The cohort included 170 women (71% of probands) with a median (IQR) age of 59 (47–67) years. Clinical characteristics were well balanced between probands assigned to genetic testing and those assigned to usual care (Table 1). Targeted genetic sequencing of LDLR, APOB, PCSK9, and LDLRAP1 identified potentially disease-causing variants, both known pathogenic (Table S1) as well as variants of unknown significance (VUS) (Table S2), in 44 probands (27.5%) in the genetic testing arm. Demographic and clinical characteristics of probands with pathogenic variants were largely comparable to those of probands with VUS (Table S3); however, compared to probands with VUS, those with pathogenic variants were younger (30 vs. 35 years, p = 0.02) and more likely to have a family history of hypercholesterolemia (87% vs. 50%, p = 0.01). Compared to those in the no pathogenic variant and usual care groups, probands in the pathogenic variant group had higher LDL-C levels and were younger, more likely to have tendon xanthomas, and a history of FH (Table S4).
Primary and secondary endpoints
Despite active attempts to encourage family-based cascade screening, only 38 (15.8%) probands overall had at least one family member contact the study team over the study period and of these, only 28 probands (11.7%) had at least one family member actually enroll in the study (Table 2). Of note, no family members were enrolled before the call from the genetic counselor in week 6. Overall, a total of 43 family members (0.2 family members per proband enrolled in the study) were enrolled over the study period (Table 2). Family member characteristics are summarized in Table S5. Family members enrolled were mostly first-degree relatives (83.7%), younger (33 vs. 59, p < 0.01), female (67.4%), and white (83.7%). The proportion of probands with enrolled relatives was 13.1% in the genetic testing arm and 8.8% in the usual care arm (p = 0.32) (Figure S2, Table 2). The number of relatives enrolled per proband in the genetic testing arm was 0.2, compared to 0.1 in usual care arm (p = 0.14) (Fig. 2, Table 2). The new cases per index case (NCIC) ratio was also comparable between the two groups (0.1 vs. 0.1, p = 0.27) (Figure S2, Table 2).
Responses to question 21 from the “Is Family screening Improved by Genetic Testing in FH” (I FIGhT FH) baseline questionnaire, question 15 from the I FIGhT FH week 26 questionnaire (genetic testing) and question 14 from the I FIGhT FH week 26 questionnaire (usual care) are presented. Only 122 probands completed follow-up questionnaires. For this analysis, only baseline responses from these probands are represented here. FH familial hypercholesterolemia.
By contrast, in subgroup analyses stratified by genetic test result, more probands in the pathogenic variant group had relatives enrolled (22.7%) compared to those in the no pathogenic variant (9.5%) and usual care (8.8%) groups (pathogenic variant vs. no pathogenic variant vs. usual care, p = 0.04) (Table 2). This trend persisted but was nonsignificant in a sensitivity analysis that excluded probands with VUS from the pathogenic variant group (pathogenic variant vs. no pathogenic variant vs. usual care, 25% vs. 10.6% vs. 8.8%; p = 0.06) (Table S6). The number of relatives enrolled per proband was also higher in the pathogenic variant group (0.4) compared to the no pathogenic variant (0.2) and usual care (0.1) groups (p = 0.02) (Table 2). The NCIC ratio in the pathogenic variant group was three times that in the usual care group, although this difference was not significant (pathogenic variant vs. no pathogenic variant vs. usual care, 0.2 vs. 0.10 vs. 0.06; p = 0.08) (Table 2).
Impact of the study on proband perceptions
Responses to questions in the baseline and week 26 follow-up questionnaires from probands that participated in both are summarized in Fig. 2, Table 3 and Supplemental S7–S9. Only 51% (122/240) of the initial cohort participated in the week 26 survey. At baseline, the majority of probands in the three groups (pathogenic variant, no pathogenic variant, and usual care) agreed that genetic factors played a causative role in their high LDL-C, that relatives were at risk of having a similar condition, and that their cholesterol could be effectively controlled with medications. More probands in the pathogenic variant group agreed that genetics played a role in their high LDL-C at baseline (p < 0.01) compared to those in the no pathogenic variant and usual care groups. Genetic testing alone had no or only a marginal effect on these perceptions at follow-up (Table 3, Table S7). However, in subgroup analyses stratified by genetic test result, compared to participants in the pathogenic variant and usual care groups, fewer subjects in the no pathogenic variant group agreed that their high LDL-C was due to genetics and that relatives were at risk of having high cholesterol at follow-up (both p < 0.01) (Table 3, Table S8). Consistent with this, probands in the pathogenic variant and usual care groups felt more strongly about the heritability of their high cholesterol compared to participants in the no pathogenic variant group at follow-up (68% vs. 47% vs. 20%, p < 0.01) (Table S8).
Family member responses to study invitation and unmeasured study impact
By follow-up, 97% of probands reported that they had shared their diagnosis of high cholesterol with an average of five relatives (Table S9). This did not differ by randomization or genetic test result. Reasons probands did not share their diagnosis with relatives are summarized in Fig. 2. Notably, the responses to this question were similar at baseline and at follow-up. In a third of the responses, probands indicated that they had shared their diagnosis with all of their relatives prior to the study. The second most common reasons were emotional/social (21% at baseline, 14% at follow-up) and geographical distance (10% at baseline, 11% at follow-up) (Fig. 2). Privacy concerns were trivial (1%) in this cohort at follow-up. Of the initial cohort, 29% (70/240) participated in the week 58 survey. From this group, 81% (57) had invited relatives and 57% (40) reported having relatives in the study. By far the most common reasons family members declined to participate in the study were already knowing their cholesterol level (25%) and being on treatment for high cholesterol (18%) (Figure S3a). Interestingly, 13% of respondents indicated that family members did not want to participate in research. When asked to share whether the study had impacted the lives of family members outside of the study, more than half of the responses indicated that family members had gotten their cholesterol checked as a result of the study (Figure S3b).
DISCUSSION
Family-based cascade screening for FH is believed to be very poorly executed in the United States. Furthermore, genetic testing for FH is not yet clinically established as standard of care. In this randomized controlled trial of any intervention to promote cascade screening in FH, we tested the hypothesis that genetic testing of probands with FH would promote more effective cascade screening of family members. We found, in a sample of 240 probands with clinical FH, that only 0.2 relatives per proband were enrolled over the study period, despite systematic repeated efforts to encourage family outreach through probands. Further, genetic testing overall did not improve family participation in cascade screening or identification of new FH cases compared to usual care over a one year period. Interestingly, however, in a subgroup analysis of probands stratified by genetic test result, the identification of an FH-causing pathogenic variant was associated with a significant increase in participation of family members in cascade screening, with a two to threefold higher enrollment in the pathogenic variant group. By contrast, relative participation was comparable in the no pathogenic variant and usual care groups, suggesting that a negative genetic result did not depress family enrollment. Taken together, these findings suggest that a positive genetic test result may improve the efficiency of cascade screening but genetic testing alone, at least in a cohort with a relatively low rate of positive genetic findings, does not.
In a systematic review of cascade screening programs for FH, cascade screening was shown to be six times more effective in identifying new FH cases when conducted using genetic testing compared to lipid testing alone.21 However, the studies that used genetic testing performed cascade screening only in individuals found to have a pathogenic variant and no studies with a head-to-head comparison were included. As such, these findings suggest that a positive genetic result may improve the efficiency of cascade screening for FH, but they do not provide evidence to support the role of genetic testing overall. In the present study, we show that genetic testing itself did not significantly improve family engagement in cascade screening, but that a positive genetic result did do so. This finding is in line with the concept that knowledge of a causal FH variant enhances proband and family member participation in screening. It should also be noted that a negative genetic test result did not result in poorer engagement in cascade screening relative to usual care. The neutral effect of genetic testing in this study may have occurred because individuals in whom a pathogenic variant was identified comprised <30% of the genetic testing group. In a setting where the pretest probability of finding a positive result on genetic testing is considerably higher, genetic testing might be expected to enhance cascade screening.
A considerable body of qualitative evidence has shown that positive genetic confirmation of FH provides reassurance, increases confidence in the role of lipid-lowering medicines, motivates initiation and adherence to lipid-lowering treatments,22,23,24 and promotes greater awareness of familial risk, over and above clinical diagnosis alone.24,25 In the Genetic Risk Assessment for FH trial (GRAFT), investigators examined the impact of genetic testing on perceptions of control over hypercholesterolemia and adherence to risk-reducing behavior among participants clinically diagnosed with FH recruited from lipid clinics in England and randomized to genetic testing and lipid testing or lipid testing alone.23 They found that the identification of a pathogenic variant was associated with stronger perceptions about the impact of risk-reducing behaviors such as taking medication. Thus, we hypothesize that genetic confirmation of FH would improve relative participation in cascade screening for FH by changing perceptions about the heritability of high cholesterol and familial risk. Further studies in FH populations with higher probability of positive genetic test results are needed to examine this hypothesis.
Our findings highlight the importance of careful communication of “negative” or equivocal genetic test results in the FH cascade screening process. We found that the failure to identify a pathogenic variant affected perceptions about the heritability of high cholesterol, personal and familial risk, and motivation to engage relatives in screening. Up to 70% of patients clinically diagnosed with FH do not have a pathogenic variant identified in genetic testing,26,27,28 and the majority of variants identified in patients with a clinical diagnosis of FH are labeled as a VUS. VUS may later be shown to be pathogenic.29 In this study, subjects with VUS were included in the pathogenic variant group and were clinically similar to those with established pathogenic variants. Further, we were able to demonstrate that almost all of the variants labeled as a VUS were likely to be pathogenic adopting an approach that has been used previously.30 Our findings call for careful discussion about the handling of patients found to have a VUS in the FH cascade screening process. While overcalling VUS is undesirable,29 in patients with a clinical diagnosis of FH, many of these are likely pathogenic and suboptimal communication about a VUS result could adversely impact attitudes about the genetic basis of elevated LDL-C and the need for cascade screening.
Finally, despite repeated encouragement of the FH probands, we found extremely low participation by family members in cascade screening. One explanation for this might be the research study format of this project. Our findings suggest that this may have deterred some family members from participation. Indeed, probands reported that family members acted on the information they received from the proband and got their cholesterol checked, sought medical care, or changed their lifestyle outside of the study. Another important explanation to consider is that the restriction on directly approaching relatives due to privacy regulations, with the requirement that the initial contact be made by probands, likely posed a significant barrier to relative engagement in this study. Direct contact of relatives is the widely preferred method of contacting relatives for cascade screening. In the Dutch national cascade screening program, health-care workers asked index patients for consent to contact their relatives and, after consent was obtained, field workers were dispatched to homes to enroll relatives.31 Using this strategy, ~23 relatives/proband were enrolled during the first 5 years of the Dutch program.12 After replacement of direct contact with indirect contact of relatives in this program, relative participation in screening declined ~8-fold.31 In a US study that used an online direct approach to recruit first-degree relatives in cascade screening for familial cancers, over a one-year period 2,280 first-degree relatives of 1,101 index subjects (2 relatives per proband) were recruited.32 One reason indirect contact might fail is proband reticence to inform family members about screening.24,33 In our study, this was most commonly attributed to feelings of emotional/social and geographical distance, as has previously been suggested.25 Direct contact of relatives is the preferred method for conducting cascade screening—we suggest that probands be offered the option of either direct or indirect contact with relatives at the time of initial screening.34
Several limitations of this study should be noted. First, our sample size was not based on a power calculation as no prior relevant empirical data was available for this, and given the poor response overall in cascade screening the study was relatively underpowered. The relatively low genetic yield on sequencing of the probands further limited our power and, given our finding of greater participation in probands with a positive genetic result, likely reduced the overall apparent efficacy of genetic testing. A post hoc power analysis using the observed family enrollment rates in the genetic testing and usual care groups suggests that 827 individuals would be needed in each group (for 1:1 enrollment) for the study to have 80% power to detect a ~4% difference at an alpha level of 0.05. Second, extended family size (beyond third-degree relatives) was not taken into account in the primary and secondary outcomes—this may have influenced the absolute number of individuals identified in each arm of the study. However, the majority of family members enrolled were first-degree relatives, family size was comparable between the two arms of the study and, post hoc analyses of our primary outcome that adjusted for family size were consistent with our primary findings. Third, men and individuals from nonwhite racial backgrounds were underrepresented in the sample of probands recruited, so the generalizability of these results will require confirmation. Fourth, family members may have pursued genetic and/or lipid testing outside of this study. However, we have no evidence to suggest this would have differentially impacted the two arms of the study and affected our primary outcome. Fifth, given the nature of the study, it was not considered for registration a priori, but was registered post hoc upon request. We will make our institutional review board (IRB) protocol and subsequent amendments available upon request. Finally, this study provides only a partial representation of the impact of cascade screening as it focuses only on family member participation, without considering cost-effectiveness of screening and uptake of therapy—both key indices of cascade screening success. We know of one study already underway in the United States (NCT03640234) that has set out to explore the broader impact of genetic testing, including its impact on relative recruitment, cost, and psychosocial factors.
In conclusion, in this randomized study of the effect of genetic testing on cascade screening for FH, we found that genetic testing overall did not improve family participation in cascade screening or the number of FH cases identified among relatives. However, we show that the identification of a FH-causing pathogenic variant in the proband may be associated with an increase in family member participation in FH cascade screening. Overall, we found that relying solely on index patients for family outreach led to very poor family member participation in cascade screening, and gained insight into the reasons for lack of participation in cascade screening. Further systematic work is needed to improve the efficiency of cascade screening for FH.
Data availability
Data including the study protocol and de-identified clinical data sets will be made available upon request.
References
Abul-Husn, N. S. et al. Genetic identification of familial hypercholesterolemia within a single U.S. health care system. Science. https://doi.org/10.1126/science.aaf7000 (2016).
Benn, M., Watts, G. F., Tybjaerg-Hansen, A. & Nordestgaard, B. G. Familial hypercholesterolemia in the danish general population: prevalence, coronary artery disease, and cholesterol-lowering medication. J. Clin. Endocrinol. Metab. 97, 3956–3964 (2012).
Mortensen, M. B., Kulenovic, I., Klausen, I. C. & Falk. E. Familial hypercholesterolemia among unselected contemporary patients presenting with first myocardial infarction: prevalence, risk factor burden, and impact on age at presentation. J. Clin. Lipidol. https://doi.org/10.1016/j.jacl.2016.06.002 (2016).
De Backer, G. et al. Prevalence and management of familial hypercholesterolaemia in coronary patients: an analysis of EUROASPIRE IV, a study of the European Society of Cardiology. Atherosclerosis. 241, 169–175 (2015).
Wald, D. S., Bangash, F. A. & Bestwick, J. P. Prevalence of DNA-confirmed familial hypercholesterolaemia in young patients with myocardial infarction. Eur. J. Intern. Med 26, 127–130 (2015).
Pang, J. et al. Frequency of familial hypercholesterolemia in patients with early-onset coronary artery disease admitted to a coronary care unit. J. Clin. Lipidol. 9, 703–708 (2015).
Nordestgaard, B. G. B. G. et al. Familial hypercholesterolaemia is underdiagnosed and undertreated in the general population: guidance for clinicians to prevent coronary heart disease. Eur. Heart J. 34, 3478–3490 (2013).
WHO Human Genetics Programme. Familial hypercholesterolaemia (FH): report of a second WHO consultation, Geneva, 4 September 1998. World Health Organization (1999).
US Centers for Disease Control and Prevention. Office of Public Health Genomics. Genomic tests by levels of evidence. http://www.cdc.gov/genomics/gtesting/file/print/tier.pdf (2016).
Knowles, J. W., Rader, D. J. & Khoury, M. J. Cascade screening for familial hypercholesterolemia and the use of genetic testing. JAMA. 318, 381–382 (2017).
Sturm, A. C. et al. Clinical genetic testing for familial hypercholesterolemia: JACC Scientific Expert Panel. J. Am. Coll. Cardiol. 72, 662–680 (2018).
Umans-Eckenhausen, M. A. W., Defesche, J. C., Sijbrands, E. J. G., Scheerder, R. L. J. M. & Kastelein, J. J. P. Review of first 5 years of screening for familial hypercholesterolaemia in the Netherlands. Lancet. 357, 165–168 (2001).
Umans-Eckenhausen, Ma. W., Defesche, J. C., van Dam, M. J. & Kastelein, J. J. P. Long-term compliance with lipid-lowering medication after genetic screening for familial hypercholesterolemia. Arch. Intern. Med. 163, 65–68 (2003).
Leren, T. P., Finborud, T. H., Manshaus, T. E., Ose, L. & Berge, K. E. Diagnosis of familial hypercholesterolemia in general practice using clinical diagnostic criteria or genetic testing as part of cascade genetic screening. Community Genet. 11, 26–35 (2008).
Nherera, L., Marks, D., Minhas, R., Thorogood, M. & Humphries, S. E. Probabilistic cost-effectiveness analysis of cascade screening for familial hypercholesterolaemia using alternative diagnostic and identification strategies. Heart. 97, 1175–1181 (2011).
Degoma, E. M. et al. Treatment gaps in adults with heterozygous familial hypercholesterolemia in the United States. Circ. Cardiovasc. Genet. 9, 240–249 (2016).
McGowan, M. P., Hosseini Dehkordi, S. H., Moriarty, P. M. & Duell, P. B. Diagnosis and treatment of heterozygous familial hypercholesterolemia. J. Am. Heart Assoc. https://doi.org/10.1161/JAHA.119.013225 (2019).
Johns, M. B. & Paulus-Thomas, J. E. Purification of human genomic DNA from whole blood using sodium perchlorate in place of phenol. Anal. Biochem. 180, 276–278 (1989).
Maglio, C. et al. Genetic diagnosis of familial hypercholesterolaemia by targeted next-generation sequencing. J. Intern. Med. 276, 396–403 (2014).
GRIFOLS. FH genetic testing. https://www.diagnostic.grifols.com/en/fh-genetic-testing/test-details (2020).
Lee, C., Rivera-Valerio, M., Bangash, H., Prokop, L. & Kullo, I. J. New case detection by cascade testing in familial hypercholesterolemia: a systematic review of the literature. Circ. Genomic Precis. Med. 12, e002723 (2019).
Senior, V., Marteau, T. M. & Weinman, J. Self-reported adherence to cholesterol-lowering medication in patients with familial hypercholesterolaemia: the role of illness perceptions. Cardiovasc. Drugs Ther. 18, 475–481 (2004).
Marteau, T. et al. Psychological impact of genetic testing for familial hypercholesterolemia within a previously aware population: a randomized controlled trial. Am. J. Med. Genet. A 128A, 285–293 (2004).
Hardcastle, S. J. et al. Patients’ perceptions and experiences of familial hypercholesterolemia, cascade genetic screening and treatment. Int. J. Behav. Med. 22, 92–100 (2015).
Hallowell, N. et al. Patients’ experiences and views of cascade screening for familial hypercholesterolemia (FH): a qualitative study. J. Community Genet. 2, 249–257 (2011).
Damgaard, D. et al. The relationship of molecular genetic to clinical diagnosis of familial hypercholesterolemia in a Danish population. Atherosclerosis 180, 155–160 (2005).
Shin, D. G. et al. Clinical features of familial hypercholesterolemia in Korea: predictors of pathogenic mutations and coronary artery disease—a study supported by the Korean Society of Lipidology and Atherosclerosis. Atherosclerosis. 243, 53–58 (2015).
Civeira, F. et al. Comparison of genetic versus clinical diagnosis in familial hypercholesterolemia. Am. J. Cardiol. 102, 1187–93, 1193.e1 (2008).
Chora, J. R., Medeiros, A. M., Alves, A. C. & Bourbon, M. Analysis of publicly available LDLR, APOB, and PCSK9 variants associated with familial hypercholesterolemia: application of ACMG guidelines and implications for familial hypercholesterolemia diagnosis. Genet. Med. 20, 591–598 (2018).
Khera, A. V. et al. Diagnostic yield and clinical utility of sequencing familial hypercholesterolemia genes in patients with severe hypercholesterolemia. J. Am. Coll. Cardiol. 67, 2578–2589 (2016).
Louter, L., Defesche, J. & Roeters van Lennep, J. Cascade screening for familial hypercholesterolemia: practical consequences. Atheroscler. Suppl. 30, 77–85 (2017).
Caswell-Jin, J. L. et al. Cascade genetic testing of relatives for hereditary cancer risk: results of an online initiative. J. Natl. Cancer Inst. 111, djy147 (2019).
Van Maarle, M. C. et al. How disturbing is it to be approached for a genetic cascade screening programme for familial hypercholesterolaemia?: psychological impact and screenees’ views. Community Genet. 4, 244–252 (2001).
Newson, A. J. & Humphries, S. E. Cascade testing in familial hypercholesterolaemia: How should family members be contacted? Eur. J. Hum. Genet. 13, 401–408 (2005).
Acknowledgements
We thank Stephanie DerOhannessian for her tremendous support with participant recruitment and liaising with vendors. We also extend a special thank you to Tracey Sikora, the phenomenal project manager who contributed tremendously to this project from start to finish. This work was funded by the Penn Center for Precision Medicine and The Stacey and Curtis Lane Fund.
Author information
Authors and Affiliations
Contributions
Conceptualization: E.A., E.M.d.G., A.R., K.D., D.J.R. Data curation: E.A., A.R., K.D.. Formal analysis: E.A., D.J.R. Funding Acquisition: E.M.d.G., D.J.R. Investigation: E.A., E.M.d.G., A.R., K.D., M.C., D.J.R. Project administration: E.A., E.M.d.G., A.R., K.D., M.C., D.J.R. Writing—original draft: E.A., E.M.d.G., A.R., K.D., M.C., D.J.R. Writing—review and editing: E.A., D.J.R.
Corresponding author
Ethics declarations
Ethics declaration
The study protocol was approved by the Institutional Review Board of the University of Pennsylvania. Informed consent was required for study participation.
Competing interests
D.J.R. is the Chief Scientific Advisor for the Familial Hypercholesterolemia Foundation and receives no compensation for this role; he also serves on Scientific Advisory Boards for Alnylam, Novartis, Pfizer, and Verve. M.C. has received institutional funding from Akcea, Regeneron and Regenxbio and consulting fees from Amryt. The other authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Ajufo, E., deGoma, E.M., Raper, A. et al. A randomized controlled trial of genetic testing and cascade screening in familial hypercholesterolemia. Genet Med 23, 1697–1704 (2021). https://doi.org/10.1038/s41436-021-01192-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41436-021-01192-z
This article is cited by
-
Family cascade screening for equitable identification of familial hypercholesterolemia: study protocol for a hybrid effectiveness-implementation type III randomized controlled trial
Implementation Science (2024)
-
Cholesterol Screening in Children: Is a Universal Approach Working?
Current Atherosclerosis Reports (2023)