Abstract
We present the first pachyonychia congenita (PC) to involve all ectodermal derivatives and the first recessive KRT17-related PC in total seven members of two consanguineous Pakistani families. This atypical PC is characterized by an unusual combination of pachyonychia, plantar keratoderma, folliculitis, alopecia, sparse eyebrows, dental anomalies and variable acanthosis nigricans of neck, dry skin, palmoplantar hyperhidrosis, recurrent blisters on soles and/or arms, rough sparse hair on scalp and keratosis pilaris. By exome sequencing we detected homozygous KRT17 c.281G>A (p.(Arg94His)) in affected individuals, and linkage mapping indicated a single locus. Heterozygous variants in KRT17 cause PC2 (PC-K17) with main characteristics of pachyonychia, subungual keratosis, palmoplantar keratoderma, hyperhidrosis, oral leukokeratosis and epidermal cysts, or steatocystoma multiplex, both with dominant inheritance. The causative variant has been reported in heterozygous state in a family afflicted with severe steatocystoma multiplex and in a sporadic PC2 case, and thus we also define a third phenotype related to the variant. Both exome sequencing and linkage mapping demonstrated recessive inheritance whereas Sanger sequencing indicated heterozygosity for the causal variant, reiterating caution for simple targeted sequencing for genetic testing. Testing parents for variants found in sibs could uncover recessive inheritance also in other KRT genes.
Similar content being viewed by others
Introduction
Clinical phenotype and inheritance pattern can vary for variants in the same gene. For example, a variant in KRT1 or KRT10 can cause either dominant or recessive epidermolytic hyperkeratosis (OMIM 113800). Alternatively, variants with recessive effect can lead to somewhat different diseases as in GUCY2D-related eye diseases and BMPR1B-related brachydactyly (OMIM 600179, OMIM 603248). It is even possible that heterozygous or biallelic variants in the same gene can cause a variety of different diseases, some with no overlapping features, as in TBC1D24 (OMIM 613577).
Pachyonychia is hypertrophic nail dystrophy resulting in extremely thick and abnormally shaped nails. Pachyonychia congenita (PC) is a very rare genodermatosis that primarily affects the nails and skin. It is characterized by malformed, discolored, and very thick nails due to hypertrophy, as well as painful and highly debilitating palmoplantar keratoderma, follicular keratosis especially on knees and elbows, oral leukokeratosis and a variety of epidermal cysts [1].
PC is known to be an autosomal dominant condition. PC1 (PC-K16), PC2 (PC-K17), PC3 (PC-K6A), and PC4 (PC-K6B) are caused by heterozygous variants in KRT16, KRT17, KRT6A, and KRT6B, respectively. Heterozygous KRT17 variants, besides PC2, can cause steatocystoma multiplex [2,3,4,5,6,7,8,9,10,11,12,13,14]. Currently around 30 disease-causing variants are known.
In two families we define a recessive and atypical PC involving all ectodermal derivatives caused by a homozygous missense KRT17 variant.
Subjects and methods
Families
Families are from nearby towns. Parents of all seven affected individuals are consanguineous (Fig. 1). We examined all affected individuals and their 24 unaffected relatives. We obtained DNA samples of 19 participants.
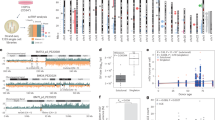
KRT17 c.281G/A genotypes are shown for individuals where DNA samples were available, deduced after evaluating the exome file integrative genome view (IGV), haplotypes and Sanger sequencing results all together. * SNP data and # exome data were generated, ? distant consanguinity, horizontal lines above symbols physical examination was performed, + variant, - reference.
Genetic studies
Exome sequencing was performed for one patient and a relative in each family to find the causal variant (Supplementary Materials). Sanger sequencing was performed to investigate the segregation of the candidate variant. Later, segregation of the disease haplotype was assessed by linkage analysis using SNP genotypes (Fig. 1).
Results
Clinical findings
We clinically investigated all seven affected individuals. The disorder was a combination of features affecting nails, skin, scalp hair, and teeth (Table 1, Supplementary Fig. 1 and Supplementary Table 1). The most conspicuous feature was pachyonychia shared by all affected individuals (Fig. 2). Generally, hands were more involved than feet. All nails of hands and feet were involved, and expression was more severe in the preaxial nails. Most of the affected individuals complained of recurrent blisters on soles and/or arms and rarely on trunk. There was no pain in the soles in any participant. Five patients complained of localized dry skin more marked on the back of hands and feet that becomes severe in winters. Sparse eyebrows and round patches of non-scarring alopecia were evident in all affected individuals. Rough and sparse scalp hair was noted in four. Hair microscopy revealed smooth hair with loose anagen, distorted and elongated bulb and attached root sheath in all patients, but there were no pili torti, nodes/internodes, twists, or loops (Supplementary Fig. 1). After detecting the causal variant, 2 years after the first evaluation, a dermatologist, a dentist and a general practitioner evaluated the disease. Dermatological findings common to all seven patients were focal plantar keratoderma, folliculitis on the ventral side of midarms and/or occipital region, and keratosis pilaris on occipital region. Acanthosis nigricans was observed in five patients and hyperpigmentation of hands and few nails in one patient (Supplementary Fig. 1). Two patients had additional features: patient I-501 had hyperpigmentation of hands and nails of thumb and hallux, and angular cheilitis, and patient II-406 had warts on dorsal side of right 5th finger. Brachydactyly of 5th finger was present bilaterally in six patients.
PC-like nails in patients 415, 505, 507, 406, and 411; focal plantar keratoderma in patients 415 and 411, 5th finger brachydactyly in patients 415, 507, and 411; decaying teeth, hypodontia and diastema in patients 505, 507, and 406; and rough and sparse hair and scalp hair loss in patients 505, 507, and 406.
Dental findings were mixed dentition and decaying teeth in all patients, and in some of them additionally missing anterior teeth, peg-shaped incisors, diastema and attrition in anterior teeth (Supplementary Table 1). All other 23 relatives examined were assessed as unaffected. Thirteen of those were tested for the detected variant, and 12 were found heterozygous whereas one did not carry the variant. A patient brother in each family had sparse eyebrows. Several of the unaffected relatives also had 5th finger brachydactyly.
Genetic findings
KRT17 c.281G>A (p.(Arg94His); NM_000422.3) was found as the only shared rare homozygous exonic variant in the exome files of a patient from each family (Supplementary Tables 2 and 3). Exome files of their unaffected relatives revealed heterozygosity (Supplementary Fig. 2). Sanger sequencing performed to validate the variant indicated heterozygosity in affected individuals and about a quarter dose in 12 others (Supplementary Fig. 3). A genome-wide search for sequence similarity to the PCR product revealed four regions (Supplementary Material).
To investigate the segregation of the haplotype harboring the causal variant, multipoint linkage analysis was performed for both families (Supplementary Fig. 4). A 5.2-Mb region was identified as the locus of the gene responsible for the disease, with a cumulative maximal multipoint LOD score of 4.44. Patients were homozygous for the same SNP genotype, and all others were heterozygous (Supplementary Fig. 5). We concluded that KRT17 c.281G>A at the locus underlies the disease.
Discussion
Patients had atypical PC2 with not much variability, characterized by an unusual combination of some PC features such as pachyonychia, focal plantar keratoderma, folliculitis, alopecia, sparse eyebrows, keratosis pilaris, and dental anomalies plus less common findings such acanthosis nigricans of neck, dry skin, palmoplantar hyperhidrosis, and recurrent blisters that heal spontaneously. The characteristics in common with the known PC conditions include pachyonychia, sparse eyebrows and sparse scalp hair. On the other hand, the condition is distinct from the known PCs considering that some PC-associated hyperkeratosis types are not found such as palmoplantar hyperkeratosis, and follicular keratosis of knees and elbows. Also, epidermoid cysts, pain in soles, leukokeratosis, coarse voice, or history of natal teeth seen in some PCs are not present.
We assessed that this first recessive PC reported to date is a new PC due to the involvement of all ectodermal derivatives plus an unusual combination of shared and variable features. We found the same known KRT17 missense variant c.281G>A in homozygous state in the families. Sharing of another very rare variant and SNP genotypes in the identified gene region indicated identity by descent. The causative variant is not listed in databases for population variation but has been reported in heterozygous state first in a family afflicted with severe steatocystoma multiplex, nails either absent or thickened, no history of natal teeth, and variable mild epidermolytic palmoplantar keratoderma [3], and later in a sporadic PC2 case with palmoplantar keratoderma and pilosebaceous cysts [4]. In the families we present, among the examined patient relatives (12 heterozygotes and four parents not tested but considered as obligate carriers) only I-416 and II-407 who was not tested have sparse eyebrows as a mild characteristics of PC. PC features are not rare in the general population; for example, sister I-502 who does not carry the variant has dry skin.
Alopecia seen in all of our patients is not a feature of PC but has been reported in two unrelated cases with severe PC due to “homozygous dominant” missense KRT17 variants which were already known to cause PC in heterozygous state [9]. First proband had c.275 A > G (p.(Asn92Ser)), the most common KRT17 variant in PC, and his four heterozygous relatives were mildly affected. Second proband was a 32-year-old male with c.280 C > T (p.(Arg94Cys)), reported in several cases of dominant PC. Father reportedly had steatocysts. Those variants are assessed to have homozygous dominant effect whereas the variant we present herein is homozygous recessive. Keratin 17 null mice exhibited age (6 months after birth) and strain-dependent alopecia, with some strains not exhibiting the phenotype at all [15]. “The examination failed to reveal obvious anomalies at a gross level in the nail”, and no dental evaluation was mentioned.
As an initial step, we performed exome sequencing to assess whether the disease gene was new. After detecting a homozygous KRT17 variant, we attempted Sanger sequencing to validate the variant, mislead by the UCSC in-silico PCR tool which indicated that the designed primers targeted a unique genomic region. Obtaining ‘heterozygous’ results which were not compatible with the exome data, we performed SNP genotyping and linkage analysis. We did not opt to perform nested PCR instead which is easier and costs less, as we wanted also to investigate whether the haplotype harboring the causal variant indeed segregated recessively with full penetrance and was the same in both families, and whether there was yet another recessive locus that could possibly harbor a genetic modifier. We identified a single homozygous locus shared by affected individuals only and detected a known variant reported in heterozygous state in a PC2 case and a steatocystoma multiplex family, and hence it is difficult to speculate why the variant has recessive effect in the families we present and the majority (11 of 12) of heterozygotes do not have any signs of the disease. On the other hand, it is not very rare that a known gene linked to a dominant disease can also be linked to a recessive disease with a different clinical phenotype, as discussed in supplementary materials.
The presented families are the first to carry a KRT17 variant which is pathogenic in homozygous state only. The variant in homozygous state manifests with a condition different from the two known KRT17-related dominant diseases, widening the KRT17-related phenotype in addition to the inheritance pattern.
There are many points that call for caution in genetic testing for PCs, as techniques are not always easy to apply and interpretation of the results not straightforward. Sanger sequencing should be carefully designed due to the presence of pseudogenes, and sequencing of directly amplified exons should not be preferred (Supplementary material) [14]. Some older sequencing results obtained with Maxam-Gilbert method could be interpreted as homozygous rather than heterozygous (Fig 4a in ref. [2]). Without a control assay, it is not possible to be sure that the restriction enzyme digestion is complete and not partial (Fig. 4c in ref. [2]). Considering that an important portion of heterozygous KRT17 variants reported are not familial but “spontaneous”, used as opposite to inherited, and thus, to mean de novo48,12 and that parents generally are not tested for the variant found in their sib, it is tempting to speculate that some of the variants reported as spontaneous could in fact be homozygous and not de novo heterozygous. To date exome sequencing has not been reported in studies with most keratin genes including KRT17 and SNP genotyping reported in only a few. Our results raise caution for dominant inheritance in PCs, and we recommend that parents be tested for the variants detected in sibs in KRT17 and even in other KRT genes, as a recessive disease poses a lower risk for next generations and genetic counseling would be accordingly.
Web resources
The URLs for data presented herein are as follows:
Homozygosity Mapper: http://www.homozygositymapper.org/
Mutalyzer: https://mutalyzer.nl/
Mutation Taster 2: http://www.mutationtaster.org/
Online Mendelian Inheritance in Man (OMIM): https://www.omim.org/
Polymorphism Phenotyping v2 (PolyPhen-2): http://genetics.bwh.harvard.edu/pph2/
Sorting Intolerant From Tolerant (SIFT): http://sift.jcvi.org/
UCSC Genome Bioinformatics (for Human Genome Browser): http://genome.cse.ucsc.edu/
CADD—Combined Annotation Dependent Depletion (CADD): https://cadd.gs.washington.edu/
M-CAP—Mendelian Clinically Applicable Pathogenicity (M-CAP): http://bejerano.stanford.edu/mcap/
UniProt—The Universal Protein Resource (UniProt): https://www.uniprot.org/
The Human Protein Atlas: https://proteinatlas.org/
InterPro: https://www.ebi.ac.uk/interpro/
Pachyonychia Congenita Project: www.pachyonychia.org
Data availability
Data will be made available upon request.
References
McLean WH, Moore CB. Keratin disorders: from gene to therapy. Hum Mol Genet. 2011;20:R189–97. https://doi.org/10.1093/hmg/ddr379.
McLean WH, Rugg EL, Lunny DP, Morley SM, Lane EB, Swensson O, et al. Keratin 16 and keratin 17 mutations cause pachyonychia congenita. Nat Genet. 1995;9:273–8. https://doi.org/10.1038/ng0395-273.
Smith FJ, Corden LD, Rugg EL, Ratnavel R, Leigh IM, Moss C, et al. Missense mutations in keratin 17 cause either pachyonychia congenita type 2 or a phenotype resembling steatocystoma multiplex. J Invest Dermatol. 1997;108:220–3. https://doi.org/10.1111/1523-1747.ep12335315.
Terrinoni A, Smith FJ, Didona B, Canzona F, Paradisi M, Huber M. et al. Novel and recurrent mutations in the genes encoding keratins K6a, K16 and K17 in 13 cases of pachyonychia congenita. J Invest Dermatol. 2001;117:1391–6. https://doi.org/10.1046/j.0022-202x.2001.01565.x.
Smith FJ, Coleman CM, Bayoumy NM, Tenconi R, Nelson J, David A, et al. Novel keratin 17 mutations in pachyonychia congenita type 2. J Invest Dermatol. 2001;116:806–8. https://doi.org/10.1046/j.1523-1747.2001.01335.x.
Hashiguchi T, Yotsumoto S, Shimada H, Terasaki K, Setoyama M, Kobayashi K, et al. A novel point mutation in the keratin 17 gene in a Japanese case of pachyonychia congenita type 2. J Invest Dermatol. 2002;118:545–7. https://doi.org/10.1046/j.0022-202x.2001.01701.x.
Xiao SX, Feng YG, Ren XR, Tan SS, Li L, Wang JM, et al. A novel mutation in the second half of the keratin 17 1A domain in a large pedigree with delayed-onset pachyonychia congenita type 2. J Invest Dermatol. 2004;122:892–5. https://doi.org/10.1111/j.0022-202X.2004.22408.x.
Wilson NJ, Leachman SA, Hansen CD, McMullan AC, Milstone LM, Schwartz ME, et al. A large mutational study in pachyonychia congenita. J Invest Dermatol. 2011;131:1018–24. https://doi.org/10.1038/jid.2011.20.
Wilson NJ, Pérez ML, Vahlquist A, Schwartz ME, Hansen CD, Irwin McLean WH, et al. Homozygous dominant missense mutation in keratin 17 leads to alopecia in addition to severe pachyonychia congenita. J Invest Dermatol. 2012;132:1921–4. https://doi.org/10.1038/jid.2011.484.
Ghazawi FM, Hassani-Ardakani K, Henriques L, Jafarian F. Identification of a novel substitution mutation (R103C) in the rod domain of the keratin 17 gene associated with pachyonychia congenita type 2. Int J Dermatol. 2019;58:233–6. https://doi.org/10.1111/ijd.14082.
Dabbagh B, Cukier O, Yeganeh M, Halal F, Dos Santos BF. Pachyonychia congenita associated with a novel variant of KRT17 presenting unusual oral manifestations. J Dent Child (Chic). 2019;86:61–63.
Samuelov L, Smith FJD, Hansen CD, Sprecher E. Revisiting pachyonychia congenita: a case-cohort study of 815 patients. Br J Dermatol. 2020;182:738–46. https://doi.org/10.1111/bjd.18794. 03.
Pavlovsky M, Peled A, Samuelov L, Malki L, Malovitski K, Assaf S, et al. Molecular epidemiology of pachyonychia congenita in the Israeli population. Clin Exp Dermatol. Nov 2020;https://doi.org/10.1111/ced.14509
Zhang B, Sun L, Fu X, Yu G, Liu H, Zhang F. Mutation analysis of the KRT17 gene in steatocystoma multiplex and a brief literature review. Clin Exp Dermatol. 2020;45:132–4. https://doi.org/10.1111/ced.14030.
McGowan KM, Tong X, Colucci-Guyon E, Langa F, Babinet C, Coulombe PA. Keratin 17 null mice exhibit age and strain-dependent alopecia. Genes Dev. 2002;16:1412–22. https://doi.org/10.1101/gad.979502.
Acknowledgements
We are grateful to families for their collaboration. We acknowledge the contribution of Dr. Muhammad Jamal Ullah to clinical evaluation of participants. Adult patients and guardians of minor patients gave written informed consent to publication of the relevant case details.
Funding
Supported by the Istanbul Technical University Research Fund (TGA-2021-42537) and URF-QAU, Pakistan (DAS-1071).
Author information
Authors and Affiliations
Contributions
Conceptualization: AT and SM; Formal analysis: MK and GN; Funding acquisition: AT and SM; Investigation: MK, MN, GN, RMKS, TM, ZH, SM, and AT; Project administration: AT and SM; Supervision: AT and SM; Validation: MK and GN; Visualization: MK, MN, GN, and RMKS; Writing - Original Draft Preparation: AT, SM, MK, and GN; Writing - Review and Editing: AT and SM.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethical approval
The procedures followed were in accordance with the ethical standards of the Ethical Review Committee of Quaid-i-Azam University (review no DAS-1070) and the Istanbul Technical University Human Research Ethical Review Board (MB62/2019).
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Koprulu, M., Naeem, M., Nalbant, G. et al. KERATIN 17-related recessive atypical pachyonychia congenita with variable hair and tooth anomalies. Eur J Hum Genet 30, 1292–1296 (2022). https://doi.org/10.1038/s41431-022-01128-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41431-022-01128-4
This article is cited by
-
Genome sequencing—do you know what you are getting into?
European Journal of Human Genetics (2022)