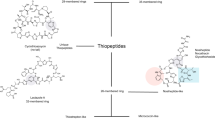
Abstract
Nucleoside antibiotics possess various biological activities such as antibacterial, antifungal, anticancer, and herbicidal activities. RIKEN scientists contributed to this area of research with two representative antifungal nucleoside antibiotics, blasticidin S and polyoxin. Blasticidin S was the first antibiotic exploited in agriculture worldwide. Meanwhile, the polyoxins discovered by Isono and Suzuki are still used globally as an agricultural antibiotic. In this review article, the research on nucleoside antibiotics mainly done by Isono and his collaborators is summarized from the discovery of polyoxin to subsequent investigations.
Similar content being viewed by others
Introduction
Pesticides containing organic mercury were widely used to suppress plant pathogens until the 1950s especially against the rice blast disease. But the use of mercury-containing agrochemicals was banned around 1960 when it became clear that the cause of the Minamata disease was the cumulative toxicity of organic mercury. Thus, the development of a safe pesticide to replace mercury was sought, and antibiotics such as blasticidin S [1, 2] and kasugamycin [3, 4] were discovered (Fig. 1).
Polyoxin
Researchers of RIKEN’s Antibiotics Laboratory led by Yusuke Sumiki and subsequently by Saburo Suzuki started the screening for antifungal antibiotics for the prevention of rice plant pathogens. A strong antifungal activity was detected in the culture broth of an actinomycete named Streptomyces cacaoi var. asoensis isolated from a soil sample collected at Mt. Aso, Kumamoto Prefecture [5]. The active substance was water-soluble and multicomponent. Although the isolation and purification were very difficult, they succeeded in obtaining the main component as a colorless crystal. With a structural feature containing many oxygen atoms, it was named polyoxin A (Fig. 2). Moreover, this compound appeared to have a nucleoside skeleton containing a hydroxymethyluracil moiety. At that time, there was no high-performance liquid chromatography (HPLC) so the numerous components were separated by an open cellulose column. By 1968, they had isolated and identified the chemical structures of all the components, polyoxins A–L (Fig. 2a) [6]. Isono pointed out at an international conference that the basic structure of polyoxin was similar to uridine diphosphate-N-acetylglucosamine which is a substrate of chitin synthetase. In 1970, Endo and Misato clearly demonstrated that polyoxin inhibited the enzymatic activity of chitin synthetase [7]. The RIKEN researchers called the polyoxins “fungus penicillin” because they inhibit the cell wall synthesis of fungi. They further isolated new derivatives, polyoxins N and O (Fig. 2b). They studied the structure–activity relationship and made derivatives by interconversion between components via sulfite decarboxylation. It was also found that the producer strain, S. cacaoi, can synthesize unnatural polyoxins by the incorporation of 5- fluoro-, 5-bromo-, and 6-azauracil-uracil into the polyoxins.
The polyoxins are effective not only for rice fungal diseases (Pyricularia oryzae, Cochliobolus miyabeanus) but also for various diseases caused by phytopathogenic filamentous fungi such as the gray mold disease of fruits (Botrytis cinerea) and the black spot disease of Japanese pear (Alternaria kikuchiana). Being a specific inhibitor of cell wall chitin, the polyoxins have no toxicity to animals and plants. In addition, they are easily degraded in the environment making them ideal green pesticides. To date, the polyoxins are still widely used as an agricultural antibiotic.
Cell wall synthesis inhibitors
The excellent selective toxicity of the polyoxins was attributed to the inhibition of cell wall synthesis; therefore, Nishii and Isono designed several new screening systems targeting enzymes for cell wall synthesis in collaboration with Kazuo Izaki of Tohoku University. They established in vitro assay systems targeting bacterial peptidoglycan synthesis, yeast chitin synthesis, and fungal beta-glucan/mannan synthesis. In these screenings, lipopeptin [8], neopeptin [9], neopolyoxin [10, 11], and phosphazomycin [12] were discovered. In 1979, mucopeptin was isolated as an inhibitor of bacterial cell wall synthesis by Kusano et al. (reported only in Japanese, Japanese Patent Application 56-139499, 1981), however, the chemical structure could not be determined due to its complexity. In 1987, Lederle Laboratories isolated BO2964 and its analogs (US Patent 4,677,071, 1987), which might be similar compounds to mucopeptin. Moreover, chemically and biologically similar compounds, pacidamycins and mureidomycins (Fig. 3), were isolated by Abbott Laboratories [13,14,15] and Sankyo Co. Ltd. [16, 17], respectively, and the structures of these nucleoside peptides were revealed. Later, the stereochemistry of these compounds and their derivatives was elucidated by chemical synthesis [18].
Nikkomycin and neopolyoxin
Zahner et al. at the University of Tubingen reported the isolation and structure determination of the nikkomycin group as a chitin synthesis inhibitor in 1976 [19]. Subsequently, the Isono group isolated neopolyoxins A–C from the polyoxin-producing strain [10, 11]. Coincidentally, neopolyoxins A and C were structurally identical to nikkomycin X and Z, respectively (Fig. 4) [10]. Isono et al. wondered how these derivatives, which differed only at the nucleobase moiety, were biosynthesized. Their effort revealed that pyrimidine undergoes a ring contraction reaction to form oxo-imidazoline [11]. On the other hand, the Zahner group continued to isolate many derivatives of nikkomycin [20, 21] and tried to develop these compounds as therapeutic agents but to no avail.
Liposidomycin and caprazamycin
The liposidomycin complex was isolated from Streptomyces griseosporeus as an inhibitor of bacterial peptidoglycan synthesis (Fig. 5). Despite the difficulty of isolating the more than 12 components of the mixture, Kimura et al. successfully accomplished the task [22, 23]. They were able to elucidate the complex nucleoside structure by collaborating with McCloskey who was in charge of the structure elucidation [24]. Both polyoxin and liposidomycin contain a uracil moiety and inhibit the fungal and bacterial cell wall synthesis, respectively, however, their mode of action is different from each other. The polyoxins inhibit fungal chitin synthesis unlike liposidomycin which inhibits bacterial peptidoglycan synthesis. Isono dubbed liposidomycin as a “bacterial polyoxin” because of their structural and target similarities.
Furthermore, the precise action mechanism of liposidomycin was shown to be as an inhibitor of UDP-N-acetylmuramoyl-pentapeptide: undecaprenyl-phosphate phospho-N-acetylmuramoyl-pentapeptide transferase (MraY), an essential membrane enzyme for bacterial cell wall biosynthesis (Fig. 6) [25]. Despite the remarkable activity of liposidomycin in the cell-free peptidoglycan biosynthesis, antibacterial activity was not strong and observed only in limited bacteria. Therefore, its development as a therapeutic agent was suspended.
Key enzymes, MraY, and MurG, for peptidoglycan biosynthesis in bacterial cell wall. MraY (PDB code: 4J72) is phospho-N-acetylmuramoyl-pentapeptide transferase and catalyzes the first step of bacterial peptidoglycan synthesis to form lipid I. MurG (PDB code: 1NLM) is UDP-N-acetylglucosamine-N-acetylmuramyl-(pentapeptide) pyrophosphoryl-undecaprenol N-acetylglucosamine transferase, and converts lipid I to lipid II
In 2003, the Bikaken group isolated caprazamycin (Fig. 7) from the culture broth of Streptomyces sp. MK730-62F2, and found that it was active against mycobacteria [26]. The caprazamycins consisted of several members, and the structures were identified by extensive NMR analyses [27]. To examine the fragments of the structures, they subjected the caprazamycins to acid treatment. Subsequently, the core moiety of caprazamycin, caprazen (CPZEN) (Fig. 8), was obtained in high yield. Semisynthetic derivatives containing the CPZEN moiety with different side chains, were then obtained [28]. One of the derivatives, CPZEN-45, showed remarkable activity against Mycobacterium tuberculosis [29]. Moreover, CPZEN-45 showed antibacterial activity against caprazamycin-resistant strains, including a strain overexpressing MraY. Finally, it was revealed that the N-acetylglucosamine-1-phosphate transferase, WecA, is the target of the caprazamycins [30].
Liposidomycin and caprazamycin have more complex chemical structures compared with CPZEN-45. Liposidomycin and caprazamycin inhibited MraY activity which recognize UDP-MurNAc-pentapeptide as a substrate. CPZEN-45 has a simpler structure and inhibited WecA activity which uses UDP-GlcNAc as a substrate (Fig. 9).
Guanine 7-N-oxide
As described above, the Isono group were interested in isolating the inhibitors of microbial cell wall synthesis. Nishii came to RIKEN from a company and established the screening system of the inhibitors of cell wall synthesis. Although he went back to the company, he kept the collaboration with Isono and they discovered guanine 7-N-oxide (Fig. 10) [31]. It was an unusual nucleobase and exhibited remarkable anticancer and antiviral effects [32]. This simple compound had never been synthesized, but the microbes had taught us its existence. Later, a collaboration of RIKEN and Kanazawa University succeeded in its chemical synthesis [33]. It was expected to be developed as an antiviral agent but the development was unsuccessful. The Isono group expanded the screening target to antitumor compounds from the mid-1980s along with antimicrobial agents.
Ascamycin
The nucleoside antibiotic, ascamycin [34], was isolated from a Streptomyces sp. together with dealanylascamycin which was identical to AT-265 (Fig. 11) [35]. AT-265 (dealanylascamycin) showed a broad range of antibacterial activity against various Gram-negative and Gram-positive bacteria. On the contrary, ascamycin showed selective toxicity against plant pathogenic Gram-negative bacteria, Xanthomonas campestris pv. citri (X. citri) and X. campestris pv. oryzae (X. oryzae). When the inhibitory activity of ascamycin and dealanylascamycin was tested using a cell-free protein synthesis system, both compounds inhibited the protein synthesis of bacteria not only in X. citri but also in Escherichia coli [36]. These data suggested that the differential antibacterial activity of ascamycin and dealanylascamycin was dependent on the membrane permeability. Based on this hypothesis, a hydrolyzing enzyme of ascamycin was purified from the membrane fraction of X. citri. Because it was a new enzyme belonging to the proline iminopeptidase family, it was named Xc-aminopeptidase (Fig. 12) [37]. Later, the gene encoding Xc-aminopeptidase was cloned from X. citri by our group [38].
When ascamycin was isolated, it was considered as a lead compound against plant pathogenic bacteria. Moreover, the aminopeptidase gene was cloned from X. citri, and a homologous gene was found in Neisseria gonorrhoeae, a Gram-negative bacterium which causes the sexually transmitted genitourinary infection, gonorrhea. Then, the inhibitory activity of ascamycin against N. gonorrhoeae was examined, but the potency was very low. It was speculated that the localization of the aminopeptidase in N. gonorrhoeae might be different from that in X. campestris. At the same time, it was reported that the aminopeptidase activity in tumor cell lines is higher than that in normal cells. This observation suggested that ascamycin might act as a prodrug of an antitumor compound, and the structure-activity relationship of ascamycin derivatives against tumor cells was examined. Many ascamycin derivatives were synthesized by substituting the amino acid moiety [39, 40] in which the prolyl derivative of ascamycin was the most potent among the aminoacyl derivatives. This result was consistent with the observation that tumor cells expressed high aminopeptidase activity, especially proline iminopeptidase activity [41].
Protein kinase inhibitors
At the beginning of the 1980s, tumor biology research was rapidly advanced and protein kinases were targeted by the screeners as antitumor agents [42,43,44]. As a representative discovery in the field of the tumor biology, Nishizuka et al. revealed that protein kinase C (PKC) is a receptor protein of the tumor promoting compounds, phorbol esters [45, 46].
Responding to those backgrounds, the screening target of the Isono group was gradually shifted from agrochemical compounds to antitumor compounds [47, 48]. During the screening for PKC inhibitors using the bleb forming assay, a deazaadenosine-type nucleoside compound, sangivamycin (Fig. 13) was found and it showed strong inhibitory activity against PKC [49]. Although sangivamycin was a known compound, it was the first report regarding the PKC inhibition. Tubercidin, a deazaadenosine nucleoside antibiotic [50], was previously isolated by RIKEN Antibiotics Laboratory, and the mode of action was revealed to be via incorporation into DNA and RNA, and inhibition of their synthesis [51]. In those days, the mode of action of the deazaadenosine antibiotics was thought to be same. In other words, they were mimicking adenosine, but recently, new biological activities of the deazaadenosine antibiotics have been reported [52,53,54].
As the bleb-forming assay was very rapid and sensitive to detect PKC inhibitors [48], indolocarbazole-type PKC inhibitors besides sangivamycin were effectively identified. Indolocarbazole-type compounds, staurosporine and K-252a, were reported as PKC inhibitors by the Kyowa Hakko group. The RIKEN group found new derivatives of the indolocarbazoles such as RK-286C, RK-1409A and B. (Fig. 14) [55,56,57,58,59,60,61]. As the chemical structure of the indolocarbazole is partially similar to adenosine, many kinase inhibitors have been developed based on the structure of indolocarbazole [62].
Phosmidosine
Phosmidosine (Fig. 15) was isolated by Uramoto et al. as a weak antifungal antibiotic from the culture of Streptomyces sp. RK-16 [63]. Later, Matsuura et al. isolated phosmidosine and its derivative, phosmidosine B, as detransforming compounds showing tumor morphology reversion activity in src-oncogene-transformed NRK (tsNRK) cells (Fig. 16) [64]. As these tsNRK cells contain the temperature sensitive v-src gene, the cell morphology was transformed at 32 °C and reverted to the normal morphology at 39 °C. As shown in Fig. 16, phosmidosine reverted the transformed morphology to the normal morphology at 32 °C. Moreover, phosmidosine arrested the cell-cycle progression at the G1 phase through the inhibition of cyclin D1 expression in cancer cells [65].
Conclusion and future direction
Many important nucleoside/nucleotide compounds such as zidovudine (AZT) [66] and lamivudine (3TC) [67] have been synthesized and developed as antiviral agents, particularly as anti-HIV agents [67, 68]. However, in this review, the antibiotics are mainly focused on those isolated as microbial metabolites.
During the golden age of antibiotic discovery, there were a lot of chances to discover novel antibiotics. Moreover, the newly discovered antibiotics were launched to the market within a few years. Nowadays, it has become too difficult to explore novel antibiotics which could be exploited as therapeutic agents. Therefore, a new approach is needed to overcome these difficulties to find a new compound from microbial fermentation broths. One strategy is the construction of a broth library and further, a semipurified (fraction) library with HPLC/MS data [69]. As high-throughput screening (HTS) requires a large number of the screening samples, most pharmaceutical companies gave up on natural products and shifted to use the chemical library prepared by combinatorial synthesis. However, the semipurified (fraction) library is applicable to HTS. In addition, HPLC/MS data of the fraction library enables a certain degree of dereplication of known compounds from the microbial fermentation broths [70, 71]. Moreover, recent advances in biosynthesis enable production of a large amount of the targeted antibiotics [72,73,74].
Microbial metabolites are still thought to be an important source for drug screening because it offers a wider chemical space (diversity) and stronger observed biological activity compared with the compounds synthesized by combinatorial chemistry. Among the antibiotics, the nucleoside antibiotics are fascinating because of their broad biological activities [75,76,77].
References
Huang KT, Misato T, Asuyama H. Selective toxicity of blasticidin S to Piricularia oryzae And Pellicularia sasakii. J Antibiot. 1964;17:71–74.
Takeuchi S, Hirayama K, Ueda K, Sakai H, Yonehara H. Blasticidin S, a new antibiotic. J Antibiot. 1958;11:1–5.
Umezawa H, Hamada M, Suhara Y, Hashimoto T, Ikekawa T. Kasugamycin, a new antibiotic. Antimicrob Agents Chemother. 1965;5:753–7.
Urabe H, Nakama T. Antifungal activity of Kasugamycin. J Antibiot B. 1967;20:424–6.
Suzuki S, et al. A new antibiotic, polyoxin A. J Antibiot. 1965;18:131.
Isono K, Asahi K, Suzuki S. Studies on polyoxins, antifungal antibiotics. 13. The structure of polyoxins. J Am Chem Soc. 1969;91:7490–505.
Endo A, Kakiki K, Misato T. Mechanism of action of the antifugal agent polyoxin D. J Bacteriol. 1970;104:189–96.
Tsuda K, et al. A new antibiotic, lipopeptin A. J Antibiot. 1980;33:247–8.
Ubukata M, Uramoto M, Uzawa J, Isono K. Structure and biological activity of neopeptins A, B and C, inhibitors of fungal cell wall glycan synthesis. Agric Biol Chem. 1986;50:357–65.
Kobinata K, et al. Neopolyoxins A, B, and C, new chitin synthetase inhibitors. Agric Biol Chem. 1980;44:1709–11.
Uramoto M, Kobinata K, Isono K. Chemistry of the neopolyoxins, pyrimidine and imidazoline nucleoside peptide antibiotics. Tetrahedron. 1980;38:1599–608.
Uramoto M, Shen Y-C, Takizawa N, Kusakabe H, Isono K. A new antifungal antibiotic phosphazomycin A. J Antibiot. 1985;5:665–8.
Karwowski JP, et al. Pacidamycins, a novel series of antibiotics with anti-Pseudomonas aeruginosa activity. I. Taxonomy of the producing organism and fermentation. J Antibiot. 1989;42:506–11.
Fernandes PB, et al. Pacidamycins, a novel series of antibiotics with anti-pseudomonas aeruginosa activity. III. Microbiologic profile. J Antibiot. 1989;42:521–6.
Chen RH, Buko AM, Whittern DN, McAlpine JB. Pacidamycins, a novel series of antibiotics with anti-Pseudomonas aeruginosa activity. II. Isolation Struct elucidation. J Antibiot. 1989;42:512–20.
Inukai M, et al. Mureidomycins A-D, novel peptidylnucleoside antibiotics with spheroplast forming activity. I. Taxonomy, fermentation, isolation and physico-chemical properties. J Antibiot. 1989;42:662–6.
Isono F, et al. Mureidomycins A-D, novel peptidylnucleoside antibiotics with spheroplast forming activity. II. Structural elucidation. J Antibiot. 1989;42:667–73.
Boojamra CG, et al. Stereochemical elucidation and total synthesis of dihydropacidamycin D, a semisynthetic pacidamycin. J Am Chem Soc. 2001;123:870–4.
Dahn U, et al. Stoffwechselprodukte von mikroorganismen. 154. Mitteilung. Nikkomycin, ein neuer hemmstoff der chitinsynthese bei pilzen. Arch Microbiol. 1976;107:143–60.
Delzer J, et al. New nikkomycins by mutasynthesis and directed fermentation. J Antibiot. 1984;37:80–82.
Decker H, Zahner H, Heitsch H, Konig WA, Fiedler HP. Structure-activity relationships of the nikkomycins. J Gen Microbiol. 1991;137:1805–13.
Isono K, et al. Liposidomycins: novel nucleoside antibiotics which inhibit bacterial peptidoglycan synthesis. J Antibiot. 1985;38:1617–21.
Kimura K, et al. Selective inhibition of the bacterial peptidoglycan biosynthesis by the new types of liposidomycins. J Antibiot. 1998;51:1099–104.
Ubukata M, Isono K, Kimura K, Nelson C, McCloskey J. The structure of liposidomycin B, an inhibitor of bacterial peptidoglycan synthesis. J Am Chem Soc. 1988;110:4416–7.
Muroi M, Kimura K, Osada H, Inukai M, Takatsuki A. Liposidomycin B inhibits in vitro formation of polyprenyl (pyro)phosphate N-acetylglucosamine, an intermediate in glycoconjugate biosynthesis. J Antibiot. 1997;50:103–4.
Igarashi M, et al. Caprazamycin B, a novel anti-tuberculosis antibiotic, from Streptomyces sp. J Antibiot. 2003;56:580–3.
Igarashi M, et al. Caprazamycins, novel lipo-nucleoside antibiotics, from Streptomyces sp. II. Structure elucidation of caprazamycins. J Antibiot. 2005;58:327–37.
Takahashi Y, et al. Novel semisynthetic antibiotics from caprazamycins A-G: caprazene derivatives and their antibacterial activity. J Antibiot. 2013;66:171–8.
Salomon JJ, et al. Biopharmaceutical in vitro characterization of CPZEN-45, a drug candidate for inhalation therapy of tuberculosis. Ther Deliv. 2013;4:915–23.
Ishizaki Y, et al. Inhibition of the first step in synthesis of the mycobacterial cell wall core, catalyzed by the GlcNAc-1-phosphate transferase WecA, by the novel caprazamycin derivative CPZEN-45. J Biol Chem. 2013;288:30309–19.
Nishii M, et al. A new antitumor antibiotic, guanine 7-N-oxide produced by Streptomyces sp. J Antibiot. 1985;38:1440–3.
Hasobe M, Saneyoshi M, Isono K. Antiviral activity and its mechanism of guanine 7-N-oxide on DNA and RNA viruses derived from salmonid. J Antibiot. 1985;38:1581–7.
Nohara F, et al. Synthesis of guanine 7-oxide, an antitumor antibiotic from species. Tetrahedron Lett. 1987;28:1287–90.
Isono K, et al. Ascamycin and dealanylascamycin, nucleoside antibiotics from Streptomyces sp. J Antibiot. 1984;37:670–2.
Takahashi E, Beppu T. A new nucleoside antibiotic AT-265. J Antibiot. 1982;35:939–47.
Osada H, Isono K. Mechanism of action and selective toxicity of ascamycin, a nucleoside antibiotic. Antimicrob Agents Chemother. 1985;27:230–3.
Osada H, Isono K. Purification and characterization of ascamycin hydrolyzing aminopeptidase from Xanthomonas citri. Biochem J. 1986;233:459–63.
Sudo T, et al. Isolation and characterization of the gene encoding an aminopeptidase from Xanthomonas campestris pv. citri. Biochem J. 1996;319:99–102.
Ubukata M, Osada H, Isono K. Synthesis and biological activity of nucleoside antibiotics, ascamycin and amino acid analogs. Nucleic Acids Res Symp Ser. 1985;16:81–83.
Ubukata M, Osada H, Magae J, Isono K. Synthesis and biological activity of aminoacyl analogs of ascamycin. Agric Biol Chem. 1988;52:1117–22.
Osada H, Isono K. Occurrence of an ascamycin dealanylating enzyme, Xc-aminopeptidase in mammalian cell membranes and susceptibility to ascamycin. J Antibiot. 1986;39:286–93.
Tamaoki T, et al. Staurosporine, a potent inhibitor of phospholipid/Ca++ dependent protein kinase. Biochem Biophys Res Commun. 1986;135:397–402.
Uehara Y, Hori M, Takeuchi T, Umezawa H. Phenotypic change from transformed to normal induced by benzoquinonoid ansamycins accompanies inactivation of p60src in rat kidney cells infected with Rous sarcoma virus. Mol Cell Biol. 1986;6:2198–206.
Umezawa H, et al. Studies on a new epidermal growth factor-receptor kinase inhibitor, erbstatin, produced by MH435-hF3. J Antibiot. 1986;39:170–3.
Castagna M, et al. Direct activation of calcium-activated, phospholipid-dependent protein kinase by tumor-promoting phorbol esters. J Biol Chem. 1982;257:7847–51.
Kikkawa U, Takai Y, Tanaka Y, Miyake R, Nishizuka Y. Protein kinase C as a possible receptor protein of tumor-promoting phorbol esters. J Biol Chem. 1983;258:11442–5.
Magae J, Watanabe C, Osada H, Cheng X-C, Isono K. Induction of morphological change of human myeloid leukemia and activation of protein kinase C by a novel antibiotic, tautomycin. J Antibiot. 1988;41:932–7.
Osada H, Magae J, Watanabe C, Isono K. Rapid screening method for inhibitors of protein kinase C. J Antibiot. 1988;41:925–31.
Osada H, Sonoda T, Tsunoda K, Isono K. A new biological role of sangivamycin; Inhibition of protein kinases. J Antibiot. 1989;42:102–6.
Anzai K, Nakamura G, Suzuki S. A new antibiotic, tubercidin. J Antibiot. 1957;10:201–4.
Acs G, Reich E, Mori M. Biological and biochemical properties of the analogue antibiotic tubercidin. Proc Natl Acad Sci USA. 1964;52:493–501.
Perlikova P, et al. 7-(2-Thienyl)-7-Deazaadenosine (AB61), a new potent nucleoside cytostatic with a complex mode of action. Mol Cancer Ther. 2016;15:922–37.
Hulpia F, et al. Revisiting tubercidin against kinetoplastid parasites: aromatic substitutions at position 7 improve activity and reduce toxicity. Eur J Med Chem. 2019;164:689–705.
Bachmaier S, et al. Nucleoside analogue activators of cyclic AMP-independent protein kinase A of Trypanosoma. Nat Commun. 2019;10:1421.
Osada H, Takahashi H, Tsunoda K, Kusakabe H, Isono K. A new inhibitor of protein kinase C, RK-286C (4’-demethylamino-4’-hydroxystaurosporine). I. Screening, taxonomy, fermentation and biological activity. J Antibiot. 1990;43:163–7.
Takahashi H, Osada H, Uramoto M, Isono K. A new inhibitor of protein kinase C, RK-286C (4’-demethylamino-4’-hydroxystaurosporine). II. Isolation, physico-chemical properties and structure. J Antibiot. 1990;43:168–73.
Usui T, et al. Uncoupled cell cycle without mitosis induced by a protein kinase inhibitor, K-252a. J Cell Biol. 1991;115:1275–82.
Koshino H, Osada H, Amano S, Onose R, Isono K. A new inhibitor of protein kinase C, RK-1409B (4’-demethylamino-4’-hydroxy-3’-epistaurosporine). J Antibiot. 1992;45:1428–32.
Koshino H, Osada H, Isono K. A new inhibitor of protein kinase C, RK-1409 (7-oxostaurosporine). II. Fermentation, isolation, physico-chemical properties, and structure. J Antibiot. 1992;45:195–8.
Osada H, Hanaoka F, Isono K. Effects of microbial inhibitors of protein kinases and phosphatases on the cell cycle progression of mouse tsFT210 cells. Second joint meeting of the AACR/JCA (Molecular oncology as a basis for new strategies in cancer therapy), Honolulu;1992.
Osada H, Koshino H, Kudo T, Onose R, Isono K. A new inhibitor of protein kinase C, RK-1409 (7-oxostaurosporine). I. Taxonomy and biological activity. J Antibiot. 1992;45:189–94.
Nakano H, Omura S. Chemical biology of natural indolocarbazole products: 30 years since the discovery of staurosporine. J Antibiot. 2009;62:17–26.
Uramoto M, Kim CJ, Shinya K, Kusakabe H, Isono K. Isolation and characterization of phosmidosine a new antifungal nucleotide antibiotic. J Antibiot. 1991;44:375–81.
Matsuura M, Onose R, Osada H. Morphology reversion activity of phosmidosine and phosmidosine B, a newly isolated derivative, on src transformed NRK cells. J Antibiot. 1996;49:361–5.
Kakeya H, et al. Inhibition of cyclin D1 expression and phosphorylation of retinoblastoma protein by phosmidosine, a nucleotide antibiotic. Cancer Res. 1998;58:704–10.
Kolata G. FDA approves AZT. Science. 1987;235:1570.
van Leeuwen R, et al. The safety and pharmacokinetics of a reverse transcriptase inhibitor, 3TC, in patients with HIV infection: a phase I study. AIDS. 1992;6:1471–5.
Staszewski S. Zidovudine and lamivudine: results of phase III studies. J Acquir Immune Defic Syndr Hum Retrovirol. 1995;10(Suppl 1):S57.
Lim C, et al. Unantimycin A, a new neoantimycin analog isolated from a microbial metabolite fraction library. J Antibiot. 2016;69:456–8.
Kato N, Takahashi S, Nogawa T, Saito T, Osada H. Construction of a microbial natural product library for chemical biology studies. Curr Opin Chem Biol. 2012;16:101–8.
Osada H. Introduction of new tools for chemical biology research on microbial metabolites. Biosci Biotechnol Biochem. 2010;74:1135–40.
Biecker, AL, Liu, X, Thorson, JS, Yang, Z & Van Lanen, SG. Biosynthetic and synthetic strategies for assembling capuramycin-type antituberculosis antibiotics. Molecules. 2019;24:433.
Hong H, Samborskyy M, Zhou Y, Leadlay PF. C-nucleoside formation in the biosynthesis of the antifungal malayamycin A. Cell Chem Biol. 2019;26:493–501. e495
Shiraishi T, Nishiyama M, Kuzuyama T. Biosynthesis of the uridine-derived nucleoside antibiotic A-94964: identification and characterization of the biosynthetic gene cluster provide insight into the biosynthetic pathway. Org Biomol Chem. 2019;17:461–6.
Yssel AEJ, Vanderleyden J, Steenackers HP. Repurposing of nucleoside- and nucleobase-derivative drugs as antibiotics and biofilm inhibitors. J Antimicrob Chemother. 2017;72:2156–70.
Chang QY, et al. The IMPDH inhibitors, ribavirin and mycophenolic acid, inhibit peste des petits ruminants virus infection. Vet Res Commun. 2018;42:309–13.
Saha, A, Dutta, S & Nandi, N. Inhibition of seryl tRNA synthetase by seryl nucleoside moiety (SB-217452) of albomycin antibiotic. J Biomol Struct Dyn. 2019. https://doi.org/10.1080/07391102.2019.1635912
Acknowledgement
I thank M. Uramoto and J. Lopez for reading the manuscript and giving helpful comments. This author received JSPS KAKENHI Grants (JP17H06412, JP17K07784, JP18H03945), a Grant-in-Aid for the Project for Cancer Research and Therapeutic Evolution (P-CREATE) from AMED and the Project of the NARO Bio-oriented Technology Research Advancement Institution (Research program on development of innovative technology).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
This article is dedicated to Dr Kiyoshi Isono on the occasion of his 88th birthday with respect and admiration for his great achievements in nucleoside antibiotics research
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations
Rights and permissions
About this article
Cite this article
Osada, H. Discovery and applications of nucleoside antibiotics beyond polyoxin. J Antibiot 72, 855–864 (2019). https://doi.org/10.1038/s41429-019-0237-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41429-019-0237-1
This article is cited by
-
Structural basis for inhibition and regulation of a chitin synthase from Candida albicans
Nature Structural & Molecular Biology (2022)
-
Cryptic phosphorylation in nucleoside natural product biosynthesis
Nature Chemical Biology (2021)