Abstract
DNA-functionalized Au nanoparticles (AuNPs) have been intensively exploited as programmable atom equivalents (PAEs) for the self-assembly of molecule-like structures. However, it remains challenging to build hierarchical PAE assemblies via discrete DNA bonds at different levels. Here, we report a strategy to program DNA bond length and bond energy on PAEs using DNA encoders carrying consecutive adenines (polyA). On AuNPs, we built three types of DNA motifs with different topologic configurations, which can form bonds for PAE self-assembly. By small-angle X-ray scattering (SAXS) analysis, we found that the bond length and flexibility between the coupled PAEs can be tuned by programming the bond structure. We also found that these bonds show different bond energies and thus differ, depending on their topologic configuration, leading to different PAE assembly efficiencies. We demonstrated that the bonds at different levels can be arranged in different directions on one nanoparticle, leading to asymmetric PAEs that allow ionic strength-controlled hierarchical assembly of multiparticle structures. This programmable bonding system may provide a new route for building complex plasmonic superstructures.
Similar content being viewed by others
Introduction
In nature, atoms and molecules react with each other via interactions at different levels, which allow bottom-up self-assembly of complex hierarchical biological structures1,2,3,4,5,6,7. For example, the primary structure of DNA is assembled from nucleotides via phosphodiester bonds, and the secondary structure (double-stranded DNA, or dsDNA) is formed via hydrogen bonds; the secondary structure further binds histones to form nucleosomes via electrostatic and van der Waals interactions8,9,10. As a result, genomic DNA can reach meter-scale length while maintaining attometer-scale precision, allowing storage of genetic information with high capacity yet almost free of errors.
In the synthetic world, self-assembled hierarchical materials are also highly desired due to their unique features that arise from multiscale ordering11. Au nanoparticles (AuNPs) decorated with DNAs have been intensively explored as programmable atom equivalents (PAEs) for the self-assembly of crystal-like12,13,14,15,16,17,18,19,20,21,22,23 or molecule-like structures24,25,26,27,28,29,30,31,32,33,34. For example, in our previous study35,36,37, we developed a series of single-stranded DNA encoders to program discrete valences and bond angles of PAEs, which allowed self-assembly of colloidal molecules with defined particle numbers and spatial arrangements. However, these structures were mostly assembled via DNA bonds at the same level. The construction of bonds on an AuNP with different bond lengths and bond energies in different directions has not been well explored, which limits the potential application of these PAEs for hierarchical assembly.
In this work, we report a strategy to program DNA bond energy and bond length on PAEs using DNA encoders containing consecutive adenines (polyA). Due to the affinity between polyA–DNA and AuNPs36,37, the DNA encoders can form different configurations (of different sizes and base numbers) on the AuNP surface, which can serve as bonds at different levels for PAE self-assembly. These bonds showed different bond energies according to the thermophoresis of the PAE assemblies. We also verified different bond lengths between the PAE assemblies by using small-angle X-ray scattering (SAXS) analysis. Finally, we demonstrated that bonds at different levels can be arranged in different directions on one PAE, which allows ionic strength-controlled hierarchical assembly of particle structures.
Materials and methods
Reagents and materials
All oligonucleotides were obtained from Sangon Biotech Co., China. The sequences used in this study are listed in Supplementary Table 1. Psoralen (trioxsalen) and bis(p-sulfonatophenyl)phenylphosphine dihydrate dipotassium salt (BSPP) were obtained from Sigma-Aldrich (St Louis, MO, USA). AuNPs of 5 and 10 nm were obtained from Ted Pella Inc. (Redding, CA, USA). Agarose gel was obtained from Bio-Rad (Hercules, CA, USA).
Methods
Preparation of BSPP-protected AuNPs
Three milligrams of BSPP was added to 10 ml of AuNPs (10 nm), and the mixture was shaken overnight at room temperature. NaCl (solid) was added slowly to this mixture while stirring until the color changed from deep burgundy to light purple. The resulting mixture was centrifuged at 10,000 × g for 10 min, and the supernatant was removed. The treated AuNPs were then resuspended in 200 µl of BSPP solution (0.3 mg ml–1) and mixed with 200 µl of methanol. The mixture was again centrifuged, the supernatant was removed, and the AuNPs were resuspended in 100 µl of BSPP solution (0.3 mg ml–1). The concentration of AuNPs was estimated from the optical absorbance at 520 nm.
Preparation of DNA encoders
The DNA encoders for monovalent or multivalent type-III bonds (comprising 2–4 DNA strands) were synthesized by mixing the same molar amount of the corresponding oligonucleotide strands (sequences listed in Supplementary Table S1). After annealing of the samples (20 μM) in 0.5× PBS (50 mM Na+, 5 mM PB, pH = 7.4), cross-linking with psoralen was performed to increase the stability of the DNA encoders38,39. The samples were cross-linked by mixing with psoralen at a 1:1 molar ratio of psoralen and DNA base pairs. The mixture solution was irradiated with 365-nm UV light for 4 h at room temperature.
Preparation of PAEs with different DNA encoders
For assembly of PAEs, 100 nM BSPP–AuNPs (10 nm) were mixed with an equal molar amount of single-stranded DNA encoders (for bonds of types I and II) or preassembled multistranded encoders (for monovalent or multivalent type-III bonds). After 1 min, citrate–HCl buffer (500 mM, pH 3.1) was rapidly added to the DNA/AuNP mixture with a final citrate concentration of 10 mM. This mixture was incubated for 15 min at room temperature.
Assembly of PAE clusters
For assembly of dimer structures, monovalent PAEs with bonds I–III were incubated with complementary monovalent PAEs in 0.5× PBS with a molar ratio of 1:1. The mixture was then annealed by decreasing the temperature from 35 °C to 15 °C overnight with a thermoincubator. For multivalent assembly, 10-nm multivalent PAEs (valence=1–4) were incubated with complementary 5-nm monovalent PAEs in 0.5× PBS at a ratio of 1:2n (n, the valence of the former). The mixture was then annealed by decreasing the temperature from 35 °C to 15 °C overnight with a thermoincubator.
Ionic strength-controlled hierarchical assembly of particle structures
Bivalent PAEs (PAE-A) with bonds I and II were mixed with the complementary PAEs (PAE-B carrying bonds I and II) at a ratio of 1:1 in NaCl solution with different Na+ concentrations (5–100 mM). The mixture was incubated at room temperature for 24 h.
Agarose gel electrophoresis separation and recovery of AuNP assembly products
Four-percent agarose (dissolved in 0.5× TBE buffer) was used for gel separation. A 50-μl sample was mixed with 50% (w./v.) sucrose and loaded in the gel. The electrophoresis separation was performed with a Bio-Rad electrophoresis system at 100 V for 50 min. The gels were photographed with a digital camera. The desired bands were cut out from the gel and immersed in electroelution buffer (0.5× PBS). After electroelution, the products were recollected and concentrated by centrifugation (10,000×g, 30 min, CT15RE, Hitachi, Japan).
Transmission electron microscopy (TEM), dynamic light scattering (DLS), and atomic force microscopy (AFM) characterization
For TEM imaging, 10 µl of sample solution was deposited on a copper grid for 10 min, after which excess solution was removed using a piece of filter paper. TEM images were obtained with a Tecnai TF20 microscope operated at an acceleration voltage of 200 kV. For the DLS characterization, the apparent hydrodynamic size of dimers I~III was measured using a Zetasizer (Nano ZS90, Malvern Instruments, Worcestershire, UK). For AFM imaging, samples were prepared by deposition of 2 μl onto freshly cleaved mica and left for adsorption by the surface for 3 min. All AFM images were then taken in tapping mode on a Multimode Nanoscope VIII instrument (Bruker, USA).
Microscale thermophoresis (MST) measurements of dissociation constants (Kd)
The measurements were conducted on a Nano Temper Monolith NT115 instrument. Binding experiments were carried out in 0.5× PBS (50 mM NaCl, 5 mM PB, pH 7.4). For the experiment, Cy5-labeled cDNA-1 was used at a concentration of 2 nM. PAEs were titrated at a 1:1 dilution beginning at 100 nM. Monolith NT.115 hydrophilic capillaries were used in all experiments. The analysis was performed at 50% LED power and 20% MST power: “Fluo. Before”, 5 s; “MST on time”, 30 s; “Fluo. After”, 5 s.
Hybridization yield and melting temperature (Tm) measurement of PAEs I~III
To determine the hybridization yield, single-stranded DNA (cDNA-2/89 nt) with the complementary sequence was incubated with PAEs I~III. The mixture was then annealed by decreasing the temperature from 35 °C to 15 °C overnight. The images were analyzed with ImageJ software. A real-time PCR instrument (StepOnePlusTM, Applied Biosystems, Life Technologies, USA) was used to measure the Tm. Cy5-labeled cDNA-1 was mixed with PAEs I~III in 0.5× PBS overnight. The thermal program was started with incubation at 10 °C for 10 min, followed by increasing the temperature from 10 °C to 95 °C at 0.1 °C every 10 s.
SAXS measurements and analysis
SAXS experiments were carried out on a bio-SAXS-BL19U2 at the Shanghai Synchrotron Radiation Facility (SSRF). We used a flow-through cell with a cylindrical quartz glass capillary of 1.5-mm diameter and made measurements at 12-keV X-ray energy (wavelength 0.1033 nm). An evacuated flight tube was used to cover the sample-to-detector distance of 2.708 m. A motorized beam stop was placed inside the flight tube. A Dectris Pilatus 1 M CMOS Detector (172 × 172-µm pixel size) was used. Silver behenate was used for calibration of the sample detector distance and beam center. The exposure time was 1 s for each sample. The beam size was 0.40 × 0.15 (H × V) mm2, and a conical mask was used to restrict the azimuthal integration to the direction with higher resolution. 1D scattering-intensity profiles I(q) were obtained from circularly averaging the 2D scattering patterns and the corresponding structure factors S(q) for the studied structures: q = 4πsinθ/λ, where 2θ is the scattering angle and λ is the wavelength of the X-ray.
For SAXS data analysis, background- (measurements of 0.5× PBS buffer) subtracted scattering data were transformed with Bio-XTAS RAW 1.2 and fitted using SasView (Version 4.1.2)40. The data were further analyzed by the ATSAS 2.7.2 package. The built-in model “linear-pearls” under the category of “Sphere” was used to fit the data of DNA–AuNP dimers. This model provides the form factor for N particles (here N = 2 for dimers) of radius R linearly joined by short bonds (or segment length or edge separation) L = (A−2R). A is the center-to-center pearl separation distance. The thickness of each bond is assumed to be negligible. Since we used only one or two DNA strands to modify an AuNP, the scattering intensity contributed by DNA was neglected, and only the scattering of the AuNPs was considered. The scattering intensity was fitted with Equation (1)
where the mass mp is (SLDpearl−SLDsolvent) × (volume of N pearls). V is the total volume41. We then calculated the pair-density distribution function (PDDF) p(r) using SasView. The PDDF distribution for an AuNP dimer is expected to show two peaks. The location of the first peak is determined by the AuNP radius, while the second peak results from the center-to-center distance.
Results
Figure 1a shows the general design principle for using polyA-based DNA encoders to build DNA bonds with programmable bond length and bond energy. A typical DNA encoder consists of polyA (consecutive adenines) domains and non-polyA domains. According to previous studies35,36,37, the polyA domain has a high affinity for Au, and thus can be absorbed on the AuNP surface. The non-polyA domain (bonding domain) serves as a DNA bond exposed in the solution for hybridization. Thus, DNA encoders with different arrangements of polyA and bonding domains form DNA bonds on AuNPs with different topologic configurations. Specifically, the type I encoder (encoder I) has a 20-nt bonding domain flanked by two 20-nt polyA domains, which forms a kissing-loop motif on the AuNP for bonding. Encoder II is a polyA domain appended with a 20-nt single-stranded (ss) overhang for bonding, which is a classic configuration for DNA bonding. Encoder III is a partially double-stranded (ds) structure with two polyA domains, a 20-bp ds stem and a 20-nt sticky end for bonding (Fig. 1a). We propose that these different topological configurations can lead to different bond lengths and bond energies.
To construct monovalent PAEs with different types of DNA bonds, we separately incubated 10-nm AuNPs with the DNA encoder I–III at room temperature via a previously described approach (detailed in “Methods”). After a 15-min incubation, we used agarose gel electrophoresis to separate and recover the PAEs functionalized with the DNA bonds. The gel bands with lowered migration rates compared with those of the naked AuNPs suggest successful formation of monovalent PAEs (Supplementary Fig. S1a). Next, we incubated the PAEs carrying homotypic bonds with complementary sequences, allowing interparticle hybridization. According to the gel and TEM images (Fig. 1b, c; Supplementary Fig. S1b, c), we successfully obtained PAE dimer structures with bonds of types I–III, which verified the bonding ability of these bonds with different configurations.
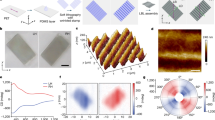
Having established the DNA bonds with programmable configurations, we further evaluated the bond length from different types of DNA bonds by measuring the PAE dimers. As shown in Fig. 2a, the expected interparticle spaces are ~2 nm (the diameter of a dsDNA), ~6.8 nm (the length of a 20-bp DNA), and ~20.4 nm (the length of a 60-bp DNA) for PAE dimers I–III, respectively. The statistical analysis of the particle interspaces from the TEM images (Supplementary Fig. S2) shows that the average interparticle distances of dimer I, dimer II, and dimer III were approximately 1.7 nm, 3.6 nm, and 6.0 nm, respectively, consistent with the expected bond-length relationship (dimer III>dimer II>dimer I), but the absolute values are smaller than expected, which might be due to the drying of the TEM samples. The DLS results (Fig. 2b) show that the hydration radii of dimers I–III were ~18 nm, ~20 nm, and ~22 nm, respectively. This size relationship is also consistent with expectations. For SAXS analysis, the SAXS-intensity patterns of the structures are plotted as functions of the magnitude of the scattering: q = 4πsinθ/λ, with wavelength λ and scattering angle 2θ (Fig. 2c). We then calculated the pair-density distribution function (PDDF) p(r) using the “linear-pearls” model (detailed in Methods). The PDDF distribution for a PAE dimer is expected to show two peaks. The position of the first peak is determined by the particle radius; the position of the second peak determines the center-to-center distance42, the variation of which can be reflected by the peak width. According to the PDDF plots (Fig. 2d), the AuNP radius (measured from the first peak position) was ~4.7 nm. For dimer I, the center-to-center distance (measured from the second peak position) was ~17.5 nm; thus, the interparticle distance was ~8.1 nm; likewise, the interparticle distances of dimer II and dimer III were ~13.1 nm and ~20.3 nm, respectively. This result is also consistent with the expected bond-length relationship. We found that the measured interparticle distance of dimer I is larger than the 2-nm theoretical interspace (the diameter of a DNA double helix, assuming that the two AuNPs were brought in proximity by the formation of an ideal dsDNA between). This phenomenon might be due to the strong steric hindrance and electrostatic repulsion between the two particles modified with BSPP and DNAs, which prevent them from being in close proximity43. We also found that the width of the second PDDF peak is positively correlated to the bond length, indicating that the longer bond shows higher bond flexibility, resulting in larger variation in interparticle distance. For a given material, the flexibility is largely determined by its geometric properties, especially the aspect ratio. Here, in this study, bond I possesses a minimal length (theoretically ~2 nm) yet a large diameter (~6.8 nm); meanwhile, bond III possesses a large length (~20.4 nm) yet a small diameter (~2 nm). Thus, bond III has a higher aspect ratio and should present higher flexibility than bond I, which is consistent with our experimental observation. Collectively, the results show that the bond-length relationship is bond III>bond II> bond I, while for bond flexibility, the relationship is reversed. In particular, compared with the commonly used linear bond configuration (bond II or bond III), bond I, relying on kissing-loop hybridization, provides the shortest bond length and the highest bond rigidity, which may facilitate fabrication of nanoparticle superstructures with higher compactness. These results suggest that polyA-based DNA bonds with programmable topological configurations can allow tailorable bond length and bond flexibility.
a Schematic of PAE dimers I–III with estimated dimensions. b DLS measurements of PAE dimers I–III (green, blue, and orange, respectively). c Measured SAXS pattern of the dimers (hollow circle, triangle, and quadrangle: experimental data; solid lines: regression curves). d PDDF plots of the corresponding structures obtained from the SAXS patterns. The first peak position reflects the AuNP-sphere radius; the second peak position reflects the center-to-center distance.
Next, we investigated the bond energies of these bonds by analyzing their hybridization and denaturation. Bond energy stands for the amount of energy required to break the bond apart. For the DNA bonds, the efficiency of DNA hybridization and denaturation can reflect their bond energy. We first analyzed the gel images of PAEs I~III incubated with complementary single-strand DNA (Supplementary Fig. S3a, b). By measuring the band intensities, we found that the yields of hybridization products (positively correlated to the bond energy) formed via bonds I–III were approximately 40, 70, and 85% (Table 1), respectively. We also used MST44 to evaluate the binding affinity of these bonds with fluorescently labeled complementary ssDNAs (Supplementary Fig. S4a–c). The dissociation constants (Kd, inversely correlated to the bond energy) derived in this manner for bonds I–III were ~8.7, 3.7, and 1.5 nM, respectively. To further measure the melting temperatures (Tm, positively correlated to the bond energy)45,46 of these bonds, we recorded their melting curves, i.e., plots of the fluorescence change rate (due to the release of fluorescent DNAs) versus increasing temperature (Supplementary Fig. S4d). The temperature corresponding to the highest fluorescence change rate can be regarded as the Tm. The Tm values measured in this way were ~52.5 °C, ~54.5 °C, and ~59 °C, corresponding to bonds I–III, respectively. Taken together, these results from three perspectives are consistent with each other (Table 1). We can conclude that the bond energy relationship of the three bond types is bond III> bond II> bond I. Notably, bond III shows the highest bond energy among the three, although the base number of the three types for hybridization (20 nt) was identical. This phenomenon might be attributed to the fact that the bonding domain (sticky end) of bond III is supported by the rigid ds stem and thus is away from the AuNP surface, leading to lower steric hindrance against DNA hybridization. The base-stacking force resulting from the ds-stem motif in bond III might also contribute to the high bond energy. Thus, bond III may be utilized for pursuing high-yield PAE assembly. Taken together, these results suggest that we can define bond energy by programming the topologic configurations of the bonds.
Having verified the assembly of monovalent PAEs mediated by homotypic bonds, we further sought to implement assembly from multivalent PAEs via heterotypic bonds (Fig. 3a; Supplementary Fig. S5). First, we sought to construct PAEs with 1–4 bonds of type III. To ensure that the AuNPs can be decorated with a prescribed number of bonds, we synthesized a series of DNA encoders with 2–4 DNA strands prior to AuNP functionalization. These multistranded encoders have 1–4 ds-stem motifs and 2–4 ss-polyA domains, which integrate 1–4 bonds of type III in one encoder structure (Supplementary Fig. S6). Thus, monomodification of AuNPs with these encoders can generate PAEs with prescribed valency (valence = 1–4). Next, we incubated these 10-nm PAEs of different valences with 5-nm monovalent PAEs with complementary bond II. In the gel and TEM images (Fig. 3b; Supplementary Fig. S7), we observed nanoparticle clusters with one 10-nm nanoparticle and 1–4 5-nm nanoparticles, corresponding to the 10-nm PAEs of valence 1–4, respectively, and the yields for these two- to five-particle structures were approximately 56.3, 40.2, 23.5, and 20.3%, respectively (Supplementary Fig. S7). These results suggest that we can use the preassembled DNA encoders to define the precise bond number as well as bond type on each AuNP, which allows the assembly of finite nanoparticle clusters with discrete PAE numbers.
Finally, we demonstrate ionic strength-controlled hierarchical assembly of nanoparticle oligomers assembled from PAEs with bonds at different levels in different directions. As shown in Fig. 4a, we designed a DNA encoder with two bonding domains and two polyA domains, allowing the combination of two bonds of different types (bond I and bond II) in one structure. Thus, functionalization of AuNPs with the DNA encoder would generate an asymmetric PAE that has two bonds with different bond lengths and bond energies in different directions. We synthesized two complementary PAEs (PAE-A and PAE-B) of this kind. Bond I on PAE-A was complementary to bond I’ on PAE-B, as was bond II to bond II’. It is known that DNA hybridization is highly dependent on the strength of cationic ions. Given that bond I and bond II have different bond energies, they require different ionic strengths for effective hybridization. Thus, we propose that by tuning the ionic strength of the solution, the two bond types can carry out sequential bonding, allowing hierarchical assembly.
a Schematic illustration of oligomer structures formed by AuNP A and B modified with bonds I and II. Bonds I and II are complementary I’ and II’, respectively. b Agarose gel electrophoresis of particle assemblies with different Na+ concentrations (5 mM, 20 mM, and 50 mM). c, d Representative TEM images and statistical distributions of oligomer length corresponding to different Na+ concentrations (N > 100). Scale bar, 20 nm.
To test this idea, the two PAEs were incubated in solutions with increasing Na+ concentrations (from 5 mM to 100 mM) at room temperature for 24 h. We then separated the products of different particle numbers by agarose gel electrophoresis (Fig. 4b; Supplementary Fig. S8a) and used TEM to verify the PAE assemblies (Fig. 4c; Supplementary Fig. S8b). We found that with 5 mM Na+, there was only one intense band in the gel with a high migration rate. The structures under this condition were found to be mostly PAE monomers under TEM, suggesting that neither bond I nor bond II could hybridize effectively under this ionic strength. Under 20 mM Na+, a band with a lower migration rate appeared. The TEM image shows that there were many PAE dimers, suggesting that under this ionic strength, bond II and bond II’ could undergo hybridization, while bond I and bond I’ could not, because bond II has a higher bond energy than bond I. When the Na+ concentration was increased to 50 mM, we observed the formation of oligomer structures, especially four-particle oligomers, which likely resulted from the bond-I-mediated assembly of the dimers, consistent with the appearance of an additional band with slower migration in the gel. The statistics of particle numbers in the oligomers (Fig. 4d) verified that the particle number increased with increasing Na+ concentration. Thus, we can implement sequential bonding of bond II and bond I by increasing the Na+ concentration. The bond-II-mediated hybridization under lower Na+ concentrations can be regarded as the first-level assembly, which forms dimer structures, while the bond-I-mediated reactions under higher Na+ concentrations can be regarded as the second-level assembly, forming oligomers (particle number ≥ 4). These results suggest that discrete bonds with programmable bond energies allow hierarchical assembly of PAEs.
Discussion
Here, we report a strategy to program DNA bond length and bond energy on PAEs using polyA-based DNA encoders. We created three types of bonds with distinct structural configurations. Bond I possesses a kissing-loop motif for bonding, bond II provides an ss overhang, and bond III has a ds-stem motif with a sticky end. We have shown that these bonds have different bond lengths and different bond energies, which are highly dependent on their topological configurations. As a result, bond I can provide compact and rigid assembly, while bond III shows high bonding efficiency. We demonstrated that these bonds at different levels in different directions allow ionic strength-controlled hierarchical assembly of multiparticle structures.
These DNA bonds show several advantages. First, they enable programming of bond length and energy in new dimensions, providing increased freedom in design. Second, these bonds allow rational control of other structural properties (e.g., steric volume and structural flexibility) as well, which may enrich the library of programmable DNA bonds. Moreover, this strategy is more facile than strategies that employ complex DNA nanostructures34 because the AuNPs are also involved in bond configurations via the polyA–Au interaction. The hierarchical assembly of plasmonic nanoparticles may have implications in the fabrication of large materials exhibiting photoelectric properties arising from nanoscopic structures (e.g., plasmon coupling, Fano resonances, and chiral response)11,47,48, which have great potential in applications such as photoelectric devices and biosensors.
References
Sanger, J. M., Mittal, B., Pochapin, M. & Sanger, J. W. Observations of microfilament bundles in living cells microinjected with fluorescently labelled contractile proteins. J. Cell Sci. 1986, 17–44 (1986).
Lane, T., Serwer, P., Hayes, S. J. & Eiserling, F. Quantized viral-DNA packaging revealed by rotating gel-electrophoresis. Virology 174, 472–478 (1990).
Bednar, J. et al. Nucleosomes, linker DNA, and linker histone form a unique structural motif that directs the higher-order folding and compaction of chromatin. Proc. Natl Acad. Sci. USA 95, 14173–14178 (1998).
Routh, A., Sandin, S. & Rhodes, D. Nucleosome repeat length and linker histone stoichiometry determine chromatin fiber structure. Proc. Natl Acad. Sci. USA 105, 8872–8877 (2008).
Conde, C. & Caceres, A. Microtubule assembly, organization and dynamics in axons and dendrites. Nat. Rev. Neurosci. 10, 319–332 (2009).
Ke, X., Zhong, G. & Zhuang, X. Actin, spectrin, and associated proteins form a periodic cytoskeletal structure in axons. Science 339, 452–456 (2013).
Garzoni, M., Okuro, K., Ishii, N., Aida, T. & Pavan, G. M. Structure and shape effects of molecular glue on supramolecular tubulin assemblies. ACS Nano 8, 904–914 (2014).
Luger, K., Mäder, A. W., Richmond, R. K., Sargent, D. F. & Richmond, T. J. Crystal structure of the nucleosome core particle at 2.8 Å resolution. Nature 389, 251–260 (1997).
Suto, R. K., Clarkson, M. J., Tremethick, D. J. & Luger, K. Crystal structure of a nucleosome core particle containing the variant histone H2A.Z. Nat. Struct. Biol. 7, 1121–1124 (2000).
Davey, C. A., Sargent, D. F., Luger, K., Maeder, A. W. & Richmond, T. J. Solvent mediated interactions in the structure of the nucleosome core particle at 1.9Å resolution. J. Mol. Biol. 319, 1097–1113 (2002).
Schreiber, R. et al. Hierarchical assembly of metal nanoparticles, quantum dots and organic dyes using DNA origami scaffolds. Nat. Nanotechnol. 9, 74–78 (2014).
Park, S. Y. et al. DNA-programmable nanoparticle crystallization. Nature 451, 553–556 (2008).
Jones, M. R. et al. DNA-nanoparticle superlattices formed from anisotropic building blocks. Nat. Mater. 9, 913–917 (2010).
Macfarlane, R. J., O’Brien, M. N., Petrosko, S. H. & Mirkin, C. A. Nucleic acid-modified nanostructures as programmable atom equivalents: forging a new “Table of Elements”. Angew. Chem. Int. Ed. 52, 5688–5698 (2013).
Zhang, C. et al. A general approach to DNA-programmable atom equivalents. Nat. Mater. 12, 741–746 (2013).
Auyeung, E. et al. DNA-mediated nanoparticle crystallization into wulff polyhedra. Nature 505, 73–77 (2014).
Lu, F., Yager, K. G., Zhang, Y., Xin, H. & Gang, O. Superlattices assembled through shape-induced directional binding. Nat. Commun. 6, 6912 (2015).
Zhang, Y. et al. Selective transformations between nanoparticle superlattices via the reprogramming of DNA-mediated interactions. Nat. Mater. 14, 840 (2015).
Kim, Y., Macfarlane, R. J., Jones, M. R. & Mirkin, C. A. Transmutable nanoparticles with reconfigurable surface ligands. Science 351, 579–582 (2016).
Liu, W. et al. Diamond family of nanoparticle superlattices. Science 351, 582–586 (2016).
Tian, Y. et al. Lattice engineering through nanoparticle-DNA frameworks. Nat. Mater. 15, 654–661 (2016).
Senesi, A. J. et al. Oligonucleotide flexibility dictates crystal quality in DNA-programmable nanoparticle superlattices. Adv. Mater. 26, 7235–7240 (2014).
Thaner, R. V. et al. The significance of multivalent bonding motifs and “bond order” in DNA-directed nanoparticle crystallization. J. Am. Chem. Soc. 138, 6119–6122 (2016).
Kraft, D. J. et al. Surface roughness directed self-assembly of patchy particles into colloidal micelles. Proc. Natl Acad. Sci. USA 109, 10787–10792 (2012).
Wang, Y. et al. Colloids with valence and specific directional bonding. Nature 491, 51–55 (2012).
Feng, L., Dreyfus, R., Sha, R. J., Seeman, N. C. & Chaikin, P. M. DNA patchy particles. Adv. Mater. 25, 2779–2783 (2013).
Edwardson, T. G. W., Lau, K. L., Bousmail, D., Serpell, C. J. & Sleiman, H. F. Transfer of molecular recognition information from DNA nanostructures to gold nanoparticles. Nat. Chem. 8, 162–170 (2016).
Liu, W. Y., Halverson, J., Tian, Y., Tkachenko, A. V. & Gang, O. Self-organized architectures from assorted DNA-framed nanoparticles. Nat. Chem. 8, 867–873 (2016).
Zhang, Y. et al. Transfer of two-dimensional oligonucleotide patterns onto stereocontrolled plasmonic nanostructures through DNA-origami-based nanoimprinting lithography. Angew. Chem. Int. Ed. 55, 8036–8040 (2016).
Xing, H. et al. Bottom-up strategy to prepare nanoparticles with a single DNA strand. J. Am. Chem. Soc. 139, 3623–3626 (2017).
Yang, Z., Liu, H. & Liu, D. Spatial regulation of synthetic and biological nanoparticles by DNA nanotechnology. NPG Asia Mater. 7, e161 (2015).
Lan, X. & Wang, Q. DNA-programmed self-assembly of photonic nanoarchitectures. NPG Asia Mater. 6, e97 (2014).
Aldaye, F. A. & Sleiman, H. F. Dynamic DNA templates for discrete gold nanoparticle assemblies: control of geometry, modularity, write/erase and structural switching. J. Am. Chem. Soc. 129, 4130–4131 (2007).
Shen, C. et al. Site-specific surface functionalization of gold nanorods using DNA origami clamps. J. Am. Chem. Soc. 138, 1764–1767 (2016).
Yao, G. et al. Programming nanoparticle valence bonds with single-stranded DNA encoders. Nat. Mater. https://doi.org/10.1038/s41563-019-0549-3 (2019).
Pei, H. et al. Designed diblock oligonucleotide for the synthesis of spatially isolated and highly hybridizable functionalization of DNA-gold nanoparticle nanoconjugates. J. Am. Chem. Soc. 134, 11876–11879 (2012).
Yao, G. et al. Clicking DNA to gold nanoparticles: poly-adenine-mediated formation of monovalent DNA-gold nanoparticle conjugates with nearly quantitative yield. NPG Asia Mater. 7, e159 (2015).
Hartman, M. R. et al. Thermostable branched DNA nanostructures as modular primers for polymerase chain reaction. Angew. Chem. Int. Ed. 52, 8699–8702 (2013).
Lin, C. et al. In vivo cloning of artificial DNA nanostructures. Proc. Natl Acad. Sci. USA 105, 17626–17631 (2008).
Nielsen, S. S. et al. BioXTAS RAW, a software program for high-throughput automated small-angle X-ray scattering data reduction and preliminary analysis. J. Appl. Crystallogr. 42, 959–964 (2009).
Dobrynin, A. V., Rubinstein, M. & Obukhov, S. P. Cascade of transitions of polyelectrolytes in poor solvents. Macromolecules 29, 2974–2979 (1996).
Hartl, C. et al. Position accuracy of gold nanoparticles on DNA origami structures studied with small-angle X-ray scattering. Nano Lett. 18, 2609–2615 (2018).
Wang, H., Li, Y., Liu, M., Gong, M. & Deng, Z. Overcoming the coupling dilemma in DNA-programmable nanoparticle assemblies by “Ag+ soldering”. Small 11, 2247–2251 (2015).
Damke, P. P. et al. Identification of the periplasmic DNA receptor for natural transformation of Helicobacter pylori. Nat. Commun. 10, 5357 (2019).
Hill, H. D., Hurst, S. J. & Mirkin, C. A. Curvature-induced base pair “slipping” effects in DNA-nanoparticle hybridization. Nano Lett. 9, 317–321 (2009).
Randeria, P. S. et al. What controls the hybridization thermodynamics of spherical nucleic acids? J. Am. Chem. Soc. 137, 3486–3489 (2015).
He, Y. et al. Hierarchical self-assembly of DNA into symmetric supramolecular polyhedra. Nature 452, 198–201 (2008).
Liu, N. & Liedl, T. DNA-assembled advanced plasmonic architectures. Chem. Rev. 118, 3032–3053 (2018).
Acknowledgements
This work was financially supported by the National Key R&D Program of China (2016YFA0201200), National Natural Science Foundation of China (21834007, 21675167, 31571014, 21775157, and 11575278), the Shanghai Municipal Science and Technology Commission (19JC1410302), the Open Large Infrastructure Research of the Chinese Academy of Sciences, the LU JIAXI International team programme supported by CAS, and the K.C. Wong Education Foundation, Shanghai Jiao Tong University. The authors are also thankful to the staff from the BL19U2 beamline of the National Facility for Protein Science Shanghai (NFPS) at the Shanghai Synchrotron Radiation Facility (SSRF) for providing assistance during data collection.
Author information
Authors and Affiliations
Contributions
J.L. and L.W. directed the research. J.L., C.F., and G.Y. conceived the study. X.C., G.Y., and Q.L. performed the experiments. X.C., R.L., and Y.L. performed TEM imaging and analysis. X.L., X.C., and H.W. performed SAXS experiments. J.L., X.C., X.L., Q.L., L.W., and C.F. analyzed the data and wrote the paper.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Chen, X., Liu, X., Yao, G. et al. PolyA-based DNA bonds with programmable bond length and bond energy. NPG Asia Mater 12, 49 (2020). https://doi.org/10.1038/s41427-020-0231-x
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41427-020-0231-x
This article is cited by
-
DNA framework-engineered chimeras platform enables selectively targeted protein degradation
Nature Communications (2023)
-
Poly-adenine-mediated spherical nucleic acids for interfacial recognition of kanamycin
Microchimica Acta (2022)