Abstract
Radiation exposure of healthy cells can halt cell cycle temporarily or permanently. In this work, we analyze the time evolution of p21 and p53 from two single cell datasets of retinal pigment epithelial cells exposed to several levels of radiation, and in particular, the effect of radiation on cell cycle arrest. Employing various quantification methods from signal processing, we show how p21 levels, and to a lesser extent p53 levels, dictate whether the cells are arrested in their cell cycle and how frequently these mitosis events are likely to occur. We observed that single cells exposed to the same dose of DNA damage exhibit heterogeneity in cellular outcomes and that the frequency of cell division is a more accurate monitor of cell damage rather than just radiation level. Finally, we show how heterogeneity in DNA damage signaling is manifested early in the response to radiation exposure level and has potential to predict long-term fate.
Similar content being viewed by others
Introduction
Signal transduction pathways modulate cellular behavior in response to changes in the internal state of a cell or its environment. These pathways are constituted by chains of interacting signaling molecules. Changes in the abundance, activity or localization of a particular molecule propagates to downstream components, ultimately connecting signals and cellular outcomes, such as proliferation, cell death and cell migration. Decades of research in molecular and cellular biology delineated signal transduction pathways that mediate response to growth factors stimulation [1], DNA damage [2, 3], changes in nutrient abundance [4] and oxygen availability [5], among others. Altogether, the qualitative dissection of signal transduction pathways provides a blueprint to understand how cells integrate information and adapt to changing environments.
However, a static and qualitative account of signal transduction fails to consider the dynamic and heterogeneous character of this process. These two features are highlighted by experiments that quantify signal transduction as it plays out in real time. The use of fluorescent reporters for the abundance and activity of key signaling molecules revealed pervasive heterogeneity in the response of individual cells to particular stimuli, and intricate patterns in the temporal evolution of signaling, ranging from all-or-none activation to frequency-modulated pulses and persistent oscillations [6, 7]. More recently, expression of multiple fluorescent reporters in the same cell advanced the possibility to quantify the way changes in cellular signaling propagate through biological networks, and ultimately influence cell fate [8,9,10,11,12]. In principle, these experiments should allow extraction of detailed quantitative features of the interaction between pairs of molecular species in a cascade, including time delays. Moreover, they should allow quantification of the extent to which variability in upstream observables can explain variability in downstream components in the pathway and cellular behaviors. These quantitative data are important since they impose constraints on quantitative dynamical models of signal transduction and are fundamental for understanding how reliably can cells process information of the presence and amount of particular signals.
Here, we develop mathematical/signal processing methods to estimate time delays and quantitative relations between pairs of nodes in signal transduction pathways. As a proof-of-principle, we use publicly available single cell time lapse imaging data on the response of individual mammalian cells to DNA damage [9, 13]. These data focus on the dynamics of the tumor suppressive transcription factor p53, and those of its downstream transcriptional target, the cell cycle inhibitor p21. In addition, we have information of the number of divisions that each cell goes through in the course of 5 days post-DNA damage, forming a three-node cascade that connects activation of p53 signaling with cell fate. These data are an ideal ground to test our methods for multiple reasons. Firstly, the pairwise interactions that we are studying (i.e., p53-p21 and p21-division) are well established in the field [14], providing a ground truth for the directionality in the quantitative relations which we uncover. Secondly, trajectories are long, enabling robust estimation of relations using multiple data points per cell and over distinct timescales (hours and days). Lastly, experiments were conducted over a range of DNA damage doses that cover the entire spectrum of cell division patterns that have been described in the literature, allowing us to test over a wide dynamic range of inputs and outputs. Collectively, these data provide a unique test case for novel mathematical data analysis aimed at dissecting quantitative interactions in signaling cascades.
The tumor suppressor p53 is often referred to the as the “guardian of the genome” for its importance in regulating a large number of target genes, its role in regulating cell cycle, inducing cell apoptosis and DNA repair, as well as senescence. An in-depth understanding of the dynamics of the p53-p21 couple is of crucial importance in understanding why a given cell may decide to continue dividing, especially in the context of cancer [15]. While not a focus in this work, the known relationship of p53 and Mdm2 is of particular importance as it gives rise to the observed oscillatory behavior in the p53 dynamics and indirectly in the dynamics of p21 [16,17,18,19,20,21,22,23]. There are also, of course, numerous other important targets of p53 such as p14ARF, Chk2, Wip1, and ATM [21, 23,24,25], which could be studied in similar fashion using the techniques introduced in this work. Cellular senescence as well as temporary instances of cell cycle arrest are also of great interest as potential therapeutic targets through the prevention of cancerous cells from proliferating [26,27,28]. Notably, for cells to remain arrested, this signaling pathway needs to be maintained [9, 28,29,30]. Among a large number of targets that p53 influences, the cyclin-dependent kinase (CDK) inhibitor p21 plays a crucial role in the ability of cells to undergo cell cycle arrest [31,32,33]. p21 interacts with CDK2 as a bistable switch with increased levels of p21 leading to cell cycle arrest, while low levels of p21 allow cells to escape G1 phase arrest [9, 34,35,36,37].
The signal processing methods described in the present paper may be used for the analysis of a number of biological phenomena described by time series, which in the present paper is dynamics of p21 and p53 in single cell datasets after radiation exposure. In particular, the proposed techniques give exact quantitative information concerning the oscillation periods of p53 and p21 as well as the signaling delay between these two measured signals, and a quantitative relationship between p21/p53 levels and the frequency of mitosis. Finally, the methods seem to indicate that the p21 and p53 dynamics cluster by the total number of observed divisions rather than radiation levels.
Methods
In this section, we give the details of the quantitative methods that were used to obtain our results using dynamic time warping and cross-correlation in order to find delays between the signals, as well as describing our use of moving averages to study the long-term trajectories of p21 and p53.
Measuring delay between two interacting oscillatory signals
Signal detrending and amplitude normalization
The preprocessing step for analyzing the relationship between p53 and p21 signals consisted of detrending the data. This can be done by fitting polynomials, preferably of low order, to the time series data. By finding an appropriate order that captures the trend of the data, one can then subtract the “trend” from the actual data to obtain the oscillatory component. In dealing with signals such as p53 and p21 that can be subjected to abrupt changes when the cells are about to divide or are undergoing division, the detrending should be applied to subsets of the overall time series data that avoids the abrupt changes.
We were able to obtain cleaner results using a method introduced in our previous work [38]. The first part of this technique, called the Detrended Autocorrelation Periodicity Scoring (DAPS) algorithm, focuses on the detrending and amplitude normalization using a sliding window embedding, which can be briefly summarized in the following steps:
-
1.
Given the sliding window of length M of an N-length time series x, the sliding windows are arranged into the columns of an M × (N − M + 1) matrix X, so that the ith column of X is SWM[x]i.
$$X = \left[ {\begin{array}{*{20}{l}} {x_1} \hfill & {x_2} \hfill & {x_3} \hfill & \cdots \hfill & {x_{N - M + 1}} \hfill \\ {x_2} \hfill & {x_3} \hfill & {x_4} \hfill & \cdots \hfill & {x_{N - M + 2}} \hfill \\ {x_3} \hfill & {x_4} \hfill & {x_5} \hfill & \cdots \hfill & {x_{N - M + 3}} \hfill \\ \vdots \hfill & \vdots \hfill & \vdots \hfill & \cdots \hfill & \vdots \hfill \\ {x_M} \hfill & {x_{M + 1}} \hfill & {x_{M + 2}} \hfill & \cdots \hfill & {x_N} \hfill \end{array}} \right]$$In other words, the ith skew-diagonal of X is a constant value equal to the ith value of the time series xi. In this work, M is chosen to be 11 data points, or 5.5 h, which is roughly the length of one period of p53 [20].
-
2.
The mean of each column of X is subtracted from each element of that column. This normalizes for linear drift in the time series.
-
3.
After point-centering, each column is normalized to be of unit norm. This controls for changes in amplitude in the signal.
-
4.
Lastly, the following operation is performed on the matrix
\(Y_{i,j} = {{{{{{{\mathrm{mean}}}}}}}}_k(\hat X_{i + k,j - k})\)with
\(1 \le i + k \le M,\)
\(1 \le j - k \le N - M + 1.\)
Finally, the ith value of the time series y is then simply the value of any of the elements in the ith skew diagonal of Y.
Quantifying delay using dynamic time warping and cross-correlation
Dynamic Time Warping (DTW) [39] is a method that measures the similarity between two time series by finding an optimal time-ordered correspondence, known as a “warping path,” which best maps every index in the first time series to every index in the second time series [40]. Once the signals have been properly detrended and normalized, we apply DTW to determine which of the proteins p53 or p21 leads or lags, as well as the actual delay between the two time series. It is likely that, because of the periodic nature of these signals, the algorithm may pick up the real delay as well as this value adjusted by integer multiples of the period of oscillation. The “warping path” of a pure delay is expected to be a straight line in this analysis with the true delay being a stronger signal than the “secondary” alignments that are offset by a certain number of periods.
Similarly, cross-correlation may be used to determine a correspondence between two time series by shifting one relative to the other. This work introduces the concept of using DTW as an alternative to cross-correlation to capture signal delays, with the hope that other applications may benefit from it even though similar results were obtained using both techniques.
Long-term trend in the signals and occupancy density
The analysis of long-term trends in the signals is performed by using moving averages (m.a.) [41]. By removing the short-term fluctuations, this transformation acts as a low-pass filter that allows highlighting the trend from the overall signal. Using a relatively small window of 9 data points or 4.5 h, all the time series shown in this work were cleanly smoothed. Since the amplitude of the oscillations may be quite large relative to the long-term trend, especially for p53, previous studies focused largely on describing the behavior of the oscillations. A primary argument of this work is to show that focusing on the trend and not the oscillations for p21 and p53 provides important and novel insights about the current state of a cell as it pertains to cell cycle arrest.
Using moving average values of p21 and p53, 2-D maps of occupancy density were constructed to better understand where these values tend to lie with respect to their likelihood of undergoing mitosis or treatment conditions. By constructing a vector of these moving averages for all time points, each time point then takes up a (x,y) coordinate in space based on (p53 m.a.,p21 m.a.). In this work, a grid was delimited by approximately the minimum and maximum values for each of the axes. Due to the large discrepancy of low and high values as well as the exponential nature of chemical reactions, a log10 (p21 m.a.) vs. log10 (p53 m.a.) was used and discretized by 0.1 increment on this scale. This discretized grid is used to count each time a coordinate lies within one of the cells. This is then normalized into the final occupancy density maps.
Results
Experimental single cell datasets and general trends
The analysis in this work is based on two datasets of individual human cells that were published in [9, 13]. These retinal pigment epithelial cells were monitored using fluorescence live-cell imaging for about a week after being subjected to radiation. The cultured cells were not cancerous and overexpressed telomerase in order to make them as normal as possible in comparison to in vivo cells. The cells had wild-type p53 and tended to exit through cell cycle arrest rather than apoptosis during the monitoring period.
The single cells in the first dataset were subjected to x-ray radiation at 0 Gy, 0.5 Gy, 1 Gy, 2 Gy, 4 Gy, and 8 Gy. Fluorescence markers for p21, p53, and geminin, as well as mitosis events, were monitored throughout the experiment. The values used in this work are fluorescence intensities (a.u.) of p53-mNeonGreen reporter for p53, the mKate2 protein to form a p21-mKate2 complex, and the CFP-hGeminin(1-110) reporter for cell cycle progression [42]. This geminin reporter indicates cell cycle transition by its degradation in G1 phase, accumulation in G1/S transition, and sharp degradation in M phase. The single cells in the second dataset were subjected to gamma irradiation at 0 Gy, 2 Gy, 4 Gy, and 10 Gy. The culture media was replenished daily, except for a subset of the cells subjected to 10 Gy irradiation that had their culture media preserved throughout the experiment. Fluorescence markers for p21 and p53, as well as the mitosis events, were tracked throughout the experiment. Since the results were quite consistent between the two types of irradiation, the x-ray results are presented in the main text, while the corresponding gamma results are provided in the Supplementary Materials. Mitosis events were recorded in a semi-automated manner using an algorithm on the recordings of the cells during the observation period [9]. Following each mitosis event, only one of the two cells was used for tracking further values in geminin, p21, and p53.
The time series data are presented in Fig. 1 for the single cells exposed to various grades of x-ray radiation and in Fig. S1 for cells exposed to gamma radiation. As shown in Fig. 1a and Fig. S1a, between two-thirds and three-quarters of the unirradiated cells divide 3–5 times during the observation period of 5 days. As the radiation levels increase from 0 to 0.5 to 1 Gy in the x-ray dataset, the fraction of cells that divide often progressively diminishes and is mostly replaced by cells that do not divide or divide only once. Due to the two datasets being exposed to different radiation sources, and the difference in intensity of the fluorescent light source at the time of the experiment, we decided to analyze the datasets independently, and we focused on results that were conserved between the experimental conditions.
Irradiated cells tend to exhibit oscillations in p53 with a short period on the order of hours [43]. There is also a longer-term oscillation observable in the data for unirradiated cells that is cell cycle dependent, as shown in panels Fig. 1b, c and Fig. S1c, d. There are, however, confounders, such as daily media changes during the experiments and the fact that cells tend to auto-fluoresce prior to mitosis that makes this observation inconclusive from the existing data. We note that cells that divide only once during the 5 days tend not to exhibit strong oscillations after mitosis. These cells tend to have dramatic increases in p53 levels observed after undergoing their sole mitosis event if they divide at all, an observation consistent with previous reports [13]. Cells that do not divide over the period of 5 days tend to exhibit lower amounts of p53 oscillations.
p21 levels exhibit much lower amounts of oscillations, and the trends in the data are of much greater importance to the local fluctuations. In other words, p21 does oscillate similarly to p53, but these oscillations constitute a smaller fraction of the overall signal. Upticks in CFP-hGeminin(1-110) correlates with temporal drops in p21 levels (shown in Fig. 1c, d). This is because this geminin reporter only accumulates during the S/G2/M parts of the cell cycle [44, 45] and p21 is degraded in the G1/S transition [33, 43]. In addition to these “gaps” in p21 during the G1/S phases, p21 levels are extremely low in unirradiated cells. p21 levels also rapidly converge to very high values for cells that undergo cell cycle arrest and never divide in the course of the experiment.
We note through additional analysis in Supplementary Sections S1 and S2 that the dynamics of p21 and p53 are not easily clustered by radiation levels. This is because even if single cells are irradiated at the same radiation level, the amount of damage they receive may vary widely. However, we show there that one can cluster p21 and p53 dynamics by the number of divisions over five days, which more closely reflects the actual damage that individual cells received, using a t-Distributed Stochastic Neighbor Embedding (t-SNE) classification technique employing the 1-Wasserstein distance. Thus, the remainder of the paper places particular emphasis on analyzing the dynamics with respect to the number of observed divisions as opposed to focusing on radiation levels which contain very heterogeneous cell behaviors.
Quantifying the delay between p21 and p53 signaling
Given that p21 lies downstream of the p53 signaling pathway, part of the observed fluctuations in p21 is directly influenced by the oscillatory component of p53. By detrending and normalizing the two signals via the Detrending Autocorrelation Periodicity Scoring (DAPS) methodology (see Section 3 below), as shown in Fig. 2a for the gamma radiation dataset, these transformed signals were used to calculate the delay between the two signals by warping or “aligning” them to one another using Dynamic Time Warping (DTW) (Section 3). By performing this step for all time series in the gamma radiation dataset, the aggregated warping paths are shown in Fig. 2b. There are mostly two main visible straight lines that cut across the graph, one indicating that the p21 signaling is lagging behind the p53 signaling by about 3.5 h and the other indicating that p21 leads p53 by 2.5 h. Because the two signals are periodic, the latter result is a natural artifact that arises from the fact that the patterns of transformed signals repeat themselves. Thus, the “correct alignment” offset by full periods of the signal will yield a local maximum in terms of correlation between the compared signals.
a, b Show histograms of the parameterizations uncovered via dynamic time warping between the detrended p53 and p21 time series, as a function of number of divisions. In (b), the normalized crosscorrelation between the two detrended time series is shown. In both cases, p53 leads p21 by 3.5 h or lags p21 by 2.5 h more often for fewer divisions, which is consistent with the period of about 6 h of p53 during cell damage.
Both the normalized cross-correlation and DTW results shown in Fig. 2b, c indicated a stronger correlation when aligning with p53 leading p21 by about 3.5 h. This holds whether looking at cells that did not divide, or divided 1, 2, or 3 times during the 5 days following radiation exposure. This is also identical to trends observed in similar results generated for the gamma radiation dataset that are shown in Fig. S6. These observations do not hold when considering cells that divided 3 + times over 5 days whether one looks at the gamma or x-ray radiation datasets. This can be partially explained by the p21 signaling being suppressed during the G1-S phases, which represent a major fraction of the cell cycle, reducing the ability to compare the two signals. The oscillations are also noisier when cells are dividing at the pace of healthy cells. Thus, these single cell datasets containing a majority of cells for which the rate of mitosis was slowed down, or for which cells were arrested, made this analysis on the delay most appropriate.
Long-term p21 and p53 trends play a key role in determining cell behavior vis-à-vis of mitosis
In Fig. 3a for the x-ray radiation dataset and Fig. S4a for the gamma radiation dataset, using a moving average (m.a.) window of 4.5 h, p21 m.a. values are plotted as a function of their respective p53 m.a. values at t = 0,12.5,25, and 50 h. After 50 h, the m.a. values of p21 and p53 remain fairly consistent. p53 values settle in a matter of hours as opposed to 1 to 2 days for the p21 values after irradiation. Videos of the datasets that show the evolution at every measured time point over 5 days are available at https://github.com/phongatran/p21p53/. There is a slight long-term drift in the two signals that makes the data points slowly disperse, which may potentially be due to a loss in fluorescence signal over time.
a Time lapses of the single cells of the moving average values of p21 vs. p53 at different time points of 0, 12.5, 25, and 50 h after irradiation. The panels are broken down either by number of total divisions over five days or by irradiation conditions. b Occupancy density of the cells with a breakdown by the total number of undergone divisions over five days. The numerical value shown in color indicates the probability for a cell to occupy a given square of 0.1 by 0.1 in the log10(p21 m.a.) vs. log10(p53 m.a.) space in a given hour.
By aggregating the m.a. values of these two markers, the occupancy densities are shown in Fig. 3b for the x-ray radiation dataset and Fig. S4b for the gamma radiation dataset. These depictions suggest that p21 levels play a more central role in determining how often cells divide over the course of 5 days after irradiation. There is a period of about a day after irradiation before the p21 levels reflect the changes due to irradiation exposure. Additional plots are available in Fig. S5, illustrating the occupancy densities broken down by radiation levels for both datasets.
To further understand the relationship between the trends in p21 and p53 with respect to cell cycle arrest, the m.a. values are centered around mitosis in Fig. 4 for cells that divide and are exposed to x-ray radiation. In Fig. 4a, the cells that do not divide have a characteristic jump in p53 m.a. values, which then subside within the first few hours after the cell receives the irradiation damage. The p21 m.a. values are constantly increasing for the next 20–30 h until they reach a plateau of about ∼3000–5500. These non-dividing cells remain in this state with little variation in geminin, p21, or p53 m.a. values.
The middle line represents the 50 percentile value for a given relative time, while the top and bottom of the shaded region represent the 75 percentile and 25 percentile values. Note that in plots (b) and (c), the cells dividing once over five days are divided into two groups: early and late single-divider cells. In (i), the mitosis events for cells dividing more than three times are aggregated and averaged out.
Single-divider cells, shown in Fig. 4b, c, undergo the process of mitosis at significantly higher values of p21 m.a. values than cells that divide more often. These cells also seem to exhibit a substantial increase of both p21 and p53 m.a. values before mitosis. These elevated levels of p53 subsist long after these cells undergo their single mitosis event. Of note, the behavior between “early single-divider cells”, defined as cells which undergo mitosis less than 3 days after exposure, exhibit distinct behavior from “late single-divider cells” (also called “escapers” in ref. [9]). Early-single divider cells have noticeably lower levels of p21 and p53. There is also a significant jump in p53 levels in late single-divider cells near the mitosis event and afterward.
Looking in detail at cells that divided exactly twice (Fig. 4d, e) or 3 times (Fig. 4f–h), the profiles are almost identical for the same number of divisions across all three markers. There does not appear to be a significant difference in the long-term trends of p21 levels before the mitosis process has taken place and afterward, once the values are stabilized. Cells that divide more than 3 times over five days in Fig. 4i exhibit in aggregate the lowest levels of p21, p53, and geminin. Note that this Fig. 4i subplot shows the behavior across all observed mitosis events, unlike the other subplots.
Cells that do not divide despite high levels of geminin are shown in Fig. 5, split into 6 clusters using k-means with correlation as the distance. These subfigures show various instances in which cells that are in the process of dividing decide to forego division or have yet to divide. These cells had the characteristics of expressing geminin levels higher than 500 a.u. for a period of more than 12.5 h, but for which no division event was recorded. There were 175 cells in the x-ray dataset. It is relatively clear that a high level of p21 is a necessary condition at the moment that the cell reverses this decision and the levels of geminin start dropping without mitosis occurring. All these cells also have an early spike in p53, which may be due to the radiation exposure at t = 0. When compared to cells that do divide in the previous panel (Fig. 4), it takes longer for geminin to reach a peak, > 20 h, from the moment geminin levels start noticeably rising as opposed to 15−20 h for dividing cells. At the end of this reversion, cells return to having high m.a. values of p53 and p21, similarly to non-dividing cells as shown in Fig. 4a. From a biological standpoint, it is probable that these cells have had an interrupted division, which may result in them having an abnormal number of chromosomes (aneuploidy), such as being tetraploid. In Fig. S8, the same results as Fig. 4 are shown for the gamma radiation dataset. The geminin levels were not measured in these single cells. The moving average trends in p21 and p53 are nearly identical across both datasets.
These cells exhibited high levels of geminin for a period of more than 12.5 h (>500 a.u.) but did not undergo mitosis. These cells were then split into 6 clusters using k-means clustering with the distance used being correlation. The middle line represents the 50 percentile value for a given relative time, while the top and bottom of the shaded region represent the 75 percentile and 25 percentile values.
Due to p21 levels being suppressed during the 15–20 h leading up to mitosis (seen on Fig. 4), Fig. 6a (x-ray irradiation dataset) and Fig. S9a (gamma irradiation dataset) focuses on the mean levels of p53 and p21 across all five days but excluding the regions in which cells are actively dividing. These excluded regions were taken as the 15 h or 30 data points leading up to a mitosis event. Looking at the distributions for mean p53 of the non-dividing regions (n.d.) as well as the corresponding mean p21 n.d., there is a correspondence between higher levels of p53 and fewer divisions as well as a correspondence between higher levels of p21 and fewer divisions over the course of five days. However, this is only true for the cells which were irradiated since cells exposed with 0 Gy did not exhibit this trend. Thus, while the decision to divide or not divide is a binary decision, the frequency of mitosis stochastically correlates to the value of p21 n.d. For p53 n.d., this signal is fainter, but still exists as one would expect since we expect p53 to orchestrate the dynamics of p21. A numerical relationship was found for both the x-ray irradiation and gamma irradiation datasets.
Swarm plots of the mean values of p21 and p53 for the x-ray dataset excluding the non-dividing regions where p21 levels are suppressed considering in (a) all five recorded days and (b) only hours 18 to 36 of the recorded data. The excluded regions are the 15 h leading to a mitosis event. Each panel is broken down into mean p53 n.d. vs. number of divisions over 5 days, mean p21 n.d. vs. number of divisions, the ratio between mean p21 n.d. and mean p53 n.d. vs. number of divisions, and mean p21 n.d. vs. mean p53 n.d.
Shown in Fig. 6b and Fig. S9b are swarm plots similar to Fig. 6a and Fig. S9a but limited to the ranges of 18–36 h from the 5 days that the data was recorded after irradiation. The decision to start this range at 18 h after radiation exposure was to allow for p21 values to settle enough into the long-term trend. Notably, for most single cells, the trends that are true over the entire interval of five days also exists within this more limited range. Thus, we hypothesize that early observations of p21 n.d. and p53 n.d. levels can be used as a predictor as to how often cells are likely to divide.
The influence of cell phase at the moment of irradiation
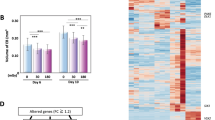
The cell population submitted to x-ray radiation were divided into two subpopulations by using a cut-off value of 100 for the geminin value at the time of irradiation. The geminin marker was employed here as the best proxy for cell cycle status at the moment of irradiation [42]. The cells were approximated to be in G1 phase under that threshold at the moment of irradiation and cells that had a threshold greater than 100 were assumed to be in S/G2 phase. Other thresholds were also considered but did not yield significant differences in our observations. By drawing histograms of key observations such as time to first division, number of total observed divisions during the 5 days following irradiation, the mean level of p53, or the mean level of p21 over that observation period, we note the differences in behavior in the two subpopulations in Fig. 7.
The overall population of x-ray irradiated cells are split into two subpopulations using a geminin cut-off value of 100. The subpopulation with cells having geminin value less or equal to 100 at the time of irradiation are shown on the four left subpanels, while the cells with geminin values higher than 100 are shown on the four right subpanels.
The key differences between those two subpopulations were that cells irradiated in G1 phase:
-
Had slower time to first division than cells irradiated in G2 or S phase;
-
Had higher number of total divisions than irradiated cells during G2 or S phase;
-
Had noticeably lower mean levels of p21 and p53 over the observation period than their G2/S counterparts.
In addition to the time to first division, which seems to simply indicate that cells irradiated later in their cell cycle divide earlier the first time, cells that are irradiated in their G2/S phases seemingly undergo more cell cycle arrest indicated by lower division rates, higher p53 levels, and higher p21 levels. Thus, we hypothesize that these cells may be suffering higher amounts of damage at similar radiation levels than their G1 irradiated counterparts.
Discussion and conclusion
Two sets of human single cells, treated with various levels of x-ray and gamma radiation, were monitored for cell division, p21, p53, and hGeminin(1-110) levels after radiation exposure [9, 13]. From the data on mitosis occurrences, it is clear that the behavior of cells within a given treatment regimen, in terms of radiation levels and type, can be quite heterogeneous with cells dividing at different rates. Using t-SNE classification with the 1-Wasserstein distance (see Supplementary Material), it was found that cells can be readily clustered using the number of cell divisions they undergo over five days, using the “histograms” of p21, and to a lesser extent, p53. This indicates that p21 dynamics plays a key role in the cell decision to undergo mitosis or not, and also affects the rate at which this occurs.
While it is known that, biologically, p21 is located downstream of the p53 signaling pathway, this work shows how p53 signaling is an integral part of the p21 response by analyzing the oscillatory components of the two signals. Through detrending and normalizing the two signals using the DAPS technique, it was shown that p53 signaling leads p21 signaling by about 3.5 h and that the period of the oscillations is about 6 h. This analysis was made possible because of the high levels of induction of p21 in damaged cells. Since p21 expression is low in the absence of radiation, and it is sharply degraded upon S-phase entry, estimation of such time delay in cells that divide normally as a result of not receiving any radiation is inconclusive.
From studying time-lapses of the moving averages of p21 and p53, we found that p53 trends tend to settle within hours of DNA damage, while p21 values may take up to a few days to fully settle into their long-term trends. Most cells that did not divide follow a pattern of expressing very high levels of p21 that builds up from the onset of the radiation.
However, in rarer cases, cells attempt to resume cell cycle progression, as evidenced by accumulation of the hGeminin(1/100) reporter and p21 degradation, but ultimately fail to undergo mitosis and establish cell cycle arrest with very high levels of p21. It is probable that these cells may have become aneuploid, since their chromosomes may have already started the process of division before being interrupted, as was previously reported [46]. The single most important characteristic of mutant p53 is aneuploidy. If these cells are only temporarily arrested in their division and eventually undergo mitosis, this may lead to catastrophic results such as cancer or trisomy. There remains an open question as to why cells would tolerate such a catastrophic scenario. Thus, while we only isolate their dynamics in this work, these cells may be of particular interest in understanding the role of p53 in cancer.
It is also made clear through this work that the cell decision to divide involves a more complex process than simply resuming division after radiation damage repair. For the datasets considered in the present work, cells that had high levels of p21 before mitosis oftentimes recovered similarly high p21 levels after mitosis. Further, we found via the mean values of p21 which exclude the parts of actively dividing cells, that the pace at which cells divide is regulated by the long-term trends of p21. This would seem to suggest that cells do not deterministically decide to cease division when considered too damaged, but that very high levels of p21 make it unlikely for cells to divide within the given observation period of five days. In summary, cell division may not be simply a binary decision which cells undertake to divide or not divide, but instead p21 levels seem to dictate how likely it is for a division to take place. Thus, long-term arrested cells may be the result of cells having extremely low probabilities of undergoing mitosis due to elevated p21 levels.
The heterogeneity of the response to DNA damage observed among cells in a culture dish, fed every day with fresh medium and growth factors, is also a function of the cell cycle stage of a given cell at the time of DNA damage [47, 48]. By using the geminin levels at the moment of irradiation, the cells were separated into a subpopulation of cells irradiated early (G1 phase) and a subpopulation of cells irradiated late in their cell cycle (G2, S, M phases). For this second subpopulation of cells irradiated later in the cell cycle, we found that levels of p21 and p53 are noticeably higher, while also observing fewer divisions over the course of the 5-days observation period. From our other analyses presented in this work, we believe that these results are consistent with a higher level of effective damage experienced by these cells. We also wish to point out that the cell cycle state at the moment of irradiation is not the sole factor determining cell fate and this is likely one of many factors that lead to the observed heterogeneity in cell fate observed in both subpopulations.
Classic studies on the function of p21 identified its role as a mediator of cell cycle arrest downstream of p53 [49,50,51]. Although it may be speculated that loss of this canonical function of p21 could confer a fitness advantage to cancer cells, with notable exceptions [52], there is a paucity of mutations affecting the p21 coding sequence [53, 54]. Furthermore, mice lacking p21 fail to develop spontaneous neoplasms within 7 months of age [55]. The low frequency of p21 genetic alterations in cancer could be explained by the prevalence of p53 alterations, as p53 orchestrates a plethora of tumor suppressive mechanisms independent of p21 [56]. In addition, studies have identified roles of p21 in the error-free progression through the cell cycle [57,58,59,60], suggesting that complete loss of this protein may be detrimental for cells in certain contexts. New methods that allow quantification of p21 protein and its relation to fate at the single cell level will allow further dissection into the roles that p21 plays in the response and recovery from DNA damage.
We should emphasize once again that while the present paper focused on the dynamics of p21/p53 using time series data, the proposed methodology employing DTW and autocorrelation with signal detrending may certainly be applied to the study of other pairs of other interacting biological oscillatory signals [61,62,63,64]. The use of moving averages can be a powerful tool to isolate long-term trends in the given signals, and hence act as an important indicator of cell state. In addition, the t-SNE using a 1-Wasserstein distance may deal with oscillatory signals better than the standard t-SNE methodology relying on traditional measures such as the Euclidean or Manhattan distances.
A potential research direction employing the proposed methodologies in the present work, is the application of these methodologies to quantify the relations in non-transformed and transformed cells [10]. Indeed, the latter analyzes similar p21/p53 relationships, but for a cancer cell line as opposed to the normal cell lines studied in this work. Other types of systems where this type of approach may be beneficial would be to study systems where the cells would be deficient in either p21 or p53, since it may unlock additional understanding on how p21 can be induced in a p53-independent manner or gain additional understanding about the way p21 deficiency may directly or indirectly p53 dynamics. We believe that the signal processing techniques introduced in this paper have the ability to reveal a deeper understanding of fundamental biological processes arising from existing and future biological experiments without necessarily adding to the burden of experimentalists.
Finally, we would like to summarize the key methodological findings of the present paper:
-
By detrending the original signals of p21 and p53 using DAPS and focusing on the oscillatory components, one may apply either the dynamic time warping or normalized cross-correlation techniques to estimate the signaling delay.
-
By smoothing the oscillatory part of the two signals using a moving average filter, we were able to describe the long-term trends of the p53 and p21 signaling after irradiation, which we utilized throughout this work. Additionally, averaging across the entire time series also brought insights on the long-term behavior of individual cells. These two approaches turned out to be especially useful dealing with signals that have a strong oscillatory component such as p53.
-
Using total divisions over the observation period as a proxy for effective radiation damage received by the individual cells, we were able to study the dynamics of these various subpopulations. Most notably, quantitative relationships between number of divisions and mean p53 value, as well as number of divisions and mean p21 value, were discovered.
Code availability
The availability of the code to reproduce the results as well as the algorithms made in this work are made available at https://github.com/phongatran/p21p53/.
Data availability
The data that support the findings of this study are available upon request through the corresponding author.
References
Dhillon AS, Hagan S, Rath O, Kolch W. MAP kinase signalling pathways in cancer. Oncogene 2007;26:3279–90.
Vogelstein B, Lane D, Levine AJ. Surfing the p53 network. Nature 2000;408:307–10.
Lakin ND, Jackson SP. Regulation of p53 in response to DNA damage. Oncogene 1999;18:7644–55.
Efeyan A, Comb WC, Sabatini DM. Nutrient-sensing mechanisms and pathways. Nature 2015;517:302–10.
Lee P, Chandel NS, Simon MC. Cellular adaptation to hypoxia through hypoxia inducible factors and beyond. Nat Rev Mol Cell Biol. 2020;21:268–83.
Purvis JE, Lahav G. Encoding and decoding cellular information through signaling dynamics. Cell 2013;152:945–56.
Kolch W, Halasz M, Granovskaya M, Kholodenko BN. The dynamic control of signal transduction networks in cancer cells. Nat Rev Cancer. 2015;15:515–27.
Albeck JG, Mills GB, Brugge JS. Frequency-Modulated Pulses of ERK activity transmit quantitative proliferation signals. Mol Cell. 2013;49:249–61.
Reyes J, Chen JY, Stewart-Ornstein J, Karhohs KW, Mock CS, Lahav G. Fluctuations in p53 signaling allow escape from cell-cycle arrest. Mol Cell. 2018;71:581–91.
Hafner A, Reyes J, Stewart-Ornstein J, Tsabar M, Jambhekar A, Lahav G. Quantifying the central dogma in the p53 pathway in live single cells. Cell Syst. 2020;10:495–505.
Bothma JP, Norstad MR, Alamos S, Garcia HG. LlamaTags: A versatile tool to image transcription factor dynamics in live embryos. Cell 2018;173:1810–1822. e16
Yang JM, Chi WY, Liang J, Takayanagi S, Iglesias PA, Huang CH. Deciphering cell signaling networks with massively multiplexed biosensor barcoding. Cell 2021;184:6193–206.
Tsabar M, Mock CS, Venkatachalam V, Reyes J, Karhohs KW, Oliver TG, et al. A Switch in p53 dynamics marks cells that escape from DSB-induced cell cycle arrest. Cell Rep. 2020;33:107995.
Engeland K. Cell cycle regulation: p53-p21-RB signaling. Cell Death \ Differ. 2022;29:946–60.
Lane DP. p53, guardian of the genome. Nature 1992;358:15–6.
Haupt Y, Maya R, Kazaz A, Oren M. Mdm2 promotes the rapid degradation of p53. Nature 1997;387:296–9.
Kim DH, Rho K, Kim S. A theoretical model for p53 dynamics: identifying optimal therapeutic strategy for its activation and stabilization. Cell Cycle. 2009;8:3707–16.
Bar-Or RL, Maya R, Segel LA, Alon U, Levine AJ, Oren M. Generation of oscillations by the p53-Mdm2 feedback loop: A theoretical and experimental study. Proc Natl Acad Sci USA. 2000;97:11250–5.
Wagner J, Ma L, Rice JJ, Hu W, Levine AJ, Stolovitzky GA. p53-Mdm2 loop controlled by a balance of its feedback strength and effective dampening using ATM and delayed feedback. IEE Proc Syst Biol. 2005;152:109–17.
Zhang T, Brazhnik P, Tyson JJ. Exploring mechanisms of the DNA-damage response: p53 pulses and their possible relevance to apoptosis. Cell Cycle. 2007;6:85–94.
Geva-Zatorsky N, Rosenfeld N, Itzkovitz S, Milo R, Sigal A, Dekel E, et al. Oscillations and variability in the p53 system. Mol Syst Biol. 2006;2:33–2006.
Batchelor E, Mock CS, Bhan I, Loewer A, Lahav G. Recurrent Initiation: A Mechanism for Triggering p53 Pulses in Response to DNA Damage. Mol Cell. 2008;30:277–89.
Stewart-Ornstein J, Lahav G p53 dynamics in response to DNA damage vary across cell lines and are shaped by efficiency of DNA repair and activity of the kinase ATM. Sci Signal. 2017;10:1–10.
Christophorou MA, Ringshausen I, Finch AJ, Swigart LB, Evan GI. The pathological response to DNA damage does not contribute to p53-mediated tumour suppression. Nature 2006;443:214–7.
Efeyan A, Serrano M. p53: guardian of the genome and policeman of the oncogenes. Cell Cycle. 2007;6:1006–10.
Xue W, Zender L, Miething C, Dickins RA, Hernando E, Krizhanovsky V, et al. Senescence and tumour clearance is triggered by p53 restoration in murine liver carcinomas. Nature 2007;445:656–60.
Collado M, Serrano M. Senescence in tumours: Evidence from mice and humans. Nat Rev Cancer. 2010;10:51–7.
Nardella C, Clohessy JG, Alimonti A, Pandolfi PP. Pro-senescence therapy for cancer treatment. Nat Rev Cancer. 2011;11:503–11.
Beauséjour CM, Krtolica A, Galimi F, Narita M, Lowe SW, Yaswen P, et al. Reversal of human cellular senescence: Roles of the p53 and p16 pathways. EMBO J. 2003;22:4212–22.
Dirac AMG, Bernards R. Reversal of Senescence in Mouse Fibroblasts through Lentiviral Suppression of p53* 210. J Biol Chem. 2003;278:11731–4.
Wade Harper J, Adami GR, Wei N, Keyomarsi K, Elledge SJ. The p21 Cdk-interacting protein Cip1 is a potent inhibitor of G1 cyclin-dependent kinases. Cell 1993;75:805–16.
Xiong Y, Hannon GJ, Zhang H, Casso D, Kobayashi R, Beach D. p21 is a universal inhibitor of cyclin kinases. Nature 1993;366:701–4.
Brugarolas J, Chandrasekaran C, Gordon JI, Beach D, Jacks T, Hannon GJ. Radiation-induced cell cycle arrest compromised by p21 deficiency. Nature 1995;377:552–7.
Bornstein G, Bloom J, Sitry-Shevah D, Nakayama K, Pagano M, Hershko A. Role of the SCFSkp2 ubiquitin ligase in the degradation of p21Cip1 in S phase. J Biol Chem. 2003;278:25752–7.
Overton KW, Spencer SL, Noderer WL, Meyer T, Wang CL. Basal p21 controls population heterogeneity in cycling and quiescent cell cycle states. Proc Natl Acad Sci. 2014;111:E4386–E4393.
Barr AR, Cooper S, Heldt FS, Butera F, Stoy H, Mansfeld J, et al. DNA damage during S-phase mediates the proliferation-quiescence decision in the subsequent G1 via p21 expression. Nat Commun. 2017;8:1–17.
Spencer SL, Cappell SD, Tsai FC, Overton KW, Wang CL, Meyer T. The proliferation-quiescence decision is controlled by a bifurcation in CDK2 activity at mitotic exit. Cell 2013;155:369.
Moosmüller C, Tralie C, Kooshkbaghi M, Belkhatir Z, Pouryahya M, Reyes J, et al. Periodicity scoring of time series encodes dynamical behavior of the tumor suppressor p53. IFAC-PapersOnLine. 2021;54:488–95. https://www.sciencedirect.com/science/article/pii/S2405896321005814
Sakoe H, Chiba S. Dynamic programming algorithm optimization for spoken word recognition. IEEE Trans Acoust. 1978;26:43–9.
Berndt DJ, Clifford J Using dynamic time warping to find patterns in time series. In: KDD Workshop. 1994. p. 359–70.
Chiarella C, He XZ, Hommes C. A dynamic analysis of moving average rules. J Econ Dyn Control. 2006;30:1729–53.
Sakaue-Sawano A, Kurokawa H, Morimura T, Hanyu A, Hama H, Osawa H, et al. Visualizing spatiotemporal dynamics of multicellular cell-cycle progression. Cell 2008;132:487–98.
Batchelor E, Loewer A, Lahav G. The ups and downs of p53: understanding protein dynamics in single cells. Nat Rev Cancer. 2009;9:371–7.
Van Der Maaten L, Hinton G. Visualizing data using t-SNE. J Mach Learn Res. 2008;9:2579–625.
Coleman KE, Grant GD, Haggerty RA, Brantley K, Shibata E, Workman BD, et al. Sequential replication-coupled destruction at G1/S ensures genome stability. Genes Dev. 2015;29:1734–46.
Krenning L, Feringa FM, Shaltiel IA, vandenBerg J, Medema RH. Transient activation of p53 in G2 phase is sufficient to induce senescence. Mol Cell. 2014;55:59–72.
Ryl T, Kuchen EE, Bell E, Shao C, Flórez AF, Mönke G, et al. Cell-cycle position of single MYC-driven cancer cells dictates their susceptibility to a chemotherapeutic drug. Cell Syst. 2017;5:237–50.
Granada AE, Jiménez A, Stewart-Ornstein J, Blüthgen N, Reber S, Jambhekar A, et al. The effects of proliferation status and cell cycle phase on the responses of single cells to chemotherapy. Mol Biol Cell. 2020;31:845–57.
El-Deiry WS, Harper JW, O’Connor PM, Velculescu VE, Canman CE, Jackman J, et al. WAF1/CIP1 is induced in p53-mediated G1 arrest and apoptosis. Cancer Res. 1994;54:1169–74.
Brown JP, Wei W, Sedivy JM. Bypass of senescence after disruption of p21 CIP1/WAF1 gene in normal diploid human fibroblasts. Science 1997;277:831–4.
Fang L, Igarashi M, Leung J, Sugrue MM, Lee SW, Aaronson SA. p21Waf1/Cip1/Sdi1 induces permanent growth arrest with markers of replicative senescence in human tumor cells lacking functional p53. Oncogene 1999;18:2789–97.
Arnoff TE, El-Deiry WS. CDKN1A/p21WAF1, RB1, ARID1A, FLG, and HRNR mutation patterns provide insights into urinary tract environmental exposure carcinogenesis and potential treatment strategies. Am J Cancer Res. 2021;11:5452.
Shiohara M, El-Deiry WS, Wada M, Nakamaki T, Takeuchi S, Yang R, et al. Absence of WAF1 mutations in a variety of human malignancies. Blood 1994;84:3781–4.
McKenzie KE, Siva A, Maier S, Runnebaum IB, Seshadri R, Sukumar S. Altered WAF1 genes do not play a role in abnormal cell cycle regulation in breast cancers lacking p53 mutations. Clin Cancer Res J Am Assoc Cancer Res. 1997;3:1669–73.
Deng C, Zhang P, Harper JW, Elledge SJ, Leder P. Mice lacking p21CIP1/WAF1 undergo normal development, but are defective in G1 checkpoint control. Cell 1995;82:675–84.
Kastenhuber ER, Lowe SW. Putting p53 in context. Cell 2017;170:1062–78.
Li R, Waga S, Hannon GJ, Beach D, Stillman B. Differential effects by the p21 CDK inhibitor on PCNA-dependent DNA replication and repair. Nature 1994;371:534–7.
Cazzalini O, Scovassi AI, Savio M, Stivala LA, Prosperi E. Multiple roles of the cell cycle inhibitor p21CDKN1A in the DNA damage response. Mutat Res Mutat Res. 2010;704:12–20.
Ruan S, Okcu MF, Ren JP, Chiao P, Andreeff M, Levin V, et al. Overexpressed WAF1/Cip1 renders glioblastoma cells resistant to chemotherapy agents 1, 3-bis (2-chloroethyl)-1-nitrosourea and cisplatin. Cancer Res. 1998;58:1538–43.
Hsu CH, Altschuler SJ, Wu LF. Patterns of early p21 dynamics determine proliferation-senescence cell fate after chemotherapy. Cell 2019;178:361–73.
Gillies TE, Pargett M, Minguet M, Davies AE, Albeck JG. Linear integration of ERK activity predominates over persistence detection in Fra-1 regulation. Cell Syst. 2017;5:549–63.
Abraham U, Schlichting JK, Kramer A, Herzel H. Quantitative analysis of circadian single cell oscillations in response to temperature. PLoS One. 2018;13:e0190004.
Gabriel CH, Del Olmo M, Zehtabian A, Jäger M, Reischl S, van Dijk H, et al. Live-cell imaging of circadian clock protein dynamics in CRISPR-generated knock-in cells. Nat Commun. 2021;12:1–15.
Sanaki-Matsumiya M, Matsuda M, Gritti N, Nakaki F, Sharpe J, Trivedi V, et al. Periodic formation of epithelial somites from human pluripotent stem cells. Nat Commun. 2022;13:1–14.
Acknowledgements
The authors thank Professor Galit Lahav from Havard Medical School for making the data available to use in this study as well as the support of Memorial Sloan Kettering Cancer Center that made this work possible.
Funding
This study was supported by AFOSR grant (FA9550-17-1-0435), a grant from National Institutes of Health (R01-AG048769), MSK Cancer Center Support Grant/Core Grant (P30 CA008748), and a grant from Breast Cancer Research Foundation (BCRF-17-193). C.M. is supported by NSF DMS grant 2111322. J.R. received support from CONACyT/Fundacion Mexico en Harvard (404476), and Harvard Graduate Merit Fellowship.
Author information
Authors and Affiliations
Contributions
All authors participated in the study concept and design. APT, CT, JR, CM, ZB, JD, and AT performed development of the methodologies and analysis of the results. JR and AL performed biological interpretation of the obtained results. APT, CT, JR, CM, ZB, and AT performed initial writing and review of the paper. APT, JR, and AT performed the revision of the paper. All authors read and approved the final paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Edited by G. Melino
Supplementary information
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Tran, A.P., Tralie, C.J., Reyes, J. et al. Long-term p21 and p53 dynamics regulate the frequency of mitosis events and cell cycle arrest following radiation damage. Cell Death Differ 30, 660–672 (2023). https://doi.org/10.1038/s41418-022-01069-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41418-022-01069-x