Abstract
Background
Oesophageal squamous cell carcinoma (ESCC) is one of the most malignant cancers worldwide. Treatment of ESCC is in progress through accurate staging and risk assessment of patients. The emergence of potential molecular markers inspired us to construct novel staging systems with better accuracy by incorporating molecular markers.
Methods
We measured H scores of 23 protein markers and analysed eight clinical factors of 77 ESCC patients in a training set, from which we identified an optimal MASAN (MYC, ANO1, SLC52A3, Age and N-stage) signature. We constructed MASAN models using Cox PH models, and created MASAN-staging systems based on k-means clustering and minimum-distance classifier. MASAN was validated in a test set (n = 77) and an independent validation set (n = 150).
Results
MASAN possessed high predictive accuracies and stratified ESCC patients into three prognostic groups that were more accurate than the current pTNM-staging system for both overall survival and disease-free survival. To facilitate clinical utilisation, we also constructed MASAN-SI staging systems based on staining indices (SI) of protein markers, which possessed similar prognostic performance as MASAN.
Conclusion
MASAN provides a good alternative staging system for ESCC prognosis with a high precision using a simple model.
Similar content being viewed by others
Introduction
Oesophageal squamous cell carcinoma (ESCC) is the fourth leading cause of cancer-related mortality, and approximately half of the world’s 500,000 new ESCC cases occur annually in China.1, 2 The survival for ESCC is poor, with a 5-year overall survival (OS) of 20.9%.3 Treatment of ESCC remains a challenging problem. However, treatment outcomes are being improved through accurate staging and risk assessment of patients.4, 5 Accurate staging techniques, including molecular staging, allow us to understand prognosis and to tailor therapy to individuals to achieve the best outcomes.
Currently, the most commonly used staging systems for ESCC is the pTNM (pathological tumour-node metastasis) staging system (the 7th edition) proposed by the American Joint Committee on Cancer (AJCC).6 The AJCC pTNM system has become a standardised staging system for evaluating cancer at a population level. However, the development of molecular biology and discovery of molecular factors that predict cancer outcome and response to treatment with better accuracy has led cancer experts to question the utility of the pTNM-staging system at the individual level.7 Molecular factors, such as protein markers, are attracting more and more attention and have been demonstrated to benefit the diagnosis and prognosis of ESCC. Incorporating molecular factors into predictive models may further improve the accuracy of the staging system.
Over the past few decades, hundreds of dysregulated proteins have been detected in ESCC patients.8 Many of them were identified to be independent prognostic factors, such as MYC,9 ANO110 and ATF3.11 On the other hand, some clinical characteristics, such as N-stage, have always been predominant prognostic factors for ESCC.12, 13 Thus, Tan et al. proposed to combine protein markers and clinical characteristics, and built a FENSAM-staging system, which possessed high-classification precision similar to the pTNM-staging system, but was much simpler for clinical use.14 However, the protein markers used to build FENSAM were still limited. The predictive power of combinations of additional newly found protein markers needs further investigation. In addition, with more and more variables available for building predictive models, the anticipated predictive performance may not increase linearly with the number of variables due to complex interactions among variables.15 How to select an optimal feature combination and build robust predictive models remains a challenging problem.
To address this problem, we examine the expression of 23 potential protein markers and eight clinical characteristics of 304 ESCC patients, and propose a novel pipeline to identify optimal feature combination for model construction. We show that the resulting MASAN-staging system yields better prognostic capability than that of the pTNM-staging system, and provides a good alternative for clinical utilisation.
Materials and methods
Patients and specimens
Two independent data sets of formalin-fixed, paraffin-embedded tissue specimens were obtained from ESCC patients undergoing curative resection at the Shantou Central Hospital. The first data set included 154 patients treated during November 2007 to January 2010, and was randomly divided into a training set (n = 77) and a test set (n = 77). The clinicopathological characteristics were comparable in these two sets (Table 1). The training set was used to construct the predictive model and test set to evaluate the predictive performance. A second independent data set included 150 patients treated during 2000–2006 (validation set). All specimens were confirmed as ESCC by pathologists in the Clinical Pathology Department of the hospital, and the cases were classified according to the seventh edition of the AJCC pTNM system6 based on surgical T-stage, N-stage and M-stage. The surgical histologic grade of tumour differentiation was based on histological criteria of the guidelines of the WHO Classification of Tumours.16 Ethical approval was obtained from the ethical committee of the Central Hospital of Shantou City and the ethical committee of the Medical College of Shantou University. Only resected samples from surgical patients with written informed consent were included.
Tissue microarrays and immunohistochemistry
Tissue microarray (TMA) construction and immunohistochemistry (IHC) staining were based on standard techniques as previously described17 (see Supplementary methods). Twenty-three markers were measured in this study (Fig. 1a and Figure S1). The detailed information on primary antibodies is listed in Table S1.
Representative images of IHC staining and scoring process. (a) Expression of ANO1, MYC, and SLC52A3 in TMAs. -: represents cases with negative or weak staining;+: represents cases with moderate staining;++: represents cases with intense staining base on manual assessment. H score represents the protein expression value of the corresponding case, evaluated by an automated quantitative pathology imaging system. (scale bars = 50 μm) (b) Scoring process: tissue, cell segmentation and spectral analysis by inform software. I, V, IX, Colour image of sample. II, VI X, Region training analysis of sample superimposed on the colour image. Red: tumour region; green: other. III, VII, XI, Composite image of cell segmentation of the tumour region, nucleus shown in green and the cytoplasm for each cell are outlined in colour around the nucleus. IV, VIII, XII: Spectral analysis based on the optical density grouping into 4 tiers: blue: 0, yellow:+, orange:++, and brown:+++. I-IV, V-VIII and IX-XII are from the same cores of the TMAs
Evaluation of IHC variables
We scored protein expression using two methods: a newly emerged technology for extracting the H score automatically18 and the traditional manual assessment-staining index (SI; see Supplementary methods).
Statistical analysis
The univariate and multivariate Cox proportional hazards (Cox PH) models were built using the R package 'survival'. The predictive performance of Cox PH models was assessed using the concordance index (C-index)19 and area under the time-dependent ROC curve (AUC),20 which were calculated using the R package 'survcomp'. The k-means clustering algorithm was used to build the MASAN-staging system. The risk scores (RS) of patients in the training set were clustered into three clusters, which corresponded to the three MASAN stages. The thresholds of the MASAN stage were determined by a minimum-distance classifier. The genetic algorithm used to select optimal feature combination was performed using the R package 'mlr'.
Results
Identification of a MASAN signature
To construct a precise survival prediction model, we collected nine clinical characteristics (Table S2) and measured the expression of 23 proteins of 304 ESCC patients from two independent cohorts (see Materials and methods). IHC analysis showed that the immunostaining patterns of the 23 biomarkers were varied (Fig. 1a and Figure S1).
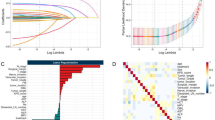
We designed a novel pipeline to identify optimal combinations of features (Fig. 2a). Initially, we used the genetic algorithm to select features from all 31 candidate features (23 proteins and 8 clinical variables) except pTNM stage. Eight features (fascin, MYC, ANO1, SLC52A3, age, smoking, G- and N-stage) with a C-index of 0.67 were identified after 100 iterations (Fig. 2b). Furthermore, an exhaustive search was performed to evaluate the predictive performance of all combinations of the eight features (Supplementary Methods). Feature combinations with both a high average C-index and a large number of times of significant stratification (located at the top right corner in Fig. 2c) were favourable signatures for survival prediction. Finally, five features (MYC, ANO1, SLC52A3, age and N-stage, MASAN) with an average C-index of 0.6514 and 993 significant stratifications were identified as the optimal feature combination (Fig. 2c).
Construction of the MASAN model. a Pipeline of feature selection. b Procedure for optimisation of the genetic algorithm. Eight features (dotted yellow line) with a C-index (blue line) of 0.67 were identified at 100 iterations. c Comparison of predictive performance of all combinations of eight features. The combinations with times of significant stratification >900 were displayed. Five features (MASAN) with an average C-index of 0.6514 and 993 significant stratifications were identified
MASAN predicts the OS of ESCC patients
We constructed a Cox PH model using MASAN as independent variables and the OS information as dependent variables (referred to as MASAN model) from the training set (Table S3). The RS for OS (RSos) of a new patient i (\(RS_{OS}^i\)) can be calculated by formula (1):
where\(E_{MYC}^i\),\(E_{ANO1}^i\) and\(E_{SLC52A3}^i\)denote the H scores of MYC, ANO1 and SLC52A3, respectively. \(E_{Age}^i\) and \(E_{N - stage}^i\)denote the age and N-stage of patient i, respectively.
To investigate the predictive ability of the MASAN model, we applied MASAN to predict RSoss of patients in the training set, test set and validation set, respectively. The RSoss yielded significant stratifications of patients, in all the three data sets, into low- and high-risk groups (P = 6.78 × 10−4, 1.07 × 10−3 and 7.57 × 10−5, respectively, Figure S2) using the median RSos in the training set as the cutoff point, indicating that the predicted RSoss were quite consistent with the actual OS.
To compare the predictive ability of the MASAN model with the pTNM-staging system, we constructed a MASAN-staging system by clustering the patients in the training set into three groups using k-means clustering on the RSoss (Table S4). Kaplan–Meier analysis showed that the survival probabilities were significantly different among three stages (OS median = 1979, 1005.5 and 427 days for MASAN stages I–III, respectively, P = 0.0001, Fig. 4a). In contrast, the pTNM-staging system classified only three patients into stage I, and had a larger P value (P = 0.0329, Fig. 3d). The median AUC was larger for the MASAN than the pTNM system (0.7130 vs. 0.6432). In fact, the time-dependent AUCs for the MASAN-staging system were larger than those for the pTNM-staging system at each time point (Fig. 4a). Figure 4d shows the ROC curves for the two systems at the 3-year time point, where the superiority of the MASAN-staging system can be clearly observed.
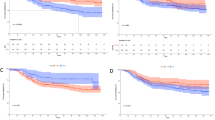
Comparison of the MASAN- and pTNM-staging systems on the OS and DFS of patients with ESCC by Kaplan–Meier analysis. a–c Kaplan–Meier curves using the MASAN system on OS for the training set (a), test set (b) and validation set (c). d–f Kaplan–Meier curves using the pTNM-staging system on OS for the training set (d), test set (e) and validation set (f). g–i Kaplan–Meier curves using the MASAN system on DFS for the training set (g), test set (h) and validation set (i). j–l Kaplan–Meier curves using the pTNM-staging system on DFS for the training set (j), test set (k) and validation set (l). P values were calculated by log-rank test
Predictive performance of the MASAN- and pTNM-staging systems on OS of patients with ESCC. a–c Time-dependent AUCs using the MASAN- and pTNM-staging systems on OS for the training set (a), test set (b) and validation set (c). d–f ROC curves of the MASAN- and pTNM-staging systems at the 3-year time point on OS for the training set (d), test set (e) and validation set (f). g Boxplots of AUCs using the MASAN- and pTNM-staging systems on the test set and validation set. ***P < 2.2 × 10−16. h Boxplots of –log (P values) using the MASAN and pTNM-staging systems on the test set and validation set. P values were calculated by the Wilcoxon-signed rank test
Furthermore, the MASAN-staging system stratified the patients into three groups with significant OS differences for both the test set (P = 0.0007, Fig. 4b) and validation set (P = 1.5 × 10−6, Fig. 4c). In contrast, the stratifications of the pTNM-staging system had less significant OS differences (P = 0.0202 and 5.13 × 10−5, respectively, Fig. 3e,f). Specifically, the pTNM-staging system classified only a few patients into stage I for both the test set (n = 5) and the validation set (n = 2). The median AUC was larger for the MASAN than the pTNM-staging system (0.7332 vs. 0.6507 for the test set, and 0.6718 vs. 0.6555 for the validation set). Time-dependent AUC curves also showed that the MASAN-staging system yielded better predictive performance than that of the pTNM-staging system (Fig. 4b, c, e and f). Moreover, multivariable analysis showed that the MASAN signature was an independent prognostic factor for OS of ESCC patients in all three data sets (P = 0.0024, 0.0120 and 0.0022, respectively; Table S5).
In addition, to ensure that the predictive performance was not dependent on the particular patient set in the test set and validation set, we randomly chose 80% of patients from the two sets as the new test set (n = 61) and validation set (n = 120). Then we compared the predictive performance of the two systems on these two new sets by median AUC and P value of the log-rank test. We repeated the procedure 500 times. Boxplots showed that both the median AUCs and –log (P values) were significantly larger for the MASAN-staging system than the pTNM-staging system on the two new sets (Wilcoxon-signed rank test, P < 2.2 × 10−16 for all four comparisons, Fig. 4g, h). Besides, we also evaluated MASAN models on patients treated with surgery alone, and obtained similar prognostic performance (Figs. S3A and 3B). This further indicates that the MASAN-staging system is robust and produces consistently better ESCC prognosis.
MASAN predicts DFS of ESCC patients
Next, we constructed a MASAN-staging system for DFS using the MASAN signature as independent variables, and the DFS information as dependent variables from the training set (Table S3). The RS for DFS (RSDFS) of a new patient i (\(RS_{DFS}^i\)) can be calculated by formula (2):
The predicted RSDFSs yielded significant stratifications of patients into low- and high-risk groups for the three data sets (P = 0.0011, 0.0037 and 6.18 × 10−5, respectively, Figure S4), indicating that the predicted RSDFSs were consistent with the actual DFS.
Next, we constructed the MASAN-staging system for DFS (Table S4). The MASAN-staging system again stratified the patients in three data sets into three stages with significant DFS differences (P = 1.1 × 10−3, 1.19 × 10−6 and 1.68 × 10−6, respectively, Fig. 3g-i). In contrast, the stratification with the pTNM-staging system was not significant for the training set (P = 0.0715, Fig. 3j) and less significant for the test set (P = 0.0026, Fig. 3k).
The median AUC was larger for the MASAN than the pTNM system for the three data sets (0.6972 vs. 0.6207, 0.7423 vs. 0.6827, and 0.6730 vs. 0.6542, respectively). Time-dependent AUC curves also showed that the MASAN system yielded better predictive performance than that of pTNM system (Figs. 5a-c and d–f). As OS, multivariable analysis of DFS showed that the MASAN signature was an in independent prognostic factor in all three data sets (P = 0.0093, 0.0002 and 0.0154, respectively; Table S5). And also, the MASAN-staging system had similar prognostic performance on patients treated with surgery alone (Figs. S3C and 3D). In addition, the permutation test also showed that the 500 AUCs and 500 –log (P values) were significantly larger for the MASAN-staging system than pTNM-staging system, respectively (Wilcoxon-signed rank test, P < 2.2 × 10−16 for all four comparisons, Fig. 5g, h).
Predictive performance of the MASAN- and pTNM-staging systems on DFS of patients with ESCC. a–c The time-dependent AUCs of the MASAN- and pTNM-staging systems on DFS for the training set (a), test set (b) and validation set (c). d–f ROC curves of the MASAN- and pTNM-staging systems at the 3-year time point on DFS for the training set (d), test set (e) and validation set (f). g Boxplots of AUCs using the MASAN- and pTNM-staging systems on the test set and validation set. ***P < 2.2 × 10−16. h Boxplots of –log (P values) of the MASAN- and pTNM-staging systems on the test set and validation set. P values were calculated by the Wilcoxon-signed rank test
MASAN-SI predicts survival outcome of ESCC patients
For the convenience of clinical utilisation, we also constructed MASAN models using the SI of protein markers (MASAN-SI; Table S6). The RS for OS (RS-SIOS) and DFS (RS-SIDFS) of a new patient i can be calculated by formulae (3) and (4), respectively:
where \(ST_{MYC}^i\),\(ST_{ANO1}^i\) and\(ST_{SLC52A3}^i\)denote the SI of MYC, ANO1 and SLC52A3, respectively.
We constructed a MASAN-SI staging system using the thresholds listed in Table S7. Similar to the MASAN-staging system, MASAN-SI stratified ESCC patients into the three data sets into three stages with significant OS differences (P = 3.0 × 10−4, 6.0 × 10−4 and 2.0 × 10−4, respectively, Figure S5A-C) and DFS differences (P = 5.5 × 10−3, 2.05 × 10−5 and 9.55 × 10−5, respectively, Figure S5G-H). The time-dependent AUCs were larger for MASAN-SI- than the pTNM-staging system in the training set (OS: Figure S5D; DFS: Figure S5J) and test set (OS: Figure S5E; DFS: Figure S5K). In the validation set, the predictive performance of the two systems was comparable, with MASAN-SI slightly better on prognosis within 3 years (Figure S5F and S5L).
Discussion
In this study, we examined the expressions of 23 potential protein markers and eight clinical characteristics of ESCC patients, from which we identified an optimal feature combination (MASAN) for precise prediction of ESCC survival outcome. We built MASAN models for both OS and DFS. The prognostic value of the MASAN models was verified in a test set and an independent validation set. Results showed that the MASAN-staging system yielded better prognostic performance than that of the pTNM-staging system.
The MASAN signature comprises both clinical factors and molecular factors. The clinical factors are essential as molecular factors alone could not accurately predict survival of ESCC patients (Figure S6A-C). In the MASAN model, coefficients are larger for N-stage than other features (formula (1)–(4)). Without N-stage, the prognostic performance was seriously deteriorated (Figure S6D-F). So N-stage is still a predominant prognostic factor, consistent with several previous studies.12,13,14 Positive expression of MYC and ANO1 has been found to be significantly correlated with poorer prognosis and suggested as potential biomarkers for ESCC patients.9, 10 In our three data sets, the expression values of ANO1 were high (>50) in only a small proportion of patients (6/77, 14/77 and 14/150, respectively). However, removing ANO1 from the MASAN model resulted in declined predictive performance, especially for DFS prediction in the validation set (Figure S6G), indicating that ANO1 plays a necessary role in the MASAN model. SLC52A3 has been suggested as a potential therapeutic target.21 Knockdown of SLC52A3 in ESCC cells results in inhibition of cell proliferation, whereas overexpression of SLC52A3 in ESCC cells promotes cell proliferation and tumourigenesis in nude mice.21 Age is also an essential factor in the MASAN model as removing age resulted in declined predictive performance (Figure S6H and 6I).
Beyond the superior predictive performance, the stratification of ESCC patients is more reasonable for MASAN-staging system than the pTNM-staging system. The MASAN-staging system stratifies more patients into the low-risk group compared to pTNM-staging system (Fig. 3). Furthermore, stratification by the MASAN-staging system possesses more consistent and higher OS for low-risk patients, and lower OS for high-risk patients, while pTNM fluctuated more widely (Table S8). DFS also had the same tendency (Table S9). Thus, the MASAN-staging system provides better guidance for making clinical decisions. More low-risk patients may avoid unnecessary treatments. Moreover, the MASAN model is based on protein markers and clinical characteristics, and is easy to use. On the basis of a simple model, MASAN provides a good alternative staging system for ESCC patients with a high precision.
Note that, although MASAN is reliable for Chinese patients, it must be careful to use it for prognosis of Caucasian patients as there exists differences between Asian and Caucasian patient populations in both clinicopathologic and molecular features.22, 23 The feasibility of MASAN or new staging models on Caucasian patients will be investigated when we have enough samples in future. Another limitation is that, as a retrospective study, the patients used in this study were mostly collected between 2000 and 2010, which lacked necessary pre-operative information for accurate clinical staging system. Thus, MASAN cannot be used as a clinical staging system. As clinical staging system is of great value for patient care, pre-operative information of ESCC patients should be included to construct novel clinical staging system with better accuracy in future.
To facilitate clinical utilisation, we constructed prognostic models using both H score (MASAN) and SI (MASAN-SI). Results show that MASAN-SI obtains similar prognostic performance as MASAN. Both models are available at http://www.licpathway.net/MASAN/index.php.
References
Torre, L. A. et al. Global cancer statistics, 2012. CA Cancer J. Clin. 65, 87–108 (2015).
Chen, W. et al. Cancer statistics in China, 2015. CA Cancer J. Clin. 66, 115–132 (2016).
Zeng, H. et al. Cancer survival in China, 2003-2005: a population-based study. Int. J. Cancer 136, 1921–1930 (2015).
Pennathur, A. & Luketich, J. D. Resection for esophageal cancer: strategies for optimal management. Ann. Thorac. Surg. 85, S751–S756 (2008).
Pennathur, A., Gibson, M. K., Jobe, B. A. & Luketich, J. D. Oesophageal carcinoma. Lancet 381, 400–412 (2013).
Rice, T. W., Blackstone, E. H. & Rusch, V. W. 7th edition of the AJCC Cancer Staging Manual: esophagus and esophagogastric junction. Ann. Surg. Oncol. 17, 1721–1724 (2010).
Amin, M. B. et al. The Eighth Edition AJCC Cancer Staging Manual: continuing to build a bridge from a population-based to a more “personalized” approach to cancer staging. CA Cancer J. Clin. 67, 93–99 (2017).
Lin, D. C., Du, X. L. & Wang, M. R. Protein alterations in ESCC and clinical implications: a review. Dis. Esophagus 22, 9–20 (2009).
Wang, W., Xue, L. & Wang, P. Prognostic value of beta-catenin, c-myc, and cyclin D1 expressions in patients with esophageal squamous cell carcinoma. Med. Oncol. 28, 163–169 (2011).
Shang, L. et al. ANO1 protein as a potential biomarker for esophageal cancer prognosis and precancerous lesion development prediction. Oncotarget 7, 24374–24382 (2016).
Xie, J. J. et al. ATF3 functions as a novel tumor suppressor with prognostic significance in esophageal squamous cell carcinoma. Oncotarget 5, 8569–8582 (2014).
Ikeda, G., Isaji, S., Chandra, B., Watanabe, M. & Kawarada, Y. Prognostic significance of biologic factors in squamous cell carcinoma of the esophagus. Cancer 86, 1396–1405 (1999).
Kuo, K. T. et al. Clinicopathologic significance of cyclooxygenase-2 overexpression in esophageal squamous cell carcinoma. Ann. Thorac. Surg. 76, 909–914 (2003).
Tan, H. et al. A novel staging model to classify esophageal squamous cell carcinoma patients in China. Br. J. Cancer 110, 2109–2115 (2014).
Ishwaran, H., Blackstone, E. H., Apperson-Hansen, C. & Rice, T. W. A novel approach to cancer staging: application to esophageal cancer. Biostatistics 10, 603–620 (2009).
Flejou, J. F. [WHO Classification of digestive tumors: the fourth edition]. Ann. Pathol. 31, S27–S31 (2011).
Xie, J. J. et al. Prognostic implication of ezrin expression in esophageal squamous cell carcinoma. J. Surg. Oncol. 104, 538–543 (2011).
Huang, W., Hennrick, K. & Drew, S. A colorful future of quantitative pathology: validation of Vectra technology using chromogenic multiplexed immunohistochemistry and prostate tissue microarrays. Hum. Pathol. 44, 29–38 (2013).
Harrell, F. E. Jr, Lee, K. L. & Mark, D. B. Multivariable prognostic models: issues in developing models, evaluating assumptions and adequacy, and measuring and reducing errors. Stat. Med. 15, 361–387 (1996).
Heagerty, P. J., Lumley, T. & Pepe, M. S. Time-dependent ROC curves for censored survival data and a diagnostic marker. Biometrics 56, 337–344 (2000).
Jiang, X. R. et al. RFT2 is overexpressed in esophageal squamous cell carcinoma and promotes tumorigenesis by sustaining cell proliferation and protecting against cell death. Cancer Lett. 353, 78–86 (2014).
Deng, J. et al. Comparative genomic analysis of esophageal squamous cell carcinoma between Asian and Caucasian patient populations. Nat. Commun. 8, 1533 (2017).
Zhang, J. et al. Comparison of clinicopathologic features and survival between eastern and western population with esophageal squamous cell carcinoma. J. Thorac. Dis. 7, 1780–1786 (2015).
Acknowledgements
We thank all the research staff for their contributions to this project. This work was supported in part by the Natural Science Foundation of China-Guangdong Joint Fund (Grants Nos. U1301227 and U1601229), the National Science Foundation of China (Grant Nos. 81472613 and 61602292), National Cohort of Oesophageal Cancer of China (Grant No. 2016YFC09014000), the China Postdoctoral Science Foundation (Grant No. 2016M602499), the University Nursing Program for Young Scholars with Creative Talents in Heilongjiang Province (Grant No. UNPYSCT-2016102), the doctoral research fund of Heilongjiang Institute of Technology (Grant No. 2014BJ16) and the Department of Education, Guangdong Government under the Top-tier University Development Scheme for Research and Control of Infectious Diseases.
Author contributions:
L.Y.X. and E.M.L. conceived the concept for this study; J.Z.H. and W.L. discussed and performed the analyses; W.L., J.Z.H. and L.Q.C. wrote the manuscript; D.K.L., S.H.W. and J.Z.H. carried out the immunohistochemical analysis; X.F.B., Y.J. and C.Q.L. implemented the MASAN website; X.E.X. and J.Z.H. were responsible for immunohistochemistry; J.Z.H. and W.L. produced the data from tissue microarrays and supervised the pathology data analysis and interpretation; J.Y.W. was responsible for follow-up tracing.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interest
The authors declare no competing interest.
Ethical approval and consent to participate
Ethical approval was obtained from the ethical committee of the Central Hospital of Shantou City and the ethical committee of the Medical College of Shantou University.
Additional information
Note: This work is published under the standard license to publish agreement. After 12 months the work will become freely available and the license terms will switch to a Creative Commons Attribution 4.0 International licence (CC BY 4.0).
Electronic supplementary material
Rights and permissions
This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Liu, W., He, Jz., Wang, Sh. et al. MASAN: a novel staging system for prognosis of patients with oesophageal squamous cell carcinoma. Br J Cancer 118, 1476–1484 (2018). https://doi.org/10.1038/s41416-018-0094-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41416-018-0094-x
This article is cited by
-
Spatial analysis of stromal signatures identifies invasive front carcinoma-associated fibroblasts as suppressors of anti-tumor immune response in esophageal cancer
Journal of Experimental & Clinical Cancer Research (2023)
-
Using a machine learning approach to identify key prognostic molecules for esophageal squamous cell carcinoma
BMC Cancer (2021)
-
Integrated single-cell transcriptome analysis reveals heterogeneity of esophageal squamous cell carcinoma microenvironment
Nature Communications (2021)
-
Large-scale and high-resolution mass spectrometry-based proteomics profiling defines molecular subtypes of esophageal cancer for therapeutic targeting
Nature Communications (2021)
-
Integration of gene interaction information into a reweighted random survival forest approach for accurate survival prediction and survival biomarker discovery
Scientific Reports (2018)