Abstract
In many anoxic environments, syntrophic acetate oxidation (SAO) is a key pathway mediating the conversion of acetate into methane through obligate cross-feeding interactions between SAO bacteria (SAOB) and methanogenic archaea. The SAO pathway is particularly important in engineered environments such as anaerobic digestion (AD) systems operating at thermophilic temperatures and/or with high ammonia. Despite the widespread importance of SAOB to the stability of the AD process, little is known about their in situ physiologies due to typically low biomass yields and resistance to isolation. Here, we performed a long-term (300-day) continuous enrichment of a thermophilic (55 °C) SAO community from a municipal AD system using acetate as the sole carbon source. Over 80% of the enriched bioreactor metagenome belonged to a three-member consortium, including an acetate-oxidizing bacterium affiliated with DTU068 encoding for carbon dioxide, hydrogen, and formate production, along with two methanogenic archaea affiliated with Methanothermobacter_A. Stable isotope probing was coupled with metaproteogenomics to quantify carbon flux into each community member during acetate conversion and inform metabolic reconstruction and genome-scale modeling. This effort revealed that the two Methanothermobacter_A species differed in their preferred electron donors, with one possessing the ability to grow on formate and the other only consuming hydrogen. A thermodynamic analysis suggested that the presence of the formate-consuming methanogen broadened the environmental conditions where ATP production from SAO was favorable. Collectively, these results highlight how flexibility in electron partitioning during SAO likely governs community structure and fitness through thermodynamic-driven mutualism, shedding valuable insights into the metabolic underpinnings of this key functional group within methanogenic ecosystems.
Similar content being viewed by others
Introduction
Anaerobic digestion (AD) is a globally important biotechnology for resource recovery and biogas production from organic waste streams. As an open fermentation process, conversion of complex organic polymers into methane within AD systems depends on coordinated activities of multiple microbial trophic guilds, including hydrolyzers, fermenters, syntrophs, and methanogenic archaea [1]. In the terminal steps of the AD food-web, acetate accounts for around 70% of the total electron flow into methane [2], and its turnover rate can have significant impacts on the AD loading capacity and process stability [3].
Acetate can be converted into methane and carbon dioxide via acetoclastic methanogenesis by archaea belonging to the genera Methanothrix and Methanosarcina [4]. Alternatively, acetate can be oxidized by syntrophic acetate oxidizing (SAO) bacteria into carbon dioxide, hydrogen and/or formate, which are substrates for methanogenesis via carbon dioxide reduction by archaeal partners [5]. SAO is not thermodynamically feasible under standard conditions, and therefore it requires the presence of methanogenic archaea to maintain low hydrogen and/or formate levels [6]. Although acetoclastic and hydrogenotrophic methanogens coexist in many AD environments, oftentimes acetoclastic methanogens are rare or absent in thermophilic systems (>50 °C) and/or in systems with elevated free ammonia (NH3) concentrations [7]. In such AD systems, SAO is likely an important pathway for methane production [8,9,10].
Despite their importance in driving methane production within many anoxic environments, little is currently known about the in situ physiologies and activities of SAO bacteria (SAOB). To date, only six strains of SAOB are available in pure culture: the thermophilic Pseudothermotoga lettingae [11] and Thermacetogenium phaeum [12]; the thermotolerant Tepidanaerobacter acetatoxydans [13]; the mesophilic Clostridium ultunense [14] and Syntrophaceticus schinkii [15]; and the alkaliphilic Candidatus Contubernalis alkalaceticum [16]. All of these isolated SAOB were obtained by enrichment or co-cultivation with methanogens, except for Candidatus Contubernalis alkalaceticum that was enriched along with sulfate reducing bacteria. Among these characterized SAOB, three species utilize the reverse Wood–Ljungdahl (acetyl-coA) pathway for acetate oxidation (Thermacetogenium phaeum, Syntrophaceticus schinkii, and Tepidanaerobacter acetatoxydans) [17,18,19], indicating the potential for metabolic diversity within this functional guild. Correspondingly, culture-independent molecular approaches, such as small subunit ribosomal RNA (SSU, or 16S rRNA) gene amplicon sequencing and lineage or gene-specific PCR, have been used to infer the identity and dynamics of putative SAOB in AD systems [20,21,22]. Results have indicated that SAOB belong to phylogenetically diverse, and in most cases uncharacterized, groups [23]. While genome-resolved metagenomic efforts have recovered genomes for putative SAOB within AD systems [24, 25], the reversibility of the Wood–Ljungdahl pathway used by some SAOB [19] further obfuscates the discrimination of SAOB from homoacetogens. This knowledge gap currently hinders our ability to develop and validate appropriate ecosystem-level models for carbon flow within SAO-dominated ecosystems, which are important for informing engineering strategies to enhance AD process stability and carbon conversion efficiency.
Stable isotope probing (SIP) is a powerful molecular approach to link genomic identity with metabolic function by detecting isotope incorporation into biomolecules during growth [26]. Given its resolving power, SIP could be a useful approach to discern the in situ ecophysiology of uncultivated SAOB. While DNA/RNA-based SIP has been applied to identify potential SAOB in AD systems [27,28,29], these efforts have so-far relied on PCR based gene sequencing, limiting new insights into the metabolic underpinnings of SAO. In a recent study, Mosbæk et al. [30] combined SIP with metaproteomics and metagenomics to identify genomes of SAOB associated with [13C]-labeled peptides within full-scale AD systems. However, the low abundance of the putative SAOB, likely due to their low energy yields [6], limited the number of 13C-labeled peptides identified within the metaproteome [30]. We posit that such genome-resolved SIP efforts could benefit from AD biomass that is enriched in SAOB for improved recovery of molecular information.
In this study, we carried out long-term enrichments of syntrophic acetate-oxidizing consortia from a thermophilic wastewater treatment plant AD system, followed by genome-resolved metaproteogenomic SIP to track carbon flow into individual populations. Genome-resolved metagenomic annotations and measurements of carbon flux into proteins were used to inform a community-scale metabolic model, which was utilized to investigate the impact of different electron shuttles on the fitness and feasibility of SAO metabolism. The results of this study shed new insights into the role of interspecies electron transfer in SAO community metabolism and composition, while highlighting how SIP-based multi-omic approaches can be used to inform community-scale models of cryptic or uncharacterized metabolisms within complex microbiomes.
Materials and methods
Anaerobic enrichment chemostat set up and operation
Duplicate sterile 5 l glass continuous stirred-tank bioreactors (R1 and R2) were pre-flushed with 80:20 N2:CO2, and were inoculated with 800 ml of sludge collected from a thermophilic (55 °C) anaerobic digester at a nearby municipal wastewater treatment plant (Vancouver, Canada: 1.7 g NH4+-N/L, 185 mg NH3/L, pH 7.5; see Supplementary Methods for calculations). The inoculum was diluted with 2400 ml of sterile anoxic basal medium prepared as described by Plugge [31] (see Supplementary Methods for details), with the total ammonium-nitrogen (TAN) concentration adjusted to 1.0 g NH4+-N/l. The bioreactors were maintained at 55 °C by an electric heating jacket, and stirred at 100 rpm by a mechanical mixer (Applikon Biosciences). The bioreactors were fed (160 ml/day) with the anoxic and sterile bicarbonate-buffered basal medium described above, with sodium acetate (75 mM) amended as the predominant carbon source. A liquid volume of 3.2 l was maintained in the bioreactors, providing a solids retention time (SRT) and hydraulic retention time (HRT) of 20 days to mimic that of the full-scale digester used as inoculum. On day 126, the SRT/HRT was increased to 30 days. The bioreactors were operated for a total of 300 days.
The volume of biogas and methane concentration were recorded in real-time by a gas meter (BlueVCount; BlueSens GmbH, Germany) and optical infrared sensor (BCP-CH4 sensor; BlueSens GmbH, Germany), respectively. The gas meter contained a one-way check valve that prevented intrusion of air into the bioreactors. The pH and temperature were measured in real-time with an in situ pH probe (InPro3250 pH; Mettler Toledo, USA). Liquid samples were periodically collected from both bioreactors to determine TAN, volatile fatty acids (VFAs), chemical oxygen demand (COD), total solids (TS) and volatile solid (VS) (Supplementary Methods).
Batch microcosms for stable isotope probing
After 300 days of chemostat operation, batch microcosms were established in 40 ml glass serum bottles flushed with 80:20 N2:CO2 by anoxically transferring 18 ml of digestate from a single bioreactor (R2) and sealing with butyl rubber septa. Four different incubation conditions were established in triplicate: (1) blank control (e.g., no amendment); (2) 50 mM [12C]-acetate; (3) 50 mM [2-13C]-acetate (e.g., methyl-labeled); (4) 50 mM [1,2-13C]-acetate (universally labeled). Acetate was added to the microcosms (2 ml) as anoxic sterilized basal medium containing [12C], [2-13C], or [1,2-13C] sodium acetate (isotope purity >98%, Cambridge Isotopes). Bottles were held at 55 °C in a shaking incubator at 100 rpm. Twelve replicate bottles were established for all universally-labeled acetate-amended microcosms, allowing for three triplicate sets to be sacrificed for protein extraction at 24, 144, and 408 h, and a single triplicate set for liquid sampling throughout for VFA analysis. Biomass was pelleted from 10 ml liquid samples via centrifugation (10,000 × g) and stored at −20 °C until protein extraction. The supernatant was filtered with 0.2 µm Titan PTFE syringe filters (Thermo Scientific) and stored at −20 °C until metabolomics analysis on NMR (see Supplementary Methods). Gas production, gas composition, carbon isotope ratios of CO2 and CH4, and VFAs were measured approximately every 3 days (Supplementary Methods). Headspace gas samples (0.5 ml) were also collected into Exetainer vials (12 ml, Labco, UK) pre-purged with N2 gas for subsequent analysis on isotope ratio mass spectrometry (IRMS) (Supplementary Methods).
Metagenomic sequencing, assembly, and binning
Samples (10 ml) from both bioreactors on operational days 0, 19, 54, 81, 111, 234, and 283 were collected for short-read metagenomic sequencing (Supplementary Table S1). DNA was extracted from these samples using the FastDNA Spin Kit For Soil (MP Biomedicals, California) with minor modifications [32]. These DNA samples were fragmented and ligated with adapters using the Nextera XT DNA Library Preparation Kit (Illumina, USA) and sequenced on a NextSeq550 System (Illumina) in 2 × 150 bp paired-end mode, generating an average of 18 ± 5 Gbp per sample (Supplementary Table S1). Additionally, a sample collected on the day of the stable-isotope probing experiment (day 300) was extracted using the DNA PowerSoil Pro MagAttract kit (Qiagen) and prepared for Nanopore sequencing using the Q20+ Ligation Sequencing Kit (SQK-Q20EA) on a R10.4 MinION flow cell.
Adapter-removal and quality trimming was performed on short-read Illumina metagenomic samples using BBMap v38.36 [33]. For read-based taxonomic profiling, metagenome k-mer signatures were generated with sourmash (v.4.8.3) [34, 35] using the “sketch dna” command (k = 31), and the resulting signatures were classified against the GTDB (release 214) [36] using the sourmash “gather” command. The output was summarized at taxonomic ranks using the sourmash “tax” command and the GTDB reference sheet (https://osf.io/wxf9z/).
Each quality-filtered short-read metagenome was individually assembled into contigs using SPAdes v3.15.4 in “metagenomic” mode [37]. Reads from each sample were reciprocally mapped to each assembly using bowtie2 v2.4.4 [38]. Binning was performed on each individual assembly with metaBAT2 v2.14 [39] using differential coverage from the mapping of all samples, and bins were de-replicated using dRep v3.2.2 [40]. Bins were assessed for completeness and contamination based on the presence of conserved single copy-core genes with CheckM v1.1.3 [41].
Nanopore long-reads were basecalled using guppy v6.0.1, yielding 10.8 Gbp of passed reads. Adapters were then trimmed using porechop v0.3.2. Long-reads were then assembled using flye v2.9 with “--nano-hq” and “--meta” settings [42]. Contigs were then polished three times using Nanopore raw reads with Racon v1.4.3 [43], followed by three rounds of polishing with medaka v1.5. Additional rounds of polishing were performed with Racon v1.4.3 and polypolish v0.5.0 using Illumina short-reads from day 283. All short-read samples were then mapped to the long-read assembly to obtain differential coverage profiles using bowtie2 v2.4.4. Archaeal and bacterial single-copy core genes were identified on all contigs using Anvi’o v.7.0 [44]. The long-read contigs, differential coverage information, contig classifications, and single copy-core gene locations were imported into R for manual binning with the mmgenome2 v2.2.1 package [45].
A final set of metagenome-assembled genomes (MAGs) (Supplementary Data 1) was obtained by de-replicating all bins across the long-read and short-read assembly sets. For genomes that shared similarity above 95% ANI, the highest quality representative MAG was chosen based on completion and contamination statistics followed by genome contiguity. Taxonomic classifications of the final set of bins were assigned using the GTDB-tk v1.7.0 and release202 database [36, 46]. Relative abundance of the dereplicated set of bins in each sample was assessed by mapping metagenomic reads to the concatenated set of bins with Bowtie2 v2.4.4 and using the “relative_abundance” method in CoverM v0.6.1. Average nucleotide identity (ANI) values for genomes of interest were calculated using FastANI [47]. Proteins were predicted with Prodigal v2.6.3 [48] and functional annotations made with KofamKOALA v1.3.0 with KEGG release 103.0 [49] and MetaPathways v2.0 [50, 51]. To predict whether annotated hydrogenases and formate dehydrogenases were electron-bifurcating, we analyzed their beta subunits for the signature amino-acid motifs identified by Losey et al. [52] (Supplementary Methods). Annotated hydrogenases were also queried with HydDB [53] to determine their functions and group classifications.
Metaproteomics sample preparation and data acquisition
Protein from cell pellet samples (200 µl) were extracted by bead beating in 100 mM ammonium bicarbonate buffer, then reduced and alkylated (Supplementary Methods). Proteins were then digested with trypsin and subsequently desalted using C18 solid phase extraction (Supplementary Methods). MS analysis was performed using 0.1 µg/µl of peptide solution injected into a Q‐Exactive HF-X mass spectrometer (Thermo Scientific), with the detailed conditions outlined in the Supplementary Methods.
Metaproteomics data analysis
Mass spectrometry (MS) data for each biological replicate at all time points (n = 18) were analyzed using an implementation of OpenMS [54] implemented in KNIME [55] (see Supplementary Methods). Briefly, MS/MS spectra were searched using the MS-GF+ tool [56] against a protein database consisting of all ORFs from the de-replicated set of MAGs, concatenated with reversed (decoy) sequences of all protein entries. Peptide spectra matches (PSMs) were filtered at a 5% false discovery rate (FDR) with Percolator [57]. For label-free quantification (LFQ) of proteins, PSMs from unlabeled (12C) samples were used for protein inference with Fido [58], followed by protein FDR filtering at 5%. Protein quantification was based on the summed intensities of all unique PSMs within a protein group.
A “total protein approach” [59, 60] was implemented to infer absolute protein levels based on the LFQ data. The relative protein abundance was determined as:
where i refers to a single protein and K refers to the set of all proteins in a given sample.
The total protein concentration was then estimated by multiplying the relative protein abundance (g/g) by the total protein concentration in the sample (g/L):
Molar concentrations of proteins were determined for use in expression profiling as:
where MWi is the molecular weight of protein i (g/mole), inferred from the amino acid sequence.
To identify labeled peptides in the MS data, we used MetaProSIP [61] implemented through OpenMS (see Supplementary Methods). The output of this analysis yielded estimates of the “labeling ratio” (LR), or the mass ratio of the labeled to the unlabeled peptide, and the “relative isotope abundance” (RIA), or atom percentage of 13C incorporated into the labeled peptide [61]. We combined the total protein concentrations along with their RIA and LR to estimate the total 13C-protein produced per MAG:
where j represents proteins within a given MAG (MAGn), \(\widehat{{{{{{{\rm{RIA}}}}}}}}\) is the mean RIA of all proteins in MAGn within a sample, and \(\widehat{{{{{{{\rm{LR}}}}}}}}\) is the mean LR of all proteins in MAGn within a sample.
Metabolic reconstruction and metabolic modeling
Metabolic reconstruction and modeling were performed using CobraPy [62]. The central carbon and energy metabolism of the three most abundant MAGs (two methanogenic organisms and one syntrophic acetate oxidizing organism) were manually reconstructed based on genome annotations (see above). All reactions were confirmed to be balanced for charge and mass. In total, the metabolic model contained 65 reactions and 82 metabolites (Supplementary Data 2) across four compartments: intracellular space of the three guilds and the extracellular space. Parsimonious flux balance analysis (pFBA), flux variability analysis (FVA), and flux sampling were used to predict flux distributions through the metabolic networks and exchange of metabolic end products between microbial populations. Gibbs free energies were calculated for the population metabolisms based on their stoichiometries predicted by pFBA and FVA, using standard energies of formation of products and reactants and adjustment based on in situ concentrations (see Supplementary Data 2).
Results and discussion
Continuous enrichment of acetate-oxidizing consortia from a full-scale anaerobic digester
An acetate oxidizing microbial community was enriched from the sludge of a full-scale anaerobic digester for 300 days using two parallel continuously-operated chemostat reactors held at 55 °C and fed with anaerobic medium containing acetate as the primary carbon source. Low acetate levels in the effluent of the reactors (~2 mM) relative to the feed (75 mM) and high percentages of methane in the headspace indicated efficient methanogenic conversion of the substrate under the steady-state conditions (Table 1).
Free ammonia is a known driver of microbial community structure in AD microbiomes [63, 64]. In particular high free ammonia levels in AD have been associated with inhibition of acetoclastic methanogenesis and a corresponding shift toward SAO [9, 65]. In mesophilic AD communities, the reported critical free ammonia level for this shift is around 140 to 280 mg NH3-N/l [65], while in thermophilic AD systems this shift toward SAO has been observed at 200 to 500 mg NH3-N/l [9, 66, 67]. Based on the temperature, pH, and total ammonia nitrogen, the free ammonia in the reactors averaged 445 mg NH3-N/l (Supplementary Text). As this level was above reported threshold concentrations for the inhibition of acetoclastic methanogenesis in thermophilic environments, we posit that favorable conditions existed for the conversion of acetate into methane through obligate mutualistic cross-feeding interactions between SAOB and methanogenic archaea.
Population succession dynamics within acetate-fed enrichment bioreactors
A long-read metagenome assembly of a single bioreactor (R2) community on day 300, as well as individual short-read assemblies of both duplicate bioreactors from 6 operational days over the 300-day period, were used to generate a de-replicated set of 60 medium and high-quality MAGs [68] (Fig. 1 and Supplementary Data 1). The recovered set of MAGs from the enrichment chemostats in this study spanned most of the phyla represented in a previously reported biogas microbiome reference database [69] (Fig. 1C).
A Conceptual schematic of 300-day chemostat enrichment period of the acetate oxidizing consortium showing the dates that short-read metagenomes were sequenced with Illumina and the long-read metagenome sequenced with Nanopore; B The relative read abundance in the short-read metagenomes of the recovered set of de-replicated MAGs within bioreactor R2, which was the bioreactor sampled for the stable isotope probing experiment. The relative read abundance is the number of reads mapped to a given genomic entity divided by the total reads in the sample. Each color in the plot represents a different genomic entity (e.g., a MAG or group of MAGs). C Phylogenetic tree of the recovered MAGs and a subset of biogas genome references from Campanaro et al. [69]. Moving from inside to outside in the colored rings: the first (inner) ring shows a phylum level taxonomic classification, the second ring indicates what study the genome originates from, the third ring shows genome completeness, the fourth shows contamination, and the fifth (outer) ring shows the number of scaffolds in the genome. The tree was constructed from ribosomal protein markers with metabolisHMM v2.21 [101] by searching for markers with hmmsearch as part of the HMMER v3.2.1 suite [102], aligning hits for each marker with muscle v3.8.31 [103], and building the phylogenetic tree with fasttree v2.1.11 [104]. The tree was visualized and metadata overlaid on the tree with EMPRESS v1.2.0. [105].
Throughout the continuous chemostat operation, the recovered set of 60 de-replicated MAGs was enriched from an initial 40% cumulative read abundance on day 0 to over 85% by day 283 (Fig. 1B and Supplementary Data 1). During the enrichment period, there was a notable washout of MAGs belonging to Coprothermobacter and Caldatribacteriota (two-tailed Student’s t test, days 0–19 (n = 4) vs. days 234–283 (n = 4); p = 4e−5 and 5e−3, respectively) (Fig. 1B). On the other hand, MAGs from the genera DTU068 and Methanothermobacter_A significantly increased in abundance throughout the chemostat operation (two-tailed Student’s t test, days 0–19 (n = 4) vs. days 234–283 (n = 4); p = 6e−4 and 1e−4, respectively), together increasing from 13% read abundance initially to over 72% after day 81 (Fig. 1B). No archaeal MAGs were recovered belonging to the acetoclastic families (Methanotrichaceae and Methanosarcinaceae) (Fig. 1B). However, short-read metagenome decomposition using k-mer signatures revealed that the mixotrophic acetoclastic genus, Methanosarcina, was initially present at roughly 6% abundance and decreased to below 0.1% by the end of operation (two-tailed Student’s t test, days 0–19 (n = 4) vs. days 234–283 (n = 4); p = 8e−3) (Supplementary Fig. S1). Therefore, a highly enriched anaerobic acetate-oxidizing consortium was obtained within 81 days of chemostat operation. The consortium was represented primarily by genomic populations belonging to DTU068 and Methanothermobacter_A lineages.
The populations belonging to Methanothermobacter_A and DTU068 underwent a dynamic succession over time during the chemostat enrichment. The Methanothermobacter_2 MAG was initially dominant at 12% read abundance vs. 1% for the Methanothermobacter_1 MAG, but it was superseded by Methanothermobacter_1 by the end of the enrichment period (25% vs. 50% read abundance, respectively; Fig. 1B). Along similar lines, the two MAGs belonging to the genus DTU068 also underwent population shifts during the enrichment period. Both DTU068_1 and DTU068_2 MAGs were initially at relatively low read abundances of 0.04% and 0.3%, respectively (Supplementary Data 1). DTU068_2 remained more abundant than DTU068_1 until day 54, reaching 5.5% read abundance vs. 0.6%, respectively. Subsequently, DTU068_1 became the dominant bacterial genome by day 283 reaching 6% read abundance, while DTU068_2 decreased to 0.5% (Fig. 1B). The cause of such population shifts within DTU068 and Methanothermobacter_A throughout the enrichment was uncertain, and could potentially be attributed to phage-host dynamics [70] and/or to the establishment of population-specific mutualistic relationships between syntrophic bacteria and methanogens [71].
The Methanothermobacter_1 and Methanothermobacter_2 MAGs represented different species with a shared ANI of 83.3% (Supplementary Fig. S2), which is below the 95% ANI cutoff considered for microbial species designation [47]. The Methanothermobacter_1 MAG shared its highest ANI of 99.5% with Methanothermobacter_A sp012840175, while Methanothermobacter_2 shared its highest ANI of 98.8% with Methanothermobacter_A sp003584625 (Supplementary Fig. S2). Notably, these closest relatives of Methanothermobacter_1 and Methanothermobacter_2 are so-far uncultured. Within the GTDB taxonomic hierarchy (release 207), the archaeal lineage of Methanothermobacter is divided into two genera, Methanothermobacter_A and Methanothermobacter. Genomes of the representative species M. tenebrarum [72] fall within the genus Methanothermobacter_A, while M. marburgensis, M. thermautotrophicus, and M. wolfeii [73, 74] fall within the Methanothermobacter genus (Supplementary Fig. S2). These cultured members of Methanothermobacter have been characterized to grow optimally at 55 to 65 °C by reducing carbon dioxide to methane using hydrogen, and sometimes formate [73], as electron donors [75]. Several species of Methanothermobacter have been isolated from thermophilic municipal sludge anaerobic digesters [73, 74], such as that used as an inoculum source in this study.
Based on genome similarity, DTU068_1 and DTU068_2 MAGs also represented distinct species with 91.6% shared ANI (Supplementary Fig. S3). DTU068_1 shared its highest ANI (99.2%) with DTU068 sp001513545, while DTU068_2 shared its highest ANI (98.6%) with DTU068 sp012840405 (Supplementary Fig. S3). DTU068 represents a so-far uncultured genus within the Thermacetogeniaceae family and Firmicutes phylum (according to GTDB release 207; Thermoanaerobacteraceae family in NCBI taxonomy). MAGs from DTU068 have been hypothesized to participate in SAO based on genome-resolved transcriptomic expression of the Wood-Ljungdahl pathway in a thermophilic (55 °C) manure-fed AD system [76]. To the best of our knowledge, members of DTU068 have not been enriched to the levels observed in this study. In a meta-analysis of 1635 MAGs recovered from 134 anaerobic digestion metagenomes [69], the maximum relative abundance of any DTU068-related MAG was 3.5%, which occurred in the second (methanogenic) phase of a two-phase thermophilic (55 °C) reactor system fed with cheese whey [77]. Based on these observations, the genus DTU068 appears to contain species that thrive in thermophilic anoxic environments and may harbor genes involved in acetate oxidation.
Time-resolved stable isotope probing metaproteogenomics of enriched community
Stable isotope probing with 13C-labeled acetate (50 mM; 3000 mg/l) was conducted on the enriched acetate oxidizing members to track their metabolism and identify potential interspecies interactions (Fig. 2A). Over the 408-h incubation, the acetate-fed microcosms produced an average of 17.5 ml CH4 (at STP) in excess of the unfed controls (Fig. 2B), corresponding to a theoretical conversion of 88% of the supplied acetate (see Supplementary Text). In concordance with this, the measured acetate concentrations in the microcosms decreased from 3000 mg/l to 280 ± 136 mg/l over 408 h (Fig. 2B and Supplementary Table S2). These results indicate a near-complete conversion of the supplied acetate to CH4 in the SIP microcosms, as well as repeatable trends among biological replicates.
A Experimental overview of the stable isotope probing (SIP) microcosms and metaproteogenomic analysis; B Cumulative methane production and acetate concentrations over time in the SIP microcosms fed with acetate, along with the unfed control (blank). Shaded regions represent the standard error of biological triplicates. C Ratio (%) of atom-percent 13CO2 to that of 13CH4 measured in the headspace of the SIP microcosms fed with 2-13C (methyl-C labeled) acetate, corrected for background 12C from dissolved inorganic carbon (see Supplementary Tables S3–S5).
The methyl carbon of acetate is converted into CO2 during SAO, but reduced to form CH4 during acetoclastic methanogenesis [78]. Therefore, we tracked 13C:12C ratios in the generated CO2 and CH4 gasses within SIP microcosms fed with 2-13C (methyl-C labeled) and 1,2-13C (universally labeled) acetate to estimate the relative activity of the SAO pathway vs. acetoclastic methanogenesis [78] (Supplementary Tables S3 and S4). The atom-percent ratio of 13CO2:13CH4 in the 2-13C acetate-fed microcosms ranged from 1.02 to 1.21 (Fig. 2C and Supplementary Table S4), indicating that most of the methyl carbon was first oxidized to CO2 via SAO before being reduced to CH4 via hydrogenotrophic methanogenesis. In support of this, we fitted the isotope-partitioning model proposed by Mulat et al. [78] to our experimental data, which suggested that the SAO pathway accounted for 98% ± 5% of the carbon flux from acetate to methane throughout the SIP microcosms (Supplementary Table S5).
At the time of the SIP incubation, metagenomic sequencing indicated that Methanothermobacter_1 was the most abundant genome (50%) based on fraction of reads mapped, followed by Methanothermobacter_2 (25%) and DTU068_1 (6%) (Fig. 1B). Correspondingly, 91% ± 4% of the metaproteome was attributed to those three MAGs throughout the SIP incubation, with Methanothermobacter_1 accounting for 50% ± 1%, followed by DTU068_1 at 23% ± 3% and Methanothermobacter_2 at 18% ± 3% (Fig. 3A). The extent of isotope incorporation into proteins indicated that the 13C atom percent labeling (i.e., relative isotope abundance, RIA) and the abundance of labeled peptides (i.e., labeling ratio, LR) increased across most community members throughout the SIP incubation (Supplementary Fig. S4). Approximately 80% of the identified 13C-labeled peptides were mapped to the three members of Methanothermobacter_1 (39%), Methanothermobacter_2 (16%), and DTU068_1 (25%) (Fig. 3B). In total, we detected 7879 13C-labeled peptides throughout the 408 h SIP incubation, which is orders of magnitude greater than a previous observation of 61 total 13C labeled peptides detected after 196 h of incubation with 100 mM 13C-acetate in a mixed anaerobic digestion community [30]. This finding highlights the benefits of long-term community enrichment prior to SIP to gain deeper insights into metabolic activities and carbon flux through SAO populations that are typically rare or in low-abundance in AD systems (Fig. 1B; [20, 24, 65, 79]).
Heatmaps of (A) relative protein abundance (log10-scaled, based on label-free quantification) and (B) the number of 13C-labeled peptides identified, for the 11 most abundant MAGs at 24, 144, and 408 h of the SIP incubation. Values from biological triplicates are shown for each time point sampled. C The 13C-labeled protein concentration (mg/L) inferred from the total protein quantification approach, relative isotope abundance (RIA), and labeling ratio (LR) of all proteins from MAGs throughout the SIP incubation. Shaded regions indicate the standard error across biological triplicates, accounting for variance in mean RIA and LR across all proteins in the genome. The MAG names used in this figure are derived from Supplementary Data 1.
To estimate carbon fluxes into different community members within the SIP incubation, we combined the total protein quantification approach [60] with the 13C atom percent labeling (RIA) and labeled-protein abundance (LR) to obtain estimated concentrations of 13C-labeled proteins per MAG over time (Fig. 3C). This analysis indicated that Methanothermobacter_1 produced the most 13C-labeled protein (8 mg/L), followed by DTU068_1 (4 mg/L) and Methanothermobacter_2 (2 mg/L), while all other MAGs accounted for 1 mg/L of 13C-labeled protein combined (Fig. 3C). As the 13C-protein was directly produced from the added 13C-acetate, we estimated an overall community biomass yield for conversion of acetate into methane of 0.01 g-VSS g-acetate−1 (0.015 g-CODbiomass g-CODacetate−1), assuming: a protein-to-biomass ratio of 0.5 g-protein g-VSS−1 [80], and a VSS-to-COD ratio of 1.42 g-CODbiomass g-VSS−1. This estimated yield is within the range reported for defined co-cultures of syntrophic propionate-oxidizing bacteria and methanogenic partners of 0.011 to 0.016 g-CODbiomass g-COD−1 using total protein measurements [81, 82]. Measurements of biomass yields for individual species within syntrophic fatty acid-oxidizing communities are sparse [83], and obtaining absolute biomass estimates for each member separately during growth typically relies on quantitative-PCR [84,85,86]. Here, we show that quantitatively tracking carbon fluxes into the biomass of individual community members using SIP metaproteomics represents a powerful approach that could help inform ecosystem level models for uncultured microbiomes.
Metabolic reconstructions and modeling of the SAO community
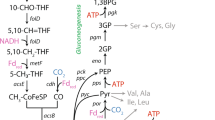
Metabolic reconstructions of the three most abundant MAGs (DTU068_1, Methanothermobacter_1, and Methanothermobacter_2) were created based on predicted functions of expressed proteins (Figs. 4 and 5). In total, DTU068_1 is predicted to oxidize acetate to formate and CO2 via the oxidative acetyl-coA pathway (i.e., reverse Wood-Ljungdahl pathway), while producing H2 and formate to maintain redox balance. Multiple hydrogen-producing enzymes were found in the DTU068_1 proteome: a NADH-dependent (Group 3b) [NiFe]-hydrogenase ([NiFe]-HydABC), a membrane-bound periplasm-facing (Group 1a) [NiFe]-hydrogenase ([NiFe]-HysAB-Cytb), [FeFe] electron-bifurcating (Group A3) hydrogenases ([FeFe]-HydABC), and a proton-translocating energy-conserving (Group 4e) hydrogenase (EchABCDEF) (Figs. 4 and 5). Formate is predicted to be produced intracellularly through formate-tetrahydrofolate ligase (Fhs), as well as extracellularly via a membrane-associated formate dehydrogenase (Fdh) complex. This membrane-associated Fdh complex and the periplasm-facing [NiFe]-hydrogenase both contain a cytochrome-b subunit (Fig. 4), and are predicted to participate in reverse electron transport from heterodisulfide reductase (HdrABC/MvhD) to drive the endergonic oxidation of methyl-THF to methylene-THF via methylene-THF reductase (MetFV) [87]. An electron-bifurcating FdhA-NuoEF complex was also found that could reversibly oxidize formate to CO2 while producing reduced ferredoxin and NADH (Fig. 5). Overall, the predicted pathway for the oxidation of acetate into CO2, H2, and formate in DTU068_1 was similar to that proposed for Thermacetogenium phaeum, except that DTU068_1 is proposed to utilize acetate kinase/phosphotransacetylase (Ack/Pta) to activate acetate to acetyl-coA rather than acetaldehyde oxidoreductase [19], as well as utilize electron-bifurcating FdhA-NuoEF and HydABC complexes for energy conservation (Figs. 4 and 5).
Total protein expression (nM; log10-scaled) for enzymes of interest throughout the acetate-fed SIP incubations in (A) DTU068_1, (B) Methanothermobacter_1, (C) Methanothermobacter_2. The vertical facets represent different sampling time points (24, 144, and 408 h), and the horizontal facets represent protein groups based on different metabolic functions and/or protein complexes. The value labeled “Median” at the bottom represents the genome-wide median protein expression. Values are shown for biological triplicates. For each protein unit, the associated gene locus is given in parentheses next to the name. Proteins in Methanothermobacter_1 and Methanothermobacter_2 that have an asterisk (*) indicate these associated subunits were identical within the two genomes, and thus the shown protein abundance represents this redundancy. Protein abbreviations: Ack acetate kinase, Acs acetyl-coA synthase/carbon monoxide dehydrogenase (CODH), CooC Acs accessory protein, Cyt cytochrome, DUF domain of unknown function, Ech energy-conserving hydrogenase, Eha energy-converting hydrogenase, Fdh formate dehydrogenase, Fhs formate-THF ligase, Fol methenyl-THF cyclohydrolase, Frh F420-reducing hydrogenase, Ftr formyl-MFR:H4MPT formyltransferase, Fwd formyl-MFR dehydrogenase, Hdr heterodisulfide reductase, Hmd H2-dependent methylene-H4MPT dehydrogenase, Hya hydrogenase, Hyd hydrogenase, Mch methenyl-H4MPT cyclohydrolase, Mer methylene-H4MPT reductase, Mtd F420-dependent methylene-H4MPT dehydrogenase, MTase methyltransferase, Mtr F420-dependent methylene-H4MPT dehydrogenase, Mvh F420-non-reducing hydrogenase, Mcr methyl-CoM reductase, Nuo NADH:ubiquinone oxidoreductase, Pta phosphotransacetylase.
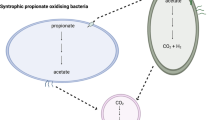
Cell diagrams showing the predicted metabolic pathways for acetate oxidation in DTU068_1 and methane generation from hydrogen/formate in Methanothermobacter_1 and Methanothermobacter_2. Values of predicted flux, obtained from parsimonious flux balance analysis, are shown in red text within boxes. Net catabolic reactions are based on stoichiometry obtained from parsimonious flux balance analysis. Protein abbreviations are defined in the legend of Fig. 4.
Both Methanothermobacter_1 and Methanothermobacter_2 contain identical pathways for methane production from CO2 (Figs. 4 and 5). A major difference lies in their utilization of formate vs. H2 as electron donors. Interspecies electron transport via formate was previously shown to occur during the syntrophic oxidation of acetate [88], propionate [89], and butyrate [90], and has been suggested to permit a greater transfer rate than hydrogen by maintaining feasible thermodynamics across a larger concentration gradient [91]. Formate was detected as one of the most abundant metabolites (besides acetate) in the SIP incubations, ranging in concentration from 3 to 7 μM (Supplementary Table S6). Moreover, the 13C-labeling ratio of formate increased over time (Supplementary Table S7), indicating that it was likely a product of acetate oxidation. Within Methanothermobacter spp., the ability of M. thermautotrophicus Z-245 to grow on formate was attributed to a fdhAB gene cluster adjacent to a formate transporter (fdhC) and a carbonic anhydrase (CA) [92]. The role of this fdhCAB gene cluster in growth on formate was recently confirmed by Fink et al. [93] using a shuttle-vector system to amend the canonical non-formate-utilizing M. thermautotrophicus ΔH with this operon, which then grew and produced methane from formate. The fdhCAB gene cluster was also shown to be essential for growth on formate in the archaeon, Methanococcus maripaludis [94]. We queried all sequenced Methanothermobacter genomes to-date and found that all species capable of growth on formate as an electron donor possess the fdhCAB gene cluster, which was not observed within the genomes of Methanothermobacter species incapable of growth on formate (Fig. 6A). Methanothermobacter_2 was found to contain this fdhCAB gene cluster (Fig. 6B), and the FdhABC protein cluster was within the 97 ± 1 percentile of its proteome expression throughout the SIP incubation (Fig. 4). In contrast, Methanothermobacter_1 did not possess the fdhCAB gene cluster (Fig. 6A), and a search of unbinned contigs and unassembled reads confirmed that the only archaeal fdhC within the metagenome belonged to Methanothermobacter_2 (Supplementary Text; Supplementary Tables S8 and S9). While both Methanothermobacter_1 and Methanothermobacter_2 MAGs encode for a FdhA unit upstream of the tungsten formylmethanofuran dehydrogenase (fwd) gene cluster, we found this fdhA-fwd gene arrangement was present in all sequenced Methanothermobacter genomes to-date (Fig. 6C), even among members known to not utilize formate as an electron donor like M. tenebrarum sp. RMAS [73], M. thermautotrophicus ΔH [74], and M. marburgensis Marburg [75]. This fdhA-fwd gene cluster arrangement was previously reported for the strain M. thermautotrophicus ΔH, and the FdhA unit was deemed a “formate dehydrogenase-like protein (FlpF)” due to an N-terminal extension of about 200 amino acids with binding motifs for two [4Fe-4S] clusters [95]. Thus, the function of this FdhA/FlpF enzyme is not clear. Therefore, we predict Methanothermobacter_2 can oxidize formate via FdhABC to reduce F420 for growth, while Methanothermobacter_1 is predicted to solely grow on H2 through its highly expressed methyl-viologen-reducing hydrogenase (MvhABDG) and an F420-reducing [NiFe]-hydrogenase (FrhABDG) (Figs. 4 and 5). As DTU068_1 is predicted to produce both H2 and formate during the oxidation of acetate (Fig. 5), we posit that this diversity of electron donors could have supported the apparent niche partitioning of methanogenic partners adapted for exclusive or preferential modes of interspecies electron transfer.
A Heatmap showing the ability to grow on formate [73, 75, 106] and the presence/absence of the fdhCAB gene cluster and fdhA-fwd gene cluster for all available sequenced Methanothermobacter genomes, along with the two Methanothermobacter MAGs from this study. The yellow borders used for the formate growth values of the two MAGs indicates that these are inferred traits based on the results of this study. B Map of the genomic arrangement of the formate dehydrogenase-carbonic anhydrase gene cluster found only within genomes of certain Methanothermobacter species, all of which are known to grow on formate, and (C) the tungsten formylmethanofuran dehydrogenase gene cluster observed in all available representative Methanothermobacter genomes. The gene clusters for the Methanothermobacter_A MAGs identified in this study are also shown. The direction of the gene arrows indicate their direction of transcription. NCBI RefSeq accessions of each representative genome is given in parenthesis below its species name. Each gene is labeled with its locus tag, which is sometimes located above/below a gene arrow for visualization purposes. fwd tungsten formylmethanofuran dehydrogenase, fdh formate dehydrogenase, CA carbonic anhydrase, moa GTP 3’,8’-cyclase, mob molybdopterin-guanine dinucleotide biosynthesis-domain protein.
To further examine potential syntrophic relationships between DTU068_1, Methanothermobacter_1, and Methanothermobacter_2, we implemented a metabolic model containing the central carbon and energy metabolism of each guild (Fig. 5). Possible flux distributions were predicted using parsimonious flux balance analysis while constraining the relative ATP yield of each of the three populations to their relative proteome contribution (Fig. 3C). With these constraints, the maximum ATP yield of the entire community was estimated at 0.706 mol ATP mol−1 acetate. The metabolic model predicts that DTU068_1 consumes 1 mol acetate and produces 1 mol formate intracellularly. Of the intracellular formate, 0.824 mol is oxidized via an electron-confurcating FdhA-NuoEF complex (Fig. 5). Further, 0.045 mol formate is produced extracellularly using the FdhA-FdxH-CytB complex. The ion motive force (IMF) in DTU068_1 is created by EchABCDEF, and consumed via ATP synthase and to drive reverse electron flow from HdrABC to CO2 and H+. In total, DTU068_1 is predicted to produce a net of 0.221 mol formate, 1.779 mol CO2, and 3.779 mol H2 from 1 mol of acetate (Fig. 5).
Overall, the guild-level metabolic model supported the hypothesis that the two methanogens underwent niche partitioning based on their preferred electron donors (e.g., H2 or formate). Methanothermobacter_2 is predicted to consume the 0.221 mol formate produced by DTU068_1, along with 0.663 mol H2 (Fig. 5). Methanothermobacter_1 is predicted to consume 3.116 mol H2 and 0.779 mol CO2. In both methanogens, H2 is consumed via the MvhADG-HdrABC and FrhABG complexes (Fig. 5). Both methanogens also utilize the tetrahydromethanopterin S-methyltransferase complex (MtrABCDEFGH) for IMF generation, which is used for ATP generation with ATP synthase. In total, Methanothermobacter_1 is predicted to generate 0.779 mol CH4 (78% of evolved CH4), and Methanothermobacter_2 is predicted to produce 0.221 mol CH4 (22% of evolved CH4). The total community ATP production of 0.706 mol ATP and the overall free energy release of −35.8 kJ suggest a net free energy release of −51.1 kJ/mol ATP under standard conditions (Fig. 5), which is likely sufficient to support growth under such energy-limited conditions in anaerobic systems [96, 97].
The predicted consumption of intracellular formate by DTU068_1 to generate reduced ferredoxin, along with the consumption of extracellular formate by Methanothermobacter_2 to drive methanogenesis (Fig. 5), raises the question of whether interspecies electron transfer via formate between these two species represented a mutualistic or competitive interaction. To assess this question, we established a community-scale metabolic model for DTU068_1 grown solely in the presence of Methanothermobacter_1 (e.g., no Methanothermobacter_2) (Supplementary Data 2). As expected, when Methanothermobacter_2 is not present, DTU068_1 is predicted to consume all of its produced formate via the intracellular electron-bifurcating FdhA-NuoEF complex to generate H2, all of which is consumed by Methanothermobacter_1 to drive methanogenesis (Supplementary Data 2). Interestingly, while the net standard free energy release from 1 mole of acetate of −35.8 kJ is identical, the predicted overall community ATP yield in this scenario is 0.765 moles, which is 8% higher than when Methanothermobacter_2 is present (Fig. 5). However, an examination of the thermodynamic feasibility of the community metabolism under both scenarios revealed that more favorable energetics for DTU068_1 and Methanothermobacter_1 are achieved across a wider range of H2 partial pressures in the presence of Methanothermobacter_2 than without (Fig. 7). Thus, interspecies electron transfer via formate from DTU068_1 to Methanothermobacter_2 does appear to be mutualistic. By shunting electrons to both formate and H2 during acetate oxidation, DTU068_1 can establish more favorable energetics for ATP production at the potential sacrifice of net community ATP yield.
Free energy yields (ΔG) per mole of ATP produced for each member of the syntrophic acetate oxidizing consortium as a function of hydrogen partial pressure (PH2), for the cases where the Methanothermobacter_2 MAG (A) is present; and (B) is not present. The dashed gray line represents the ATP phosphorylation potential measured in cells performing acetogenesis from H2 and CO2 (−32.1 kJ/mol-ATP) [96]. Shaded regions represent the ranges of hydrogen partial pressure that would support ATP synthesis by DTU068_1 and Methanothermobacter_1 (yellow region) or DTU068_1 and Methanothermobacter_2 (red region). The free energy values were calculated based on reaction stoichiometry predicted by the parsimonious flux balance analysis model for both cases (Supplementary Data 2), assuming environmentally-relevant concentrations of acetate (50 mM), formate (7.5 μM), methane (0.5atm) and carbon dioxide (0.5atm).
While many community-scale models consider the optimization of ATP or biomass yield to be the overarching objective of microbial community metabolism [98, 99], the above finding indicates that anaerobic microbes surviving near thermodynamic limits of life [6] may optimize the energetic favorability of their community metabolism. Previous modeling efforts informed by multi-omics on syntrophic communities of defined isolates have identified electron transfer via different metabolites to be favorable on conditional bases [100]. Our current work using metaproteogenomics-informed SIP builds upon those findings by identifying flexibility in central metabolic processes and electron partitioning that likely governs community composition and fitness through thermodynamic-driven mutualism in so-far uncultured microbes. These insights provide a more nuanced data-driven perspective on community-level modeling of obligate cross-feeding metabolisms driving carbon flux in anoxic ecosystems, such as the case of SAO in AD processes, as well as in the design of synthetic communities for high-value product generation from waste streams.
Data availability
All raw metagenomes and de-replicated MAGs are available on NCBI at the Bioproject Accession PRJNA885503 (Supplementary Table S1). Metaproteomic MS data are available on ProteomeXchange as dataset PXD042127. All data files including assemblies, annotated genomes, and metaproteomics results are available on OSF at https://osf.io/kdnms/.
Code availability
All analysis code, including the metabolic model, is available in the repository: https://github.com/ZielsLab/SAOB_Metaproteogenomics.
References
Nobu MK, Narihiro T, Mei R, Kamagata Y, Lee PKH, Lee P-H, et al. Catabolism and interactions of uncultured organisms shaped by eco-thermodynamics in methanogenic bioprocesses. Microbiome. 2020;8:111.
Conrad R. Contribution of hydrogen to methane production and control of hydrogen concentrations in methanogenic soils and sediments. FEMS Microbiol Ecol. 1999;28:193–202.
Conklin AS, Chapman T, Zahller JD, Stensel HD, Ferguson JF. Monitoring the role of aceticlasts in anaerobic digestion: activity and capacity. Water Res. 2008;42:4895–904.
Jetten MSM, Stams AJM, Zehnder AJB. Methanogenesis from acetate: a comparison of the acetate metabolism in Methanothrix soehngenii and Methanosarcina spp. FEMS Microbiol Lett. 1992;88:181–97.
Zinder SH, Koch M. Non-aceticlastic methanogenesis from acetate: acetate oxidation by a thermophilic syntrophic coculture. Arch Microbiol. 1984;138:263–72.
Schink B. Energetics of syntrophic cooperation in methanogenic degradation. Microbiol Mol Biol Rev. 1997;61:262–80.
Ziels RM, Karlsson A, Beck DAC, Ejlertsson J, Yekta SS, Bjorn A, et al. Microbial community adaptation influences long-chain fatty acid conversion during anaerobic codigestion of fats, oils, and grease with municipal sludge. Water Res. 2016;103:372–82.
Schnürer A, Zellner G, Svensson BH. Mesophilic syntrophic acetate oxidation during methane formation in biogas reactors. FEMS Microbiol Ecol. 1999;29:249–61.
Sun L, Müller B, Westerholm M, Schnürer A. Syntrophic acetate oxidation in industrial CSTR biogas digesters. J Biotechnol. 2014;171:39–44.
Schnürer A, Nordberg Å. Ammonia, a selective agent for methane production by syntrophic acetate oxidation at mesophilic temperature. Water Sci Technol. 2008;57:735–40.
Balk M, Weijma J, Stams AJM. Thermotoga lettingae sp. nov., a novel thermophilic, methanol-degrading bacterium isolated from a thermophilic anaerobic reactor. Int J Syst Evol Microbiol. 2002;52:1361–8.
Hattori S, Kamagata Y, Hanada S, Shoun H. Thermacetogenium phaeum gen. nov., sp. nov., a strictly anaerobic, thermophilic, syntrophic acetate-oxidizing bacterium. Int J Syst Evol Microbiol. 2000;50:1601–9.
Westerholm M, Roos S, Schnürer A. Tepidanaerobacter acetatoxydans sp. nov., an anaerobic, syntrophic acetate-oxidizing bacterium isolated from two ammonium-enriched mesophilic methanogenic processes. Syst Appl Microbiol. 2011;34:260–6.
Schnürer A, Schink B, Svensson BH. Clostridium ultunense sp. nov., a mesophilic bacterium oxidizing acetate in syntrophic association with a hydrogenotrophic methanogenic bacterium. Int J Syst Evol Microbiol. 1996;46:1145–52.
Westerholm M, Roos S, Schnürer A. Syntrophaceticus schinkii gen. nov., sp. nov., an anaerobic, syntrophic acetate-oxidizing bacterium isolated from a mesophilic anaerobic filter. FEMS Microbiol Lett. 2010;309:100–4.
Timmers PHA, Vavourakis CD, Kleerebezem R, Damsté JSS, Muyzer G, Stams AJM, et al. Metabolism and occurrence of methanogenic and sulfate-reducing syntrophic acetate oxidizing communities in haloalkaline environments. Front Microbiol. 2018;9:3039.
Manzoor S, Bongcam-Rudloff E, Schnürer A, Müller B. Genome-guided analysis and whole transcriptome profiling of the mesophilic syntrophic acetate oxidising bacterium Syntrophaceticus schinkii. PLoS ONE. 2016;11:e0166520.
Oehler D, Poehlein A, Leimbach A, Müller N, Daniel R, Gottschalk G, et al. Genome-guided analysis of physiological and morphological traits of the fermentative acetate oxidizer Thermacetogenium phaeum. BMC Genomics. 2012;13:723.
Keller A, Schink B, Müller N. Energy-conserving enzyme systems active during syntrophic acetate oxidation in the thermophilic bacterium Thermacetogenium phaeum. Front Microbiol. 2019;10:2785.
Westerholm M, Müller B, Singh A, Karlsson Lindsjö O, Schnürer A. Detection of novel syntrophic acetate‐oxidizing bacteria from biogas processes by continuous acetate enrichment approaches. Micro Biotechnol. 2017;11:680–93.
Hori T, Sasaki D, Haruta S, Shigematsu T, Ueno Y, Ishii M, et al. Detection of active, potentially acetate-oxidizing syntrophs in an anaerobic digester by flux measurement and formyltetrahydrofolate synthetase (FTHFS) expression profiling. Microbiology. 2011;157:1980–9.
Müller B, Sun L, Westerholm M, Schnürer A. Bacterial community composition and fhs profiles of low- and high-ammonia biogas digesters reveal novel syntrophic acetate-oxidising bacteria. Biotechnol Biofuels. 2016;9:48.
Werner JJ, Garcia ML, Perkins SD, Yarasheski KE, Smith SR, Muegge BD, et al. Microbial community dynamics and stability during an ammonia-induced shift to syntrophic acetate oxidation. Appl Environ Microbiol. 2014;80:3375–83.
Dyksma S, Jansen L, Gallert C. Syntrophic acetate oxidation replaces acetoclastic methanogenesis during thermophilic digestion of biowaste. Microbiome. 2020;8:105.
Nobu MK, Narihiro T, Rinke C, Kamagata Y, Tringe SG, Woyke T, et al. Microbial dark matter ecogenomics reveals complex synergistic networks in a methanogenic bioreactor. ISME J. 2015;9:1710–22.
Radajewski S, Ineson P, Parekh NR, Murrell JC. Stable-isotope probing as a tool in microbial ecology. Nature. 2000;403:646–9.
Ito T, Yoshiguchi K, Ariesyady HD, Okabe S. Identification of a novel acetate-utilizing bacterium belonging to Synergistes group 4 in anaerobic digester sludge. ISME J. 2011;5:1844–56.
Zheng D, Wang H-Z, Gou M, Nobu MK, Narihiro T, Hu B, et al. Identification of novel potential acetate-oxidizing bacteria in thermophilic methanogenic chemostats by DNA stable isotope probing. Appl Microbiol Biotechnol. 2019;103:8631–45.
Hao L, Lü F, Mazéas L, Desmond-Le Quéméner E, Madigou C, Guenne A, et al. Stable isotope probing of acetate fed anaerobic batch incubations shows a partial resistance of acetoclastic methanogenesis catalyzed by Methanosarcina to sudden increase of ammonia level. Water Res. 2015;69:90–9.
Mosbæk F, Kjeldal H, Mulat DG, Albertsen M, Ward AJ, Feilberg A, et al. Identification of syntrophic acetate-oxidizing bacteria in anaerobic digesters by combined protein-based stable isotope probing and metagenomics. ISME J. 2016;10:2405–18.
Plugge CM. Anoxic media design, preparation, and considerations. Methods Enzymol. 2005;397:3–16.
Nierychlo M, Andersen KS, Xu Y, Green N, Jiang C, Albertsen M, et al. MiDAS 3: an ecosystem-specific reference database, taxonomy and knowledge platform for activated sludge and anaerobic digesters reveals species-level microbiome composition of activated sludge. Water Res. 2020;182:115955.
Bushnell B, Rood J, Singer E. BBMerge—accurate paired shotgun read merging via overlap. PLoS ONE. 2017;12:e0185056.
Brown CT, Irber L. sourmash: a library for MinHash sketching of DNA. J Open Source Softw. 2016;1:27.
Pierce NT, Irber L, Reiter T, Brooks P, Brown CT. Large-scale sequence comparisons with sourmash. F1000Research. 2019;8:1006.
Parks DH, Chuvochina M, Waite DW, Rinke C, Skarshewski A, Chaumeil P-A, et al. A standardized bacterial taxonomy based on genome phylogeny substantially revises the tree of life. Nat Biotechnol. 2018;36:996.
Nurk S, Meleshko D, Korobeynikov A, Pevzner PA. metaSPAdes: a new versatile metagenomic assembler. Genome Res. 2017;27:824–34.
Langmead B, Salzberg SL. Fast gapped-read alignment with Bowtie 2. Nat Methods. 2012;9:357–9.
Kang DD, Froula J, Egan R, Wang Z. MetaBAT, an efficient tool for accurately reconstructing single genomes from complex microbial communities. PeerJ. 2015;3:e1165.
Olm MR, Brown CT, Brooks B, Banfield JF. DRep: a tool for fast and accurate genomic comparisons that enables improved genome recovery from metagenomes through de-replication. ISME J. 2017;11:2864–8.
Parks DH, Imelfort M, Skennerton CT, Hugenholtz P, Tyson GW. CheckM: assessing the quality of microbial genomes recovered from isolates, single cells, and metagenomes. Genome Res. 2015;25:1043–55.
Kolmogorov M, Bickhart DM, Behsaz B, Gurevich A, Rayko M, Shin SB, et al. metaFlye: scalable long-read metagenome assembly using repeat graphs. Nat Methods. 2020;17:1103–10.
Vaser R, Sović I, Nagarajan N, Šikić, M. Fast and accurate de novo genome assembly from long uncorrected reads. Genome Research. 2017;27:737–46.
Eren AM, Esen ÖC, Quince C, Vineis JH, Morrison HG, Sogin ML, et al. Anvi’o: an advanced analysis and visualization platform for ‘omics data. PeerJ. 2015;3:e1319.
Karst SM, Kirkegaard RH, Albertsen M. mmgenome: a toolbox for reproducible genome extraction from metagenomes. bioRxiv. 2016. https://doi.org/10.1101/059121.
Chaumeil P-A, Mussig AJ, Hugenholtz P, Parks DH. GTDB-Tk: a toolkit to classify genomes with the Genome Taxonomy Database. Bioinformatics. 2019;36:1925–7.
Jain C, Rodriguez-R LM, Phillippy AM, Konstantinidis KT, Aluru S. High throughput ANI analysis of 90K prokaryotic genomes reveals clear species boundaries. Nat Commun. 2018;9:5114.
Hyatt D, Chen G-L, Locascio PF, Land ML, Larimer FW, Hauser LJ. Prodigal: prokaryotic gene recognition and translation initiation site identification. BMC Bioinform. 2010;11:119.
Aramaki T, Blanc-Mathieu R, Endo H, Ohkubo K, Kanehisa M, Goto S, et al. KofamKOALA: KEGG ortholog assignment based on profile HMM and adaptive score threshold. bioRxiv. 2019. https://doi.org/10.1101/602110.
Konwar KM, Hanson NW, Pagé AP, Hallam SJ. MetaPathways: a modular pipeline for constructing pathway/genome databases from environmental sequence information. BMC Bioinform. 2013;14:202.
Hanson NW, Konwar KM, Wu S-J, Hallam SJ. MetaPathways v2.0: a master-worker model for environmental Pathway/Genome Database construction on grids and clouds. 2014 IEEE Conference on Computational Intelligence in Bioinformatics and Computational Biology. 2014; IEEE. p. 1–7.
Losey NA, Poudel S, Boyd ES, McInerney MJ. The beta subunit of non-bifurcating NADH-dependent [FeFe]-hydrogenases differs from those of multimeric electron-bifurcating [FeFe]-hydrogenases. Front Microbiol. 2020;11:1109.
Søndergaard D, Pedersen CNS, Greening C. HydDB: a web tool for hydrogenase classification and analysis. Sci Rep. 2016;6:34212.
Röst HL, Sachsenberg T, Aiche S, Bielow C, Weisser H, Aicheler F, et al. OpenMS: a flexible open-source software platform for mass spectrometry data analysis. Nat Methods. 2016;13:741–8.
Berthold MR, Cebron N, Dill F, Gabriel TR, Kötter T, Meinl T, et al. KNIME—the Konstanz information miner: version 2.0 and beyond. ACM SIGKDD Explor Newsl. 2009;11:26–31.
Kim S, Pevzner PA. MS-GF+ makes progress towards a universal database search tool for proteomics. Nat Commun. 2014;5:5277.
Käll L, Canterbury JD, Weston J, Noble WS, MacCoss MJ. Semi-supervised learning for peptide identification from shotgun proteomics datasets. Nat Methods. 2007;4:923–5.
Serang O, MacCoss MJ, Noble WS. Efficient marginalization to compute protein posterior probabilities from shotgun mass spectrometry data. J Proteome Res. 2010;9:5346–57.
Wiśniewski JR, Rakus D. Multi-enzyme digestion FASP and the ‘Total Protein Approach’-based absolute quantification of the Escherichia coli proteome. J Proteom. 2014;109:322–31.
Delogu F, Kunath BJ, Evans PN, Arntzen MØ, Hvidsten TR, Pope PB. Integration of absolute multi-omics reveals dynamic protein-to-RNA ratios and metabolic interplay within mixed-domain microbiomes. Nat Commun. 2020;11:4708.
Sachsenberg T, Herbst F-A, Taubert M, Kermer R, Jehmlich N, von Bergen M, et al. MetaProSIP: automated inference of stable isotope incorporation rates in proteins for functional metaproteomics. J Proteome Res. 2015;14:619–27.
Ebrahim A, Lerman JA, Palsson BO, Hyduke DR. COBRApy: COnstraints-Based Reconstruction and Analysis for Python. BMC Syst Biol. 2013;7:74.
Ziels RM, Svensson BH, Sundberg C, Larsson M, Karlsson A, Yekta SS. Microbial rRNA gene expression and co-occurrence profiles associate with biokinetics and elemental composition in full-scale anaerobic digesters. Micro Biotechnol. 2018;11:694–709.
De Vrieze J, Saunders AM, He Y, Fang J, Nielsen PH, Verstraete W, et al. Ammonia and temperature determine potential clustering in the anaerobic digestion microbiome. Water Res. 2015;75:312–23.
Westerholm M, Moestedt J, Schnürer A. Biogas production through syntrophic acetate oxidation and deliberate operating strategies for improved digester performance. Appl Energy. 2016;179:124–35.
Hao L, Fan L, Chapleur O, Guenne A, Bize A, Bureau C, et al. Gradual development of ammonia-induced syntrophic acetate-oxidizing activities under mesophilic and thermophilic conditions quantitatively tracked using multiple isotopic approaches. Water Res. 2021;204:117586.
Fotidis IA, Karakashev D, Angelidaki I. The dominant acetate degradation pathway/methanogenic composition in full-scale anaerobic digesters operating under different ammonia levels. Int J Environ Sci Technol. 2014;11:2087–94.
Bowers RM, Kyrpides NC, Stepanauskas R, Harmon-Smith M, Doud D, Reddy TBK, et al. Minimum information about a single amplified genome (MISAG) and a metagenome-assembled genome (MIMAG) of bacteria and archaea. Nat Biotechnol. 2017;35:nbt.3893.
Campanaro S, Treu L, Rodriguez-R LM, Kovalovszki A, Ziels RM, Maus I, et al. New insights from the biogas microbiome by comprehensive genome-resolved metagenomics of nearly 1600 species originating from multiple anaerobic digesters. Biotechnol Biofuels. 2020;13:25.
Zhang J, Gao Q, Zhang Q, Wang T, Yue H, Wu L, et al. Bacteriophage–prokaryote dynamics and interaction within anaerobic digestion processes across time and space. Microbiome. 2017;5:57.
Embree M, Liu JK, Al-Bassam MM, Zengler K. Networks of energetic and metabolic interactions define dynamics in microbial communities. Proc Natl Acad Sci. 2015;112:15450–5.
Nakamura K, Takahashi A, Mori C, Tamaki H, Mochimaru H, Nakamura K, et al. Methanothermobacter tenebrarum sp. nov., a hydrogenotrophic, thermophilic methanogen isolated from gas-associated formation water of a natural gas field. Int J Syst Evol Microbiol. 2013;63:715–22.
Wasserfallen A, Nölling J, Pfister P, Reeve J, Conway, de Macario E. Phylogenetic analysis of 18 thermophilic Methanobacterium isolates supports the proposals to create a new genus, Methanothermobacter gen. nov., and to reclassify several isolates in three species, Methanothermobacter thermautotrophicus comb. nov., Methanothermobacter wolfeii comb. nov., and Methanothermobacter marburgensis sp. nov. Int J Syst Evol Microbiol. 2000;50:43–53.
Zeikus JG, Wolfe RS. Methanobacterium thermoautotrophicus sp. n., an anaerobic, autotrophic, extreme thermophile. J Bacteriol. 1972;109:707–13.
Boone DR. Methanothermobacter. Bergey’s manual of systematics of archaea and bacteria. In: Whitman WB, editor. John Wiley & Sons, Ltd.; 2015. p. 1–8. https://doi.org/10.1002/9781118960608.
Treu L, Campanaro S, Kougias PG, Zhu X, Angelidaki I. Untangling the effect of fatty acid addition at species level revealed different transcriptional responses of the biogas microbial community members. Environ Sci Technol. 2016;50:6079–90.
Fontana A, Campanaro S, Treu L, Kougias PG, Cappa F, Morelli L, et al. Performance and genome-centric metagenomics of thermophilic single and two-stage anaerobic digesters treating cheese wastes. Water Res. 2018;134:181–91.
Mulat DG, Ward AJ, Adamsen APS, Voigt NV, Nielsen JL, Feilberg A. Quantifying contribution of synthrophic acetate oxidation to methane production in thermophilic anaerobic reactors by membrane inlet mass spectrometry. Environ Sci Technol. 2014;48:2505–11.
Wei Y, Wu Y, Zhang L, Zhou Z, Zhou H, Yan X. Genome recovery and metatranscriptomic confirmation of functional acetate-oxidizing bacteria from enriched anaerobic biogas digesters. Environ Pollut. 2020;265:114843.
Stouthamer AH. A theoretical study on the amount of ATP required for synthesis of microbial cell material. Antonie Van Leeuwenhoek. 1973;39:545–65.
Uhlenhut F, Schlüter K, Gallert C. Wet biowaste digestion: ADM1 model improvement by implementation of known genera and activity of propionate oxidizing bacteria. Water Res. 2018;129:384–93.
Scholten JCM, Conrad R. Energetics of syntrophic propionate oxidation in defined batch and chemostat cocultures. Appl Environ Microbiol. 2000;66:2934–42.
Junicke H. Comment on “A compilation and bioenergetic evaluation of syntrophic microbial growth yields in anaerobic digestion” by Patón, M. and Rodríguez, J. Water Res. 2020;173:115347.
Westerholm M, Dolfing J, Schnürer A. Growth characteristics and thermodynamics of syntrophic acetate oxidizers. Environ Sci Technol. 2019;53:5512–20.
Junicke H, Abbas B, Oentoro J, van Loosdrecht M, Kleerebezem R. Absolute quantification of individual biomass concentrations in a methanogenic coculture. AMB Express. 2014;4:35.
Ziels RM, Beck DAC, Martí M, Gough HL, Stensel HD, Svensson BH. Monitoring the dynamics of syntrophic β-oxidizing bacteria during anaerobic degradation of oleic acid by quantitative PCR. FEMS Microbiol Ecol. 2015;91:fiv028.
Rosenbaum FP, Müller V. Energy conservation under extreme energy limitation: the role of cytochromes and quinones in acetogenic bacteria. Extremophiles. 2021;25:413–24.
Hattori S, Luo H, Shoun H, Kamagata Y. Involvement of formate as an interspecies electron carrier in a syntrophic acetate-oxidizing anaerobic microorganism in coculture with methanogens. J Biosci Bioeng. 2001;91:294–8.
de Bok FAM, Luijten MLGC, Stams AJM. Biochemical evidence for formate transfer in syntrophic propionate-oxidizing cocultures of Syntrophobacter fumaroxidans and Methanospirillum hungatei. Appl Environ Microbiol. 2002;68:4247–52.
Schmidt A, Müller N, Schink B, Schleheck D. A proteomic view at the biochemistry of syntrophic butyrate oxidation in Syntrophomonas wolfei. PLoS ONE. 2013;8:e56905.
Storck T, Virdis B, Batstone DJ. Modelling extracellular limitations for mediated versus direct interspecies electron transfer. ISME J. 2016;10:621–31.
Nölling J, Reeve JN. Growth- and substrate-dependent transcription of the formate dehydrogenase (fdhCAB) operon in Methanobacterium thermoformicicum Z-245. J Bacteriol. 1997;179:899–908.
Fink C, Beblawy S, Enkerlin AM, Mühling L, Angenent LT, Molitor B. A shuttle-vector system allows heterologous gene expression in the thermophilic methanogen Methanothermobacter thermautotrophicus ΔH. mBio. 2021;12:e02766–21.
Wood GE, Haydock AK, Leigh JA. Function and regulation of the formate dehydrogenase genes of the methanogenic archaeon Methanococcus maripaludis. J Bacteriol. 2003;185:2548–54.
Hochheimer A, Schmitz RA, Thauer RK, Hedderich R. The tungsten formylmethanofuran dehydrogenase from Methanobacterium thermoautotrophicum contains sequence motifs characteristic for enzymes containing molybdopterin dinucleotide. Eur J Biochem. 1995;234:910–20.
Spahn S, Brandt K, Müller V. A low phosphorylation potential in the acetogen Acetobacterium woodii reflects its lifestyle at the thermodynamic edge of life. Arch Microbiol. 2015;197:745–51.
Müller V, Hess V. The minimum biological energy quantum. Front Microbiol. 2017;8.
Scarborough MJ, Hamilton JJ, Erb EA, Donohue TJ, Noguera DR. Diagnosing and predicting mixed-culture fermentations with unicellular and guild-based metabolic models. mSystems. 2020;5:e00755-20.
Zomorrodi AR, Maranas CD. OptCom: a multi-level optimization framework for the metabolic modeling and analysis of microbial communities. PLoS Comput Biol. 2012;8:e1002363.
Nagarajan H, Embree M, Rotaru A-E, Shrestha PM, Feist AM, Palsson BØ, et al. Characterization and modelling of interspecies electron transfer mechanisms and microbial community dynamics of a syntrophic association. Nat Commun. 2013;4:2809.
McDaniel EA, Anantharaman K, McMahon KD. metabolisHMM: phylogenomic analysis for exploration of microbial phylogenies and metabolic pathways. bioRxiv. 2019. https://doi.org/10.1101/2019.12.20.884627.
Eddy SR. Accelerated profile HMM searches. PLoS Comput Biol. 2011;7:e1002195.
Edgar RC. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 2004;32:1792–7.
Price MN, Dehal PS, Arkin AP. FastTree 2—approximately maximum-likelihood trees for large alignments. PLoS ONE. 2010;5:e9490.
Cantrell K, Fedarko MW, Rahman G, McDonald D, Yang Y, Zaw T, et al. EMPress enables tree-guided, interactive, and exploratory analyses of multi-omic data sets. mSystems. 2021;6:e01216-20.
Kosaka T, Toh H, Toyoda A. Complete genome sequence of a thermophilic hydrogenotrophic methanogen, Methanothermobacter sp. strain CaT2. Genome Announc. 2013;1:e00672-13.
Acknowledgements
This work was performed under the auspices of the Natural Sciences and Engineering Research Council (NSERC) of Canada Discovery Grants program (RMZ and SJH) and the Genome British Columbia Sector Innovation Program (SIP06; EAM, DGM, RMZ, SJH). The metaproteomics, metabolomics, and short-read metagenomics was performed on a project awarded to RMZ and SJH (https://doi.org/10.46936/lser.proj.2020.51366/60000180) from the Environmental Molecular Sciences Laboratory, a DOE Office of Science User Facility sponsored by the Biological and Environmental Research program under Contract No. DE-AC05-76RL01830. PSS was also supported in part by the NSERC CREATE Ecosystem Services, Commercialization Platforms and Entrepreneurship (ECOSCOPE) training program at the University of British Columbia. We thank Metro Vancouver staff for kindly providing the anaerobic digester sludge sample.
Author information
Authors and Affiliations
Contributions
EAM prepared samples for metagenomics, metabolomics, and metaproteomics, performed bioinformatics and data analysis, and helped to write to the manuscript. MS generated and ran the metabolic models, and helped write the manuscript. DGM operated the bioreactors, performed microcosm incubations, and collected samples. XL performed long-read metagenomics and helped with bioinformatics analysis. PSS assisted with bioinformatics analysis. HO performed protein extractions and prepared peptide samples for analysis. RY, DH, and EE performed metabolomics analysis and data processing. KA performed peptide analysis on LC-MS/MS. LMM prepared short-read DNA libraries and performed sequencing. ML helped with the proteomics workflow. SJH assisted with experimental design and conception, as well as manuscript writing. RMZ helped with experimental conception and design, data analysis, metabolic reconstruction and modeling, and manuscript writing. All authors contributed to the revision of the manuscript.
Corresponding author
Ethics declarations
Competing interests
With the exception of SJH, the authors declare no competing interests associated with this work. SJH is a co-founder of Koonkie Inc., a bioinformatics consulting company that designs and provides scalable algorithmic and data analytics solutions in the cloud. Koonkie Inc. was not involved in any aspect of this research.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
McDaniel, E.A., Scarborough, M., Mulat, D.G. et al. Diverse electron carriers drive syntrophic interactions in an enriched anaerobic acetate-oxidizing consortium. ISME J 17, 2326–2339 (2023). https://doi.org/10.1038/s41396-023-01542-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41396-023-01542-6