Abstract
Despite important ecological roles posited for virocells (i.e., cells infected with viruses), studying individual cells in situ is technically challenging. We introduce here a novel correlative microscopic approach to study the ecophysiology of virocells. By conducting concerted virusFISH, 16S rRNA FISH, and scanning electron microscopy interrogations of uncultivated archaea, we linked morphologies of various altiarchaeal cells to corresponding phylogenetic signals and indigenous virus infections. While uninfected cells exhibited moderate separation between fluorescence signals of ribosomes and DNA, virocells displayed complete cellular segregation of chromosomal DNA from viral DNA, the latter co-localizing with host ribosome signals. A similar spatial separation was observed in dividing cells, with viral signals congregating near ribosomes at the septum. These observations suggest that replication of these uncultivated viruses occurs alongside host ribosomes, which are used to generate the required proteins for virion assembly. Heavily infected cells sometimes displayed virus-like particles attached to their surface, which agree with virus structures in cells observed via transmission electron microscopy. Consequently, this approach is the first to link genomes of uncultivated viruses to their respective structures and host cells. Our findings shed new light on the complex ecophysiology of archaeal virocells in deep subsurface biofilms and provide a solid framework for future in situ studies of virocells.
Similar content being viewed by others
Most in situ studies on environmental prokaryotic lytic viruses focus on free virions or their genomes [1]. Recently, the scientific community has begun to investigate ecological roles of the virocells, which are cells that have fallen victim to viral infection and are subject to metabolic conversion [2]. Host molecular machinery is typically reprogrammed rapidly to facilitate virus replication [3], and can even result in distinct cellular compartmentalization for reproduction in the case of highly evolved bacterial jumbo phages [4]. Viruses that infect archaea often possess distinct virion structures, biochemical properties, egress mechanisms, and coordinated virocell takeover motifs (as reviewed in [5,6,7]). As all of these reports have arisen from isolated archaeal cultures, the scientific community lacks an understanding of the structure and organization of virocells of the uncultivated majority [1].
Recently, advances in fluorescence in situ hybridization (FISH) enabled researchers to detect an uncultivated virus via tagging the viral genome [8, 9] and Schaible et al. to correlated FISH imaging with scanning electron microscopy (SEM) for identification of specific prokaryotes using 16S rRNA tagging [10]. However, the combination of virusFISH (i.e., virus-targeted direct-geneFISH) with SEM has been proposed [11] but not been established, as virusFISH protocols necessitate 300 base pairs double-stranded (ds) probes to enter target cells and thus a harsh temperature treatment of cells, which results in disintegration of the cellular ultrastructure (Supplementary Fig. S1). Here we report a novel correlative microscopic approach to study the ecophysiology of indigenous, uncultivated archaeal virocells by conducting concerted virusFISH, 16S rRNA-based FISH, and SEM analyses. Applied to uncultivated archaea from the deep biosphere, we linked morphologies of various Candidatus Altiarchaeum hamiconexum cells to corresponding nucleic acid signals.
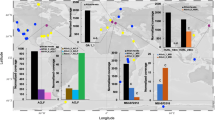
Carbon fixing members of the uncultivated archaeal genus Ca. Altiarchaeum dominate deep subsurface ecosystems worldwide [12] with uncultivated lytic ds-DNA viruses infecting these archaea and causing profound consequences on carbon cycling in the deep biosphere [8]. The genome of the ds-DNA virus primarily targeting Ca. A. hamiconexum and tagged via virusFISH, however, has not been predicted to encode any viral hallmark genes in a previous study [8], which is not surprising since most archaeal viruses encode for novel or previously unknown proteins [5]. By modeling protein tertiary structure from predicted viral genes with AlphaFold [13] and correlating results to annotations of function, we identified a novel auxiliary metabolic gene (i.e., Ni-Fe hydrogenase), a gene encoding a capsid protein, and other virus-associated proteins encoded in the circular viral genome (Supplementary Fig. S2, Supplementary Table S1). Given this in silico evidence confirming the viral nature of the tagged genome, we examined the ultrastructure of Ca. A. hamiconexum virocells using a novel correlative microscopic approach.
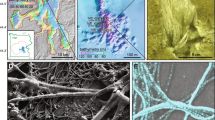
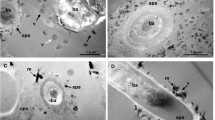
Conserving cellular ultrastructure is particularly challenging when preparing samples for (virus)FISH and subsequent SEM. A distinct characteristic of Ca. A. hamiconexum are their extracellular hami structures, which are extracellular cell surface appendages with a basal barbwire structure and nano-grappling terminal hooks to interconnect cells [14]. By carefully balancing fixation reagent concentrations with critical point drying (for details please see Supplementary Methods, Supplementary Fig. S1 illustrates insufficient sample preservation for comparison), we successfully preserved the hami, and thus ultrastructure of Ca. A. hamiconexum cells throughout virusFISH preparation (Fig. 1). Using gridded coverslips to ensure proper correlation of fluorescence and SEM signals (Fig. 1a, Supplementary Fig. S3), spatial separation between ribosomes and chromosomal DNA was apparent in most uninfected cells. Although this phenomenon has previously been observed for archaea larger than altiarchaeal cells (i.e., Asgardarchaeota) [15], we observe a comparable segregation in ribocells (cells showing no infection) of small Altiarchaea of the DPANN superphylum (700–800 nm diameter). A similar segregation was observed in cells undergoing division, with DNA localizing about the outer poles of each of the daughter cells. At the same time, our approach enabled the infrequent detection of small DNA-filled vesicles (diameter: 250–300 nm, n = 3) attached to Ca. A. hamiconexum ribocells (Fig. 1b). We hypothesize that these vesicles play an important role in inter-kingdom lateral gene transfer, which has recently been reported in Ca. Altiarchaea [16].
Blue fluorescence signal corresponds to DAPI, green to the SMARCH714 probe [20] labeling the 16S rRNA of Ca. A. hamiconexum (Atto488) and red the virusFISH probes [8] labeling the viral genome (Alexa594); a Overview of single infected cell between multiple non-infected cells demonstrating the successful correlation of the two imaging techniques due to sample preservation. b Uninfected single cell. Arrow points at a putative vesicle. c Uninfected dividing cells. d Infected swollen cell. e Dividing infected cells. f Single infected cell. g Overlay of the fluorescence images of f. Arrows point at spherical viral signals. Scale bars: 1 µm (a), 500 nm (b–g). We provide a color-blind friendly image of this figure as Supplementary Fig. S4; single channel images are provided as Supplementary Figs. S5–S8.
Altiarchaeal cells infected with uncultivated viruses (infection rate 5% [11]) were 10–20% larger and slightly more spherical than their ribocell counterparts (Fig. 2b; 30 infected and 145 non-infected cells; Wilcoxon test, two-sided, p value < 0.001; Supplementary Table S2, Supplementary Figs. S9, S10). Virocell enlargement has been reported previously in pure culture, with infected Sulfolobus islandicus cells increasing their size by several orders of magnitude [6]. In dividing Ca. A. hamiconexum cells, virus signals accumulated along the cell division septum, indicating that viral reproduction occurs in the cytosol between the two daughter cells (Fig. 1d, e). This spatial separation of chromosomal DNA, ribosomes, and virion synthesis was also observed in individual cocci (Fig. 1f, g). Upon comparing these observations with ultrathin sections of Ca. A. hamiconexum cells, we confirmed a distinct segregation of viral reproduction activities from other cytosolic spaces (Fig. 2c, Supplementary Fig. S12). An accumulation of virus signals arranged in sphere-like intracellular structures observed via fluorescence microscopy suggested an organized packing of virions (Figs. 1g and 2a). However, virocells did not exhibit these sphere-like structures in most instances, but rather a dense agglomeration of virus signals (e.g., virus signals in Fig. 2a). These observations most likely reflect different stages of the viral reproduction cycle, yet all of these are spatially separated from host chromosomes.
a Overlays of fluorescence and scanning electron micrographs (sampled in 2022). Arrows point at putative extracellular VLPs. Blue fluorescence signal corresponds to DAPI, green to the SMARCH714 probe [20] labeling the 16S rRNA of Ca. A. hamiconexum (Atto488) and red the virusFISH probes [8] labeling the viral genome (Alexa594). b Boxplot indicating the measured size of the VLPs measured in SEM and TEM (average diameter of VLPs in TEM 53 nm (n = 56) vs. SEM 65 nm (n = 71) t-test p value = 2.7·10−11) The dashed boxplot within the SEM column represents the hypothetical data after subtraction of the 4 nm Pt/Pd layer (significance of t-test above boxplots). In comparison to the measured archaeal cell size (30 infected and 145 non-infected cells; Wilcoxon test, p value < 0.001) (significance of Wilcoxon test above boxplots). c TEM images (sampled in 2018 [8]). Arrows point at putative VLPs within Ca. A. hamiconexum cells. Scale bars 500 nm.
Strong virus signals not segregating into individual spheres often co-occurred with changes to the host cell’s surface structure. These alterations manifested as small virus-like particles (VLPs) attached to the surfaces of Ca. A. hamiconexum cells (Fig. 2a). While the virion architecture of these uncultivated viruses remains unknown, these structures agree with transmission electron microscopy (TEM) images herein and elsewhere [8, 17]. We compared the size of intracellular putative VLPs of TEM thin sections and the extracellular VLPs identified via correlative microscopy finding a significant difference in size [average diameter of VLPs in TEM 53 nm (n = 56) vs. SEM 65 nm (n = 71) t-test p value = 2.7·10−11; Supplementary Table S3, Fig. 2, Supplementary Figs. S11, S12]. Although the difference in size might be due to the application of different imaging techniques or the different states of virus maturation (e.g., intracellular and extracellular, presence of the capsids), the sizes of the VLPs are in the expected range of archaeal viruses [18]. However, the correlative microscopic method described here links metagenomic data to the structure of uncultivated viruses, and thus enabled elucidating morphological traits of viruses whose existence is known only from sequence data. We suggest that this novel approach could be easily adapted to other sample matrices than subsurface biofilms, which per se represent a challenge due to the complexity of the biofilm matrix.
Microbes of the deep biosphere remain some of the most enigmatic biological entities on Earth due to limitation in sampling opportunities of the deep subsurface. While "omics" approaches are mostly used to decipher microbial traits in the deep subsurface [19], advances at single-cell level to understand ecophysiology of these organisms and associated viral infections remain the exception. Our novel protocol enabling the ability to spot rare infection events of individual cells presents a major step forward in understanding the little characterized deep biosphere at individual virocell level and demonstrates specific morphologies from vesicles to size alteration of uncultivated archaea. The spatial-functional distribution within virocells reveals a high degree of sub-cellular organization in deep-branching archaea, which might be required for the viruses to replicate and warrants future investigation necessitating sophisticated method development tied to high-resolution imaging, like TEM [9]. While the molecular mechanisms underlying such virus replication in deep-branching archaea remain unknown, we suggest that sub-cellular organizations in ribocells and virocells play an important role in microbial physiology also outside well-characterized model systems and might encompass the entire tree of archaea.
References
Turzynski V, Monsees I, Moraru C, Probst AJ. Imaging techniques for detecting prokaryotic viruses in environmental samples. Viruses. 2021;13:2126.
Howard-Varona C, Lindback MM, Bastien GE, Solonenko N, Zayed AA, Jang H, et al. Phage-specific metabolic reprogramming of virocells. ISME J. 2020;14:881–95.
Forterre P. The virocell concept and environmental microbiology. ISME J. 2013;7:233–6.
Chaikeeratisak V, Nguyen K, Khanna K, Brilot AF, Erb ML, Coker JKC, et al. Assembly of a nucleus-like structure during viral replication in bacteria. Science. 2017;355:194–7.
Prangishvili D, Bamford DH, Forterre P, Iranzo J, Koonin EV, Krupovic M. The enigmatic archaeal virosphere. Nat Rev Microbiol. 2017;15:724–39.
Liu J, Cvirkaite-Krupovic V, Baquero DP, Yang Y, Zhang Q, Shen Y, et al. Virus-induced cell gigantism and asymmetric cell division in archaea. Proc Natl Acad Sci USA 2021;118:e2022578118.
Baquero DP, Liu J, Prangishvili D. Egress of archaeal viruses. Cell Microbiol. 2021;23:e13394.
Rahlff J, Turzynski V, Esser SP, Monsees I, Bornemann TLV, Figueroa-Gonzalez PA, et al. Lytic archaeal viruses infect abundant primary producers in Earth’s crust. Nat Commun. 2021;12:4642.
Jahn MT, Lachnit T, Markert SM, Stigloher C, Pita L, Ribes M, et al. Lifestyle of sponge symbiont phages by host prediction and correlative microscopy. ISME J. 2021;15:2001–11.
Schaible GA, Kohtz AJ, Cliff J, Hatzenpichler R. Correlative SIP-FISH-Raman-SEM-NanoSIMS links identity, morphology, biochemistry, and physiology of environmental microbes. ISME Commun. 2022;2:52.
Turzynski V, Griesdorn L, Moraru C, Soares AR, Simon SA, Stach TL, et al. Virus-host dynamics in archaeal groundwater biofilms and the associated bacterial community composition. Viruses. 2023;15:910.
Probst AJ, Weinmaier T, Raymann K, Perras A, Emerson JB, Rattei T, et al. Biology of a widespread uncultivated archaeon that contributes to carbon fixation in the subsurface. Nat Commun. 2014;5:5497.
Jumper J, Evans R, Pritzel A, Green T, Figurnov M, Ronneberger O, et al. Highly accurate protein structure prediction with AlphaFold. Nature. 2021;596:583–9.
Moissl C, Rachel R, Briegel A, Engelhardt H, Huber R. The unique structure of archaeal ‘hami’, highly complex cell appendages with nano-grappling hooks. Mol Microbiol. 2005;56:361–70.
Avci B, Brandt J, Nachmias D, Elia N, Albertsen M, Ettema T, et al. Spatial separation of ribosomes and DNA in Asgard archaeal cells. ISME J. 2021;16:606–10.
Bornemann TLV, Adam PS, Turzynski V, Schreiber U, Figueroa-Gonzalez PA, Rahlff J, et al. Genetic diversity in terrestrial subsurface ecosystems impacted by geological degassing. Nat Commun. 2022;13:284.
Probst AJ, Moissl-Eichinger C. “Altiarchaeales”: uncultivated archaea from the subsurface. Life. 2015;5:1381–95.
Prangishvili D, Garrett RA, Koonin EV. Evolutionary genomics of archaeal viruses: unique viral genomes in the third domain of life. Virus Res. 2006;117:52–67.
Probst AJ, Ladd B, Jarett JK, Geller-McGrath DE, Sieber CMK, Emerson JB, et al. Differential depth distribution of microbial function and putative symbionts through sediment-hosted aquifers in the deep terrestrial subsurface. Nat Microbiol. 2018;3:328–36.
Moissl C, Rudolph C, Rachel R, Koch M, Huber R. In situ growth of the novel SM1 euryarchaeon from a string-of-pearls-like microbial community in its cold biotope, its physical separation and insights into its structure and physiology. Arch Microbiol. 2003;180:211–7.
Acknowledgements
The work carried out herein was funded by the German Academic Scholarship Foundation (given to IB). AJP acknowledges support from the Ministry of Culture and Science of North Rhine-Westphalia (Nachwuchsgruppe “Dr. Alexander Probst”) and German Research Foundation (DFG grant PR1603/2-1). This project has received funding from the European Research Council (ERC) under the European Union’s Horizon 2020 research and innovation program (grant agreement No. 863664). We thank Sabrina Eisfeld, Ines Pothmann, Maximiliane Ackers, and Agathe Materla for lab management and technical assistance. We thank the team at the Imaging Center Essen and Jennifer Grünert for technical assistance, training on the microscopes, and constructive discussions leading to superior sample preparation protocols and the colleagues of the former Group for Aquatic Microbial Ecology at the University of Duisburg-Essen for initial support and discussion.
Funding
Open Access funding enabled and organized by Projekt DEAL.
Author information
Authors and Affiliations
Contributions
IB performed all imaging; IB AK MH established the SEM sample preparation protocol; IB VT JR and CM optimized the protocol for the gridded coverslip dish; AS and IB performed data analysis in R; CM VT IB and AJP discussed the interpretation of the FM results; IB and AJP discussed SEM images and performed interpretation of correlative microscopy; SPE VT IB and JR performed sampling; SPE critically reviewed the Figures with IB; PN PM PW performed viral gene annotation with input from SPE; IB AK AJP conceptualized the study; IB and AJP wrote the paper with input and revisions from all co-authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Banas, I., Esser, S.P., Turzynski, V. et al. Spatio-functional organization in virocells of small uncultivated archaea from the deep biosphere. ISME J 17, 1789–1792 (2023). https://doi.org/10.1038/s41396-023-01474-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41396-023-01474-1