Abstract
The phylogenetic resolution at which microorganisms display geographic endemism, the rates at which they disperse at global scales, and the role of humans on global microbial dispersal are largely unknown. Answering these questions is necessary for interpreting microbial biogeography, ecology, and macroevolution and for predicting the spread of emerging pathogenic strains. To resolve these questions, I analyzed the geographic and evolutionary relationships between 36,795 bacterial and archaeal (“prokaryotic”) genomes from ∼7000 locations around the world. I find clear signs of continental-scale endemism, including strong correlations between phylogenetic divergence and geographic distance. However, the phylogenetic scale at which endemism generally occurs is extremely small, and most “species” (defined by an average nucleotide identity ≥ 95%) and even closely related strains (average nucleotide identity ≥ 99.9%) are globally distributed. Human-associated lineages display faster dispersal rates than other terrestrial lineages; the average net distance between any two human-associated cell lineages diverging 50 years ago is roughly 580 km. These results suggest that many previously reported global-scale microbial biogeographical patterns are likely the result of recent or current environmental filtering rather than geographic endemism. For human-associated lineages, estimated transition rates between Europe and North America are particularly high, and much higher than for non-human associated terrestrial lineages, highlighting the role that human movement plays in global microbial dispersal. Dispersal was slowest for hot spring- and terrestrial subsurface-associated lineages, indicating that these environments may act as “isolated islands” of microbial evolution.
Similar content being viewed by others
Introduction
Endemism caused by geographic isolation, i.e., the restriction of a taxon to a specific region of the world due to dispersal limitation, can be a major driver of biogeographic patterns and evolutionary dynamics and has long been a topic of great interest [1,2,3]. Depending on the relative rates of evolutionary divergence and geographic dispersal, endemism by geographic isolation can be observed at very coarse taxonomic levels, the restriction of the entire mammalian family Macropodidae (kangaroos, wallabies and others) to the Australasian region [4] being a striking example. In bacteria and archaea (henceforth “prokaryotes”), whose dispersal is generally less restricted than in larger organisms, endemism is not observed at these high taxonomic levels, but it is unknown at what phylogenetic resolution prokaryotes do display endemism, or how fast they disperse at global scales relative to their evolutionary divergence. For example, it is intensely debated whether prokaryotic taxa are globally distributed even at the species level [5,6,7,8,9,10]. The phylogenetic resolution of prokaryotic endemism and rates of global prokaryotic dispersal have far reaching implications for prokaryotic biogeography, ecology, and macroevolution. A global distribution of prokaryotic species could limit the potential for prokaryotic speciation and thus constrain global prokaryotic diversity [5]. Indeed, if global dispersal is sufficiently fast, prokaryotic speciation would be bound to effectively occur “sympatrically”, i.e., without geographic barriers to gene flow [11]. A global distribution of prokaryotic species may also reduce the risk of extinction in the face of local or global environmental change, since at any given time it is likely that some areas in the world remain suitable for a species to persist and fast colonization of these areas would act as a rescue mechanism [12]. Rates of prokaryotic dispersal also dictate how fast genetic material, such as genes involved in antibiotic resistance, pathogenicity or bioremediation, can spread across space. Lastly, knowing the rates of global prokaryotic dispersal is necessary for a proper interpretation of biogeographic patterns, such as species-area or distance-similarity relationships [13, 14].
Surveys of prokaryotic 16S rRNA gene sequences suggest that at the typical considered resolution (i.e., when clustered at 97%) most prokaryotic taxa are globally distributed [15,16,17,18,19]. This means that over the time scales needed for the 16S rRNA gene to diverge by 3% (∼30–150 Myr [20, 21]), a prokaryotic lineage is likely to have dispersed around the globe. These findings only provide a coarse one-sided bound on the phylogenetic resolution at which prokaryotes display endemism, and do not clarify the actual rates at which prokaryotes disperse at global scales. Indeed, for many taxa global dispersal appears to be much faster than the rate at which the 16S rRNA gene and other single marker genes can measurably diverge, thus necessitating the consideration of multiple genes or whole genomes [22, 23]. A notable exception are extremophilic (e.g., hot spring-associated) taxa, where geographic endemism at the species or strain level has been observed in specific clades using marker genes or genomics [6, 24,25,26,27,28,29,30,31], although even in these cases the phylogenetic resolution of endemism and rates of dispersal are poorly understood.
Here, to determine the precise phylogenetic resolution at which prokaryotes display geographic endemism at global scales and to estimate actual rates of global prokaryotic dispersal in the context of a process-based model, I present a phylogeographic analysis of 36,795 high-quality georeferenced prokaryotic genomes from ∼7000 locations around the world. I distinguished between human-associated genomes, other (i.e., non-human-associated) terrestrial genomes, and marine genomes, since substantial differences are expected between these environments and to facilitate comparison with previous environment-specific studies. Among non-human-associated terrestrial genomes, I also performed focused analyses of genomes found in hot springs, lakes (henceforth “terrestrial lakes”) or the subsurface (henceforth “terrestrial subsurface”), as global microbial dispersal might be particularly slow in these environments.
Results and discussion
A collection of thousands of georeferenced genomes
To examine the global distribution patterns of prokaryotic clades at phylogenetic resolutions beyond those permitted by single-marker-gene sequences, I analyzed 36,795 georeferenced quality-filtered whole or draft genomes (≥ 90% complete, ≤ 1% contamination) retrieved from NCBI GenBank [32]. The majority of genomes were >99% complete with <0.2% contamination (Supplementary Figs. S1 and S2). The genomes originated from a diverse range of environments across 183 countries, such as the surface and deep ocean, lakes, soil, human and other animal guts, hot springs, sediments, caves, wells, hydrothermal vents, glaciers, aquifers and rivers (world maps in Supplementary Fig. S3, summaries in Supplementary Table S1). The evolutionary divergence between genomes was, wherever meaningful, measured in two alternative ways: First, divergence between genomes was measured in terms of average nucleotide difference (AND) across their shared genes, which permits a much higher resolution for delineating closely related strains (compared to single genes). Note that here AND is calculated as 100%-ANI, where ANI is the average nucleotide identity, a widely used measure of microbial relatedness [33,34,35]. An AND threshold of ∼5%, in particular, is currently a commonly used measure for delineating prokaryotic species [33,34,35,36,37], thus facilitating the interpretation of biogeographic patterns and comparisons to previous studies. Second, divergence between genomes was measured in terms of absolute time (years) based on time-calibrated phylogenies, which were constructed from 120 bacterial and 122 archaeal largely universal single-copy proteins [38, 39] and dated based on multiple constraints (details in Methods and Supplementary S.1.3).
Geographic location exhibits a clear phylogenetic signal
Phylogenetic conservatism in numerical traits (i.e., closely related lineages exhibit more similar traits) is often confirmed based on a positive correlation between pairwise trait differences and evolutionary divergences, while correcting for the non-independence of related taxa [40]. By analogy, here I considered the geographic location of a lineage as an evolving trait, measured the “trait difference” between two genomes in terms of their geographic (great-circle) distance, and examined whether this distance correlates with phylogenetic divergence [41]. I found that the AND between genomes correlated strongly and significantly with their geographic distance (Fig. 1), regardless of environment (P < 0.05 in all cases, overview in Supplementary Table S2, methods details in Supplementary S.1.5). Spearman rank correlations ranged from ρ = 0.266 for marine genomes up to ρ = 0.655 for hot spring genomes. Significant correlations were also observed in all environments between temporal divergence and geographic distance (ρ > 0.1, P < 0.05). Note that quantitative comparisons of these correlations between environments should be avoided, since correlations also depend on the phylogenetic coverage available (e.g., whether mostly distantly related organisms are included in the data or not). A correlation between phylogenetic and geographic distance is in principle not surprising, since dispersal inevitably occurs at a non-infinite rate. What these significant correlations show, however, is that dispersal rates are sufficiently slow to cause a detectable phylogenomic signal in a clade’s geographic location, thus allowing estimation of dispersal rates through comparative genomics (see below).
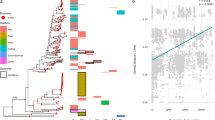
Geographic distances (vertical axes) and average nucleotide differences (AND, horizontal axes) between genome pairs (one dot per genome pair), restricted to human-associated (A), non-human-associated terrestrial (B), marine (C), hot-spring-associated (D), terrestrial lake-associated (E) and terrestrial subsurface-associated (F) prokaryotes. Curves show the expected geographic distance as a function of AND, based on a fitted Spherical Brownian Motion diffusion model (see Methods for details). The range of ANDs shown is restricted to closely related genome pairs for clarity. Note the different AND-axis range for each environment.
Continental endemism
To determine the phylogenetic resolution at which prokaryotes display geographic endemism over large scales, I calculated the fraction of genome pairs that were found on opposite hemispheres depending on the maximum allowed AND (Fig. 2). For any given AND threshold, this fraction corresponds to the probability that two genomes with AND at or below this threshold would be located on opposite hemispheres (henceforth denoted α). Two genomes were considered on “opposite hemispheres” if one could split Earth into two hemispheres, with one genome being in the center of the first hemisphere and the other genome being anywhere within the other hemisphere; this criterion is equivalent to their distance being >1/4 of Earth’s circumference (for alternative distance thresholds see Supplementary Fig. S4). If below a certain AND threshold clades are restricted to areas much smaller than one hemisphere, one would expect α to be close to zero below that threshold. Further, I compared α to the expected probability under the null model of infinitely fast dispersal (henceforth denoted αo), i.e., where relatedness has no influence on geographic distance, while accounting for the number of genomes sampled from each location (see Methods for details).
Estimated probability (α, thick gray curves) that two randomly chosen genomes would be located on opposite hemispheres (i.e., at a distance >1/4 of Earth’s circumference), as a function of their maximum allowed average nucleotide difference (AND, %), for (A) human-associated, (B) non-human-associated terrestrial, (C) marine, (D) hot spring, (E) terrestrial lake and (F) terrestrial subsurface genomes. In other words, at any given AND value on the horizontal axis, the curve’s ordinate specifies what fraction of genome pairs, with AND at or below that value, is located on different hemispheres. Note that in (F) no two genomes analyzed in this study with AND ≤ 5% were found in opposite hemispheres. In each figure, the thin horizontal line shows the expected probability under the null model of infinitely fast dispersal (i.e., choosing genome pairs at random regardless of relatedness). Observe that genome pairs with very small ANDs are unlikely to be found on separate hemispheres, although strong differences exist between environments. For the same analyses but using random genome subsets of equal size for each environment, see Supplementary Fig. S7. For a similar analysis of continental endemism see Supplementary Figs. S6 and S8. For analyses showing α for alternative distance thresholds (i.e., other than 1/4 of Earth’s circumference) see Supplementary Fig. S4.
I found that two randomly chosen genomes generally had a non-negligible probability α of being on opposite hemispheres, except for very small AND thresholds or for hot spring- and terrestrial subsurface-associated genomes. For example, for human-associated genomes and an AND threshold of 0.1%, the estimated probability α is about 0.2, which means that two randomly chosen genomes with AND ≤ 0.1% have a ∼20% chance of being on opposite hemispheres (Fig. 2A). Considering that for any given prokaryotic cell there likely exist millions or more cells within this relatedness radius, the probability that at least one of them will be located on the opposite hemisphere is thus nearly 1. If we define “globally distributed” clades to be those clades that have members in opposite hemispheres, then this implies that even if human-associated clades were delineated at an AND cutoff as low as 0.1%, nearly all of them would be globally distributed (Fig. 3 and Supplementary Fig. S5). This conclusion is consistent with previous studies that showed a global distribution for specific human-associated bacterial species [22, 42]. Similar conclusions can be reached for other terrestrial non-human-associated as well as for marine clades (Fig. 2B, C). Reciprocally, a non-global distribution corresponds to the situation where all (or nearly all) members of a clade are located on the same hemisphere, which corresponds to the situation where α ≈ 0. As can be seen in Fig. 2, the AND threshold at which α is nearly zero depends on the environment: Marine clades clearly exhibit the lowest AND threshold (∼0.0003% AND), followed by human-associated clades (∼0.005% AND), non-human-associated terrestrial (∼0.01% AND), and terrestrial lake-associated (∼0.4% AND) clades. Hot spring and terrestrial subsurface clades deviate strikingly from the terrestrial average, exhibiting an AND threshold above 5%. These differences between environments might be due to a variety of reasons, including differences in dispersal mechanisms, and differences in the availability of suitable environments (e.g., hot springs are more geographically restricted than the ocean).
Geographic locations of prokaryotic genomes (circles) in each of the two largest human-associated species-level genome bins (SGBs, i.e., genome clusters with an average nucleotide difference cutoff of 5%) analyzed in this study (A) Escherichia coli, (B) Staphylococcus aureus). Straight lines connect genomes with average nucleotide difference (AND) ≤ 0.1% (i.e., average nucleotide identity ≥ 99.9%). Representative strains and the number of genomes (n) in each SGB are noted in the sub-titles. See Supplementary Fig. S5 for additional SGBs.
The phylogenetic scale at which most examined marine and terrestrial clades are restricted to one hemisphere is extremely small, and certainly smaller than could possibly be resolved with 16S rRNA gene sequences. Indeed, a single mutation in the ∼1550 bp-long 16S rRNA gene is expected to take 0.7–3 Myr [20, 21], which based on a genome-wide nucleotide substitution rate of 0.7–30% per Myr [20, 43] corresponds to an AND of at least 0.49%. These findings explain previous observations that marine taxa, when clustered at 97% similarity in the 16S rRNA gene (generally a much coarser threshold than 0.1% AND), are essentially globally distributed [16, 18]. For human-associated clades and at an AND threshold around 5%, commonly used to operationally define prokaryotic species [33,34,35], α is nearly equal to αo (Fig. 2A); this suggests that the global distribution of human-associated prokaryotic species is nearly indistinguishable from the extreme scenario of infinitely fast dispersal.
Analogous results were also obtained for continental endemism, i.e., examining the probability of genome pairs being located on different continents (Supplementary Fig. S6). Similar results were also obtained when the same number of genomes was analyzed from each environment, suggesting that the above conclusions are not overly sensitive to sampling effort (Supplementary Figs. S7 and S8). Note that the estimated probability α constitutes an average over all sampled clades. It is possible that some broad clades (e.g., species or higher level) may be geographically restricted to a single continent, although the data suggest that such cases are rare and mostly found in extreme environments such as hot springs and the subsurface. It is also in principle possible that many non-sampled clades deviate substantially from these patterns, for example if current sequencing efforts are somehow biased toward fast dispersers.
Rates of global diffusive dispersal
The above results show that most examined prokaryotic clades only display geographic endemism at very fine phylogenetic resolutions, but they do not clarify the actual rates at which prokaryotes disperse at global scales. To determine the rates of dispersal in terms of physically interpretable quantities, an explicit process-based model for continuous dispersal across space is needed. While a variety of phylogeographic process-based models are used in the literature, most are based on a planar approximation of space and are thus unsuitable for describing dispersal at global scales, where Earth’s spherical geometry becomes important [41, 44]. Here I used phylogeographic Spherical Brownian Motion (SBM) models, which describe the dispersal of individual lineages over time as a diffusion-like process, governed by a single diffusion coefficient D (henceforth “diffusivity”), while accounting for Earth’s spherical geometry [41, 45]. Under diffusive dispersal, cells are allowed to move in any random direction and may over time partly or fully reverse previous movements; hence, such dispersal cannot be described by a linear velocity. Intuitively, the diffusivity D determines the rate at which the expected squared distance traversed by a lineage over short times increases, measured in units km2 yr−1 [46]; note that the perhaps better known “infinitesimal variance” σ2 is equal to 2D. The diffusivity can be estimated from phylogeographic data by comparing the evolutionary distances between closely related genomes to their corresponding geographic distances, via maximum-likelihood and using “independent contrasts” to account for phylogenetic correlations between genomes [40, 41]. To account for geographic sampling biases, which could influence diffusivity estimates, an iterative correction approach was used based on parametric bootstrapping (see Methods for details), although in most cases the effects of such biases were only moderate.
The highest diffusivity was estimated for human-associated prokaryotes (D = 1211 km2 yr−1, overview in Supplementary Table S3). Based on this diffusivity, after 100 years a single cell lineage is expected to have traversed a net distance of ∼580 km, or otherwise said, two extant independently dispersing cells coalescing 50 years ago (thus having patristic distance 100 years) are expected to be on average ∼580 km apart (see Supplementary Fig. S9 for the expected distance over different time intervals, for mathematical formulas see [45, 46]). It should be kept in mind that dispersal rates of human-associated prokaryotic lineages have likely increased over time (e.g., due to increased global human traffic), and hence these diffusivity estimates should be seen as average rates over recent times. Also note that the dispersal of human-associated prokaryotic lineages need not necessarily resemble the dispersal of individual humans, since prokaryotes can be transferred between humans as well as between humans and objects. For other terrestrial and marine prokaryotes, estimated diffusivities were somewhat lower albeit within the same order of magnitude (D = 455 km2 yr−1 and D = 346 km2 yr−1, respectively). This means that the expected net distance traversed after 100 years is about 370 km for non-human-associated terrestrial prokaryotes and 325 km for marine prokaryotes. Note, however, that the estimation uncertainty for marine prokaryotes was high, with the 95% confidence interval of the diffusivity including values multiple orders of magnitude higher than the maximum-likelihood estimate (Supplementary Table S3). Much lower diffusivities were estimated with high confidence for terrestrial lakes and hot springs (D = 1.29 km2 yr−1 and D = 1.01 km2 yr−1, respectively). By far the lowest diffusivity was estimated for the terrestrial subsurface (D = 0.042 km2 yr−1). This means that even after 10,000 years a terrestrial subsurface lineage is expected to only traverse a distance of ∼35 km. A slower dispersal of hot spring-, lake- or subsurface-associated lineages is consistent with the stronger geographic endemism observed earlier (Fig. 2). For hot springs, in particular, a low diffusivity is also consistent with previous reports of endemism in thermophilic microorganisms [6, 24,25,26,27, 29, 31, 47].
Strong differences were also found between bacteria and archaea. For almost all considered environments, estimated bacterial diffusivities were two or more orders of magnitude greater than archaeal diffusivities (Supplementary Tables S4 and S5). For example, diffusivities for non-human-associated terrestrial bacteria and archaea were 633 km2 yr−1 and 0.113 km2 yr−1, respectively, while diffusivities for hot spring bacteria and archaea were 3.95 km2 yr−1 and 0.0348 km2 yr−1, respectively. This suggests that bacteria and archaea generally differ in their ability to disperse over long distances. One potential explanation might be that cyst formation, sporulation and the ability to survive in variable environments— all of which facilitate passive long-distance transport across barriers of inhospitable habitats—are more common in bacteria than in archaea [48]. A notable exception was the terrestrial subsurface, where archaea exhibited higher diffusivities than bacteria, although the 95% confidence intervals of the estimates nearly overlapped between the two clades. Also note that estimates for human-associated archaea were not possible due to a scarcity of genomes.
The above diffusivities represent averages across many clades. Dispersal rates could, however, vary substantially between clades even in the same environment, for example depending on a clade’s ecological niche. To investigate this potential variation, I also estimated the diffusivity individually for species-level genome bins (SGBs), i.e., clusters of genomes with an AND ≤ 5%, commonly assumed to correspond to prokaryotic species [33, 35,36,37]; this analysis was only performed for human-associated prokaryotes, where a sufficient number of large SGBs was available. I found that diffusivities can vary by 4–5 orders of magnitude between SGBs (Fig. 4A), although for most SGBs diffusivities were estimated in the range 10–103 km2 yr−1. To further examine variation between clades, and to specifically compare the dispersal rates of human-associated to other terrestrial prokaryotes while controlling for broad taxonomy, I also estimated the diffusivities for individual prokaryotic phyla (Fig. 4B). Similarly, to control for broad phenotype I also estimated the diffusivities for individual metabolic functional groups, i.e., organisms sharing specific metabolisms (as inferred using the tool FAPROTAX [49]; Fig. 4C). Within each phylum examined, and within each functional group examined, human-associated lineages had higher diffusivities than other terrestrial lineages. This is consistent with my earlier conclusion that human-associated prokaryotes tend to disperse faster than non-human-associated terrestrial prokaryotes.
A Diffusivities (vertical axis) estimated for individual human-associated SGBs (one point per SGB), plotted against the SGB’s size (number of genomes, horizontal axis). B Diffusivities estimated for individual prokaryotic phyla (circles: human-associated genomes, triangles: other terrestrial genomes). C Diffusivities estimated for individual metabolic phenotype groups, i.e., genomes estimated to be capable of performing specific metabolic functions (circles: human-associated genomes, triangles: other terrestrial genomes). In all figures, only shown are SGBs (A) or phyla (B) or phenotypes (C) for which the diffusivity could be estimated based on at least ten genome pairs, and for which the lower and upper bound of the 95% confidence interval differed by less than a factor of 5 (i.e., the upper bound is at most five times greater than the lower bound).
Transition rates between continents
A caveat of diffusion models is that they may not always be an appropriate description of long-range dispersal, for example driven by human intercontinental air traffic or tropospheric air flow [50]. To more accurately determine the rate at which prokaryotes disperse between continents, I fitted phylogeographic Markov chain (Mk) models [51], which describe the transitions between continents over time as a probabilistic process whose rates depend on the pair of continents considered (Fig. 5A, B). To avoid biases caused by sample size differences, the same number of genomes was used for each continent (randomly subsampled where needed). For human-associated lineages, the highest per-lineage transition rate was estimated between Europe and North America (∼5.5 Myr−1), followed by Europe and Africa and then Asia and Africa (Fig. 5A). For non-human-associated terrestrial lineages, transition rates are generally much lower than for human-associated lineages; for example, the maximum estimated per-lineage transition rate, between Asia and North America (0.026 Myr−1, Fig. 5B), is two orders of magnitude lower than for human-associated lineages. This suggests that human movement greatly accelerates the global circulation of human-associated prokaryotes, consistent with previous findings for certain human pathogens [42]. Oceania (i.e., Australia, New Zealand, and other nearby islands) generally exhibits the lowest transition rates to/from other continents, and this holds true for both human-associated and other terrestrial lineages, resembling common observations for larger organisms such as animals [4].
Probabilistic transition rates (per-lineage per time) between continents for (A) human-associated and (B) terrestrial non-human-associated lineages, estimated by fitting a continuous-time stochastic Markov model for discrete trait evolution to time-calibrated phylogenies built from the genomes. Larger and darker circles correspond to faster transition rates. Note the different scales in (A) and (B). Models were fitted using rarefied datasets, i.e., with the same number of genomes per continent.
Note that the estimated intercontinental transition rates, as well as the diffusivities estimated earlier, should be interpreted on a per-lineage basis, i.e., they describe the rate at which a single cell lineage disperses over time and over generations. These estimates do not specify the rate at which an entire clade with multiple members expands its geographical boundaries over time, since the latter depends on the size of the clade and the correlations between trajectories of its members. Most bacterial species likely comprise billions or more individual cells [19, 52]; if each of these cells is considered an independently dispersing lineage, it becomes clear that the expected time needed for at least one member to colonize a new continent is orders of magnitude lower than when considering any single lineage. For example, a hypothetical human-associated bacterial species comprising 108 independently dispersing members initially located entirely in Europe, is expected to disperse to North America (i.e., have at least one member arrive in North America) within less than a day.
Human influence on non-human-associated prokaryote dispersal
It is possible that the dispersal of non-human-associated prokaryotes may also be accelerated by human activity, for example through cargo ships or automobiles. In that case, the diffusivities estimated for non-human-associated prokaryotes—even if correct—may be much higher than their natural (“background”) diffusivities, i.e., what would be expected prior to the appearance of humans, thus complicating the interpretation of microbial biogeography over geological time scales. Such “hitchhiking” effects are expected to be particularly strong in coastal areas (where ship densities are highest) and in densely populated areas on land. To assess the extent of this effect, I re-estimated diffusivities using marine genomes sampled from non-coastal areas and using non-human-associated terrestrial genomes sampled from remote areas (Supplementary Fig. S3G, H). Non-coastal areas were defined as being at least 370 km away from any major coast, corresponding to the standard width of exclusive economic zones, within which maritime activity tends to be higher [53]. Remote terrestrial areas were defined as those where population density in 2020 was estimated to be <10 humans per land km2. While the diffusivities estimated for these restricted genome sets differed somewhat from the nonrestricted estimates, there was no strong evidence that dispersal in the former was substantially slower than in the latter. Indeed, for the non-human-associated terrestrial prokaryotes the restricted diffusivity estimate was only about 19% lower than the unrestricted estimate, and for marine prokaryotes the restricted diffusivity estimate was only about 21% lower than the unrestricted estimate. In both cases the 95%-confidence intervals of the restricted and unrestricted estimates overlapped (details in Supplementary Table S3). Hence, while human activity probably contributes to the diffusive dispersal of non-human-associated prokaryotes, its overall contribution seems to be relatively moderate when compared to the naturally occurring dispersal.
Conclusions
An old, heavily contested and yet persistent hypothesis in microbial biogeography is that “everything is everywhere” and that “the environment selects” [9]. Much uncertainty exists over the validity of this hypothesis, partly because different studies investigating the importance of dispersal limitation consider different phylogenetic/taxonomic resolutions or different time scales (reviewed in [13]). The present work clarifies the phylogenetic resolution and temporal scales at which prokaryotes display geographic endemism at global scales, and quantifies overall global prokaryotic dispersal rates in the context of an explicit dispersal model. On the one hand, I find that geographic endemism is sufficiently strong to be detectable via whole-genome comparisons. On the other hand, even at clearly sub-species resolutions (e.g., ≤1% AND), nearly all prokaryotic clades on Earth’s surface seem to be globally distributed, that is, are not confined to a specific continent or hemisphere (Figs. 2, S6, S4) consistent with previous 16S rRNA-based studies [14,15,16,17,18,19]. Note that the goal of this study was not to determine the actual mechanisms by which prokaryotes disperse, which can include ocean currents, subsurface fluids, wind [50] or movement of animal hosts, nor does it clarify if the few observed cases of substantial endemism (notably in hot springs and terrestrial subsurface) are entirely driven by dispersal limitation and not environmental filtering; answering these questions remains a separate major task. Similarly, the dispersal models fitted here (SBM and Mk) are inevitably simplifications of reality, and are by no means an attempt to fully capture microbial biogeographic patterns (this would necessitate the incorporation of a multitude of physical and ecological processes, accounting for environmental heterogeneities and dispersal barriers). As such, the estimated diffusivities and Mk transition rates between continents should be seen as “effective” measures of dispersal rates, averaged over many clades and underlying processes. It should also be kept in mind that in the case of serious model violations such effective measures may be less meaningful [54, 55].
The finding that most prokaryotic clades on Earth’s surface are globally distributed suggests that local disturbances or rapid climatic shifts are unlikely to cause an extinction of a large fraction of prokaryotic species, in contrast to larger organisms. Further, many global-scale biogeographical patterns previously observed at coarse phylogenetic resolutions (e.g., as permitted by the 16S rRNA gene; [13]) are probably driven by environmental filtering—either current or in the recent past—rather than geographic isolation. Note that this does not imply that dispersal limitation is irrelevant, especially at short (ecological) time scales, since that would be equivalent to dispersal being essentially infinitely fast (which, as shown here, is not the case). For example, a recently formed or altered lake may not yet exhibit all bacterial species that could in principle live there, because migration (e.g., from other similar lakes) and establishment inevitably take time, thus leading to short-term historical contingency effects [56]. From the above analyses it becomes clear that the time scales over which historical contingencies matter will differ between environments, and may in some cases be long enough to have evolutionary implications. Indeed, estimated diffusivities were particularly low for the terrestrial subsurface and hot springs, which seem to constitute “isolated islands” of prokaryotic evolution.
Methods details
Genomes, metadata, and phylogenies
Complete or draft genomic sequences were downloaded from GenBank [32] on January 14, 2021, based on the following criteria: Only genomes with a contig-N50 above 5000, and with the “excluded_from_refseq” entry either being empty or only containing the terms “derived from metagenome”, “missing tRNA genes”, “derived from environmental source”, “derived from single cell”, “unverified source organism”, “partial”, “genus undefined”, were downloaded. The original sample coordinates were extracted from the corresponding BioSample’s “latitude”, “longitude” and/or “lat_lon” fields or (for a small number of isolate genomes) from the literature. Only georeferenced genomes, i.e., with available sample coordinates, were kept. Protein-coding genes were predicted for each georeferenced genome using prodigal v2.6.3 [57]. The quality of each genome was assessed based on the presence of multiple single-copy marker genes using checkM v1.1.3 [58], with option “reduced_tree”. Only genomes with an estimated completeness ≥90% and a contamination level ≤1% were kept, thus yielding 36,795 high-quality georeferenced genomes. The taxonomic identities of genomes were taken from GenBank, based on each genome’s taxid. An overview of genome qualities and completeness is provided in Supplementary Fig. S1. An overview of taxonomic coverages (genomes per taxon) is shown in Supplementary Fig. S10. Genome accession numbers, coordinates and other metadata are provided in Supplementary File 1. Pairwise geographic distances between genomes were calculated in terms of the great-circle distance, assuming that the Earth is approximately a sphere with radius R⊕ = 6371 km [59]. The environment type of each genome was determined based on its geographic coordinates, based on metadata provided by GenBank and using the tool FAPROTAX [49] (see Supplementary S.1.1 for details). The metabolic phenotype of each genome was predicted using the tool FAPROTAX [49] (see Supplementary S.1.2 for details). The human population density (estimated humans per land km2 for the year 2020) at every genome’s sampling location was obtained from the Gridded Population of the World data provided by the Socioeconomic Data and Applications Center (SEDAC), version 4.11, grid resolution 2.5 min [60], using bilinear interpolation between grid points where necessary. The distance of every genome’s sampling location to the nearest major coastline (including major islands) was calculated based on the Natural Earth coastlines database, version 4.1.0, resolution 110 m, accessed November 22, 2020 (www.naturalearthdata.com). A time-calibrated phylogenetic tree of genomes was built separately for each domain (bacteria and archaea) using multiple domain-specific universal marker genes. Briefly, genes were identified and aligned using the GTDB-Tk workflow v1.3.0 [39], trees were built from the concatenated alignments using FastTree v2.1.11 [61], and trees were dated with PATHd8 v1.0 [62] using multiple timing constraints from the literature (details in Supplementary S.1.3). Note that while this approach reflects current standard practice [38, 63], prokaryotic timetrees may still exhibit errors, for example due to violated molecular clock assumptions and a scarcity in dating constraints.
Estimating diffusivity in terms of km2 yr−1
Based on the dated trees and the genome coordinates, I fitted an SBM model [45, 64] for diffusion-like geographic dispersal, using the R package castor v1.6.6 [41, 65]. SBM models are analogous to the widely used Brownian Motion models of continuous dispersal [66,67,68,69], with the difference that SBM models do not simply encode geographic locations as two independent (i.e., orthogonal) numeric coordinates but instead account for Earth’s spherical geometry [41]. An SBM model is defined by a single “diffusivity” parameter D, which is equal to half the infinitesimal variance of Brownian Motion (typically denoted σ2) [70]. To estimate environment-specific diffusivities, the archaeal and bacterial trees were pruned to the subset of genomes associated with a particular environment, prior to model fitting. Note that, strictly speaking, each of these pruned trees may include old ancestral nodes specialized to a different environment than their tips, however this is expected to have a negligible effect on the estimated diffusivity since castor constructs independent contrasts mostly from recently diverged tip pairs. For additional details on how D is estimated, and how geographic sampling biases are accounted for, see Supplementary S.1.4. For diffusivity estimates not accounting or accounting for geographic sampling biases see Supplementary Tables S6 and S3, respectively.
Calculating average nucleotide differences
Pairwise average nucleotide identities (ANIs) between genomes were calculated as follows: First, approximate ANIs between all genomes were calculated using mash v2.2 [71]. Next, for any genome pair with a mash-based ANI ≥ 85% (roughly 3.9% of genome pairs), I recomputed the ANI using fastANI v1.3 [36]. The reason for this two-step approach is that while fastANI is slightly more accurate than mash, it is orders of magnitude slower; hence, calculating all possible pairwise ANIs with fastANI would have been practically unfeasible (and in fact unnecessary, since my analyses focus largely on ANIs > 90%). See Supplementary Fig. S11 for a comparison of mash vs. fastANI, and Supplementary Fig. S12 for the distribution of ANIs. Only fastANI-based ANIs were considered in the subsequent analyses. Throughout this paper, AND is defined as 100% minus ANI, and expressed in %.
Species-level genome bins
Species-level genome bins (SGBs) were constructed by clustering genomes at an AND cutoff of 5% using a modification of the approach taken by Pasolli et al. [72], as follows. First, bifurcating trees were constructed based on pairwise ANDs and using the BIONJ∗ algorithm [73] implemented in the R package ape v5.4-1 (function bionjs) [74]. For computational efficiency, prior to clustering, genomes were split into smaller disjoint subsets of moderately to closely related genomes, based on an AND cutoff threshold of 15%. BIONJ∗ trees were rooted using the midpoint method [75]. Next, tips in the BIONJ∗ trees (corresponding to genomes) were grouped into SGBs based on a maximum pairwise distance of 5% AND, using the function collapse_tree_at_resolution in the R package castor [65].
Assessing continental endemism
To examine the phylogenetic resolution of continental endemism (Supplementary Fig. S6), I proceeded as follows. For every genome, I assigned a country code based on the genome’s geographic coordinates using the python package reverse_geocoder v1.5.1 and then converted the country code to the corresponding continent name using the python package pycountry_convert v0.7.2 [76]. For any given environment type (e.g., human-associated), and for any given AND threshold, I considered all genome pairs with an AND at or below the threshold and determined which fraction of those genome pairs (denoted α) was located on separate continents. To determine the expectation for this fraction under the null hypothesis of infinitely fast dispersal (denoted αo), I randomly chose 1,000,000 genome pairs from the same environment regardless of their relatedness, and calculated the fraction of such pairs located on separate continents. Because randomly chosen genomes retained their original geographic location and environment, this null model accounts for broad environmental constraints and for geographic sampling biases, i.e., the fact that the number of genomes sampled is not equal among continents. To examine the fraction of genome pairs located on opposite hemispheres (Fig. 2) I proceeded in a similar manner, with the difference that two genomes were considered to be on opposite hemispheres if their distance was greater than πR⊕/2, where R⊕ is Earth’s radius. The null model corresponding to infinitely fast dispersal was implemented by randomly choosing 1,000,000 genome pairs regardless of their relatedness, and calculating the fraction of such pairs located on separate hemispheres. To examine whether the conclusions from this analysis are sensitive to sample sizes and biased due to differences in the number of genomes from each environment, I repeat the analyses using rarefied data, i.e., using a random subset of 400 genomes per environment. The results are shown in Supplementary Figs. S6, S7 and S8, and confirm the main conclusions presented in the main article regarding the approximate AND thresholds at which endemism appears in each environment.
Estimating intercontinental transition rates (Mk modeling)
To estimate intercontinental transition rates of prokaryotic lineages (Fig. 5), I fitted a continuous-time Markov chain model of discrete trait evolution (“Mk” model) [51] via maximum-likelihood using the castor function fit_mk, treating each continent as a distinct state. To reduce sampling biases between continents, the same number of genomes was considered for every continent through random subsampling. Antarctica was omitted from the analyses because very few genomes were available from there. To reduce the risk of overfitting, the transition rate between any two continents was assumed to be symmetric, i.e., to only dependent on the two continents but not on the direction (thus, the fitted model had 21 free parameters). To reduce the risk of converging to a local non-global maximum of the likelihood function, fitting was repeated 100 times with random start parameters. Note that an Mk model was not fitted to hot spring, lake and subsurface-associated lineages because the available data did not sufficiently cover all continents. An Mk model was also not fitted to marine lineages, as a separation of ocean space into discrete continents is much less meaningful.
Data availability
All data are available as supplementary material and on public repositories (accession numbers in Supplementary File 1).
Code availability
All software used in this paper have been described in the Methods and are freely available online.
References
Kruckeberg AR, Rabinowitz D. Biological aspects of endemism in higher plants. Annu Rev Ecol Syst. 1985;16:447–79.
Ceballos G, Brown JH. Global patterns of mammalian diversity, endemism, and endangerment. Conserv Biol. 1995;9:559–68.
Mueller GM, Schmit JP, Leacock PR, Buyck B, Cifuentes J, Desjardin DE, et al. Global diversity and distribution of macrofungi. Biodivers Conserv. 2007;16:37–48.
Prideaux GJ, Warburton NM. An osteology-based appraisal of the phylogeny and evolution of kangaroos and wallabies (macropodidae: Marsupialia). Zool J Linn Soc. 2010;159:954–87.
Finlay BJ, Clarke KJ. Ubiquitous dispersal of microbial species. Nature. 1999;400:828.
Whitaker RJ, Grogan DW, Taylor JW. Geographic barriers isolate endemic populations of hyperthermophilic archaea. Science. 2003;301:976–978.
Whitfield J. Is everything everywhere? Science. 2005;310:960–61.
Boenigk J, Pfandl K, Garstecki T, Harms H, Novarino G, Chatzinotas A. Evidence for geographic isolation and signs of endemism within a protistan morphospecies. Appl Environ Microbiol. 2006;72:5159–64.
DeWit R, Bouvier T. ‘Everything is everywhere, but, the environment selects’; what did Baas Becking and Beijerinck really say? Environ Microbiol. 2006;8:755–8.
van der Gast CJ. Microbial biogeography: the end of the ubiquitous dispersal hypothesis? Environ Microbiol. 2015;17:544–6.
Whittaker KA, Rynearson TA. Evidence for environmental and ecological selection in a microbe with no geographic limits to gene flow. Proc Natl Acad Sci USA. 2017;114:2651–56.
Louca S, Shih PM, Pennell MW, Fischer WW, Parfrey LW, Doebeli M. Bacterial diversification through geological time. Nat Ecol Evol. 2018;2:1458–67.
Martiny JBH, Bohannan BJ, Brown JH, Colwell RK, Fuhrman JA, Green JL, et al. Microbial biogeography: putting microorganisms on the map. Nat Rev Microbiol. 2006;4:102–12.
Martiny JBH, Eisen JA, Penn K, Allison SD, Horner-Devine MC. Drivers of bacterial β-diversity depend on spatial scale. Proc Natl Acad Sci USA. 2011;108:7850–54.
Jungblut AD, Lovejoy C, Vincent WF. Global distribution of cyanobacterial ecotypes in the cold biosphere. ISME J. 2010;4:191–202.
Gibbons SM, Caporaso JG, Pirrung M, Field D, Knight R, Gilbert JA. Evidence for a persistent microbial seed bank throughout the global ocean. Proc Natl Acad Sci USA. 2013;110:4651–55.
Ramirez KS, Leff JW, Barberán A, Bates ST, Betley J, Crowther TW, et al. Biogeographic patterns in below-ground diversity in New York City’s Central Park are similar to those observed globally. Proc R Soc Lond B Biol Sci. 2014;281:20141988.
Gonnella G, Böhnke S, Indenbirken D, Garbe-Schönberg D, Seifert R, Mertens C, et al. Endemic hydrothermal vent species identified in the open ocean seed bank. Nat Microbiol. 2016;1:16086 EP.
Louca S, Mazel F, Doebeli M, Parfrey WL. A census-based estimate of Earth’s bacterial and archaeal diversity. PLoS Biol. 2019;17:e3000106.
Ochman H, Wilson A. Evolution in bacteria: evidence for a universal substitution rate in cellular genomes. J Mol Evol. 1987;26:74–86.
Kuo CH, Ochman H. Inferring clocks when lacking rocks: the variable rates of molecular evolution in bacteria. Biol Direct. 2009;4:35–35.
Roberts MS, Cohan FM. Recombination and migration rates in natural populations of Bacillus subtilis and Bacillus mojavensis. Evolution. 1995;49:1081–94.
van Gremberghe I, Leliaert F, Mergeay J, Vanormelingen P, Van der Gucht K, Debeer AE, et al. Lack of phylogeographic structure in the freshwater cyanobacterium Microcystis aeruginosa suggests global dispersal. PLoS ONE. 2011;6:e19561.
Papke RT, Ramsing NB, Bateson MM, Ward DM. Geographical isolation in hot spring cyanobacteria. Environ Microbiol. 2003;5:650–9.
Hongmei J, Aitchison JC, Lacap DC, Peerapornpisal Y, Sompong U, Pointing SB. Community phylogenetic analysis of moderately thermophilic cyanobacterial mats from China, the Philippines and Thailand. Extremophiles. 2005;9:325–32.
Miller SR, Castenholz RW, Pedersen D. Phylogeography of the thermophilic cyanobacterium Mastigocladus laminosus. Appl Environ Microbiol. 2007;73:4751–59.
Takacs-Vesbach C, Mitchell K, Jackson-Weaver O, Reysenbach AL. Volcanic calderas delineate biogeographic provinces among Yellowstone thermophiles. Environ Microbiol. 2008;10:1681–89.
Reno ML, Held NL, Fields CJ, Burke PV, Whitaker RJ. Biogeography of the Sulfolobus islandicus pan-genome. Proc Natl Acad Sci USA. 2009;106:8605–10.
Bahl J, Lau MCY, Smith GJD, Vijaykrishna D, Cary SC, Lacap DC, et al. Ancient origins determine global biogeography of hot and cold desert cyanobacteria. Nat Commun. 2011;2:163.
Anderson RE, Kouris A, Seward CH, Campbell KM, Whitaker RJ. Structured populations of Sulfolobus acidocaldarius with susceptibility to mobile genetic elements. Genome Biol Evol. 2017;9:1699–710.
Podar PT, Yang Z, Björnsdóttir SH, Podar M. Comparative analysis of microbial diversity across temperature gradients in hot springs from Yellowstone and Iceland. Front Microbiol. 2020;11:1625.
Clark K, Karsch-Mizrachi I, Lipman DJ, Ostell J, Sayers EW. Genbank. Nucleic Acids Res. 2015;44:D67–D72.
Konstantinidis KT, Tiedje JM. Genomic insights that advance the species definition for prokaryotes. Proc Natl Acad Sci USA. 2005;102:2567–72.
Kim M, Oh HS, Park SC, Chun J. Towards a taxonomic coherence between average nucleotide identity and 16S rRNA gene sequence similarity for species demarcation of prokaryotes. Int J Syst Evol Microbiol. 2014;64:346–51.
Olm MR, Crits-Christoph A, Diamond S, Lavy A, Carnevali PBM, Banfield JF. Consistent metagenome-derived metrics verify and delineate bacterial species boundaries. mSystems. 2020;5:e00731-19.
Jain C, Rodriguez-R LM, Phillippy AM, Konstantinidis KT, Aluru S. High throughput ANI analysis of 90K prokaryotic genomes reveals clear species boundaries. Nat Commun. 2018;9:5114.
Shapiro BJ. What microbial population genomics has taught us about speciation. In: Polz MF, Rajora OP, editors. Population Genomics: Microorganisms. Cham, Switzerland: Springer International Publishing; 2019. p. 31–47.
Parks DH, Chuvochina M, Waite DW, Rinke C, Skarshewski A, Chaumeil PA, et al. A standardized bacterial taxonomy based on genome phylogeny substantially revises the tree of life. Nat Biotechnol. 2018;36:996–1004.
Chaumeil PA, Mussig AJ, Hugenholtz P, Parks DH. GTDB-Tk: a toolkit to classify genomes with the Genome Taxonomy Database. Bioinformatics 2020;36:1925–27.
Felsenstein J. Phylogenies and the comparative method. Am Nat. 1985;125:1–15.
Louca S. Phylogeographic estimation and simulation of global diffusive dispersal. Syst Biol. 2021;70:340–59.
Comas I, Coscolla M, Luo T, Borrell S, Holt KE, Kato-Maeda M, et al. Out-of-Africa migration and Neolithic coexpansion of Mycobacterium tuberculosis with modern humans. Nat Genet. 2013;45:1176–82.
Denef VJ, Banfield JF. In situ evolutionary rate measurements show ecological success of recently emerged bacterial hybrids. Science. 2012;336:462–6.
Bouckaert R, Cartwright R. Phylogeography by diffusion on a sphere: whole world phylogeography. PeerJ. 2016;4:e2406.
Brillinger DR. A particle migrating randomly on a sphere. In: Selected Works of David Brillinger. Cham, Switzerland: Springer; 2012. p. 73–87.
Ghosh A, Samuel J, Sinha SA. “Gaussian” for diffusion on the sphere. Europhys Lett. 2012;98:30003.
Castenholz RW. The biogeography of hot spring algae through enrichment cultures. SIL Commun. 1978;21:296–315. 1953-1996
Valentine DL. Adaptations to energy stress dictate the ecology and evolution of the archaea. Nat Rev Micro. 2007;5:316–23.
Louca S, Parfrey LW, Doebeli M. Decoupling function and taxonomy in the global ocean microbiome. Science. 2016;353:1272–77.
Smith DJ, Jaffe DA, Birmele MN, Griffin DW, Schuerger AC, Hee J, et al. Free tropospheric transport of microorganisms from Asia to North America. Micro Ecol. 2012;64:973–85.
Pagel M. Detecting correlated evolution on phylogenies: a general method for the comparative analysis of discrete characters. Proc R Soc Lond B Biol Sci. 1994;255:37–45.
Whitman WB, Coleman DC, Wiebe WJ. Prokaryotes: the unseen majority. Proc Natl Acad Sci USA. 1998;95:6578–83.
Anderson D. The regulation of fishing and related activities in exclusive economic zones. In: Modern Law Sea, Publications on Ocean Development, vol. 59, chap. 11. Leiden, The Netherlands: Brill Nijhoff; 2008. p. 209–27.
Bullock JM, Clarke RT. Long distance seed dispersal by wind: measuring and modelling the tail of the curve. Oecologia. 2000;124:506–21.
Brynjarsdóttir J, O’Hagan A. Learning about physical parameters: the importance of model discrepancy. Inverse Probl. 2014;30:114007.
Bell T. Experimental tests of the bacterial distance-decay relationship. ISME J. 2010;4:1357–65.
Hyatt D, Chen GL, LoCascio PF, Land ML, Larimer FW, Hauser LJ. Prodigal: prokaryotic gene recognition and translation initiation site identification. BMC Bioinforma. 2010;11:119.
Parks DH, Imelfort M, Skennerton CT, Hugenholtz P, Tyson GW. Assessing the quality of microbial genomes recovered from isolates, single cells, and metagenomes. Genome Res. 2014;25:1043–55.
Chambat F, Valette B. Mean radius, mass, and inertia for reference Earth models. Phys Earth Planet Inter. 2001;124:237–53.
Data NS, (SEDAC) AC Gridded Population of the World, Version 4 (GPW v4): Population Density, Revision 11. Tech. rep., Palisades, NY: Center for International Earth Science Information Network - CIESIN - Columbia University. 2018. Accessed November 23, 2020.
Price MN, Dehal PS, Arkin AP. FastTree 2: approximately maximum-likelihood trees for large alignments. PLoS ONE. 2010;5:e9490.
Britton T, Anderson CL, Jacquet D, Lundqvist S, Bremer K. Estimating divergence times in large phylogenetic trees. Syst Biol. 2007;56:741–52.
Zhu Q, Mai U, Pfeiffer W, Janssen S, Asnicar F, Sanders JG, et al. Phylogenomics of 10,575 genomes reveals evolutionary proximity between domains bacteria and archaea. Nat Commun. 2019;10:5477.
Perrin F. Étude mathématique du movement brownien de rotation. In: Annales scientifiques del’École Normale Supérieure, vol. 45. Paris, France: Elsevier; with 1928. p. 1–51.
Louca S, Doebeli M. Efficient comparative phylogenetics on large trees. Bioinformatics. 2018;34:1053–55.
Bloomquist EW, Lemey P, Suchard MA. Three roads diverged? routes to phylogeographic inference. Trends Ecol Evol. 2010;25:626–32.
Lemey P, Rambaut A, Welch JJ, Suchard MA. Phylogeography takes a relaxed random walk in continuous space and time. Mol Biol Evol. 2010;27:1877–85.
Faria NR, Suchard MA, Rambaut A, Lemey P. Toward a quantitative understanding of viral phylogeography. Curr Opin Virol. 2011;1:423–9.
Faria NR, Suchard MA, Abecasis A, Sousa JD, Ndembi N, Bonfim I, et al. Phylodynamics of the HIV-1 CRF02_AG clade in Cameroon. Infect Genet Evol. 2012;12:453–60.
Lange K. Diffusion processes. In: Applied Probability, chap. 11. New York, NY: Springer New York; 2010. p. 269–95.
Ondov BD, Treangen TJ, Melsted P, Mallonee AB, Bergman NH, Koren S, et al. Mash: fast genome and metagenome distance estimation using minhash. Genome Biol. 2016;17:132.
Pasolli E, Asnicar F, Manara S, Zolfo M, Karcher N, Armanini F, et al. Extensive unexplored human microbiome diversity revealed by over 150,000 genomes from metagenomes spanning age, geography, and lifestyle. Cell 2019;176:649–62.
Criscuolo A, Gascuel O. Fast NJ-like algorithms to deal with incomplete distance matrices. BMC Bioinforma. 2008;9:166.
Paradis E, Claude J, Strimmer K. APE: analyses of phylogenetics and evolution in R language. Bioinformatics. 2004;20:289–90.
Kinene T, Wainaina J, Maina S, Boykin LM, Kliman RM. Methods for rooting trees, vol. 3. Oxford: Academic Press; 2016. p. 489–93.
van Rossum G. Python tutorial. Tech. Rep. CS-R9526, Amsterdam: Centrum voor Wiskunde en Informatica (CWI); 1995.
Acknowledgements
SL was supported by a startup grant by the University of Oregon. The author thanks Qusheng Jin for valuable comments.
Author information
Authors and Affiliations
Contributions
All work was performed by S.L.
Corresponding author
Ethics declarations
Competing interests
The author declares no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Louca, S. The rates of global bacterial and archaeal dispersal. ISME J 16, 159–167 (2022). https://doi.org/10.1038/s41396-021-01069-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41396-021-01069-8
This article is cited by
-
Decoding populations in the ocean microbiome
Microbiome (2024)
-
A genus in the bacterial phylum Aquificota appears to be endemic to Aotearoa-New Zealand
Nature Communications (2024)
-
Dispersal changes soil bacterial interactions with fungal wood decomposition
ISME Communications (2023)
-
High speciation rate of niche specialists in hot springs
The ISME Journal (2023)
-
Trait biases in microbial reference genomes
Scientific Data (2023)