Abstract
Iron (Fe), an essential element for plant growth, is abundant in soil but with low bioavailability. Thus, plants developed specialized mechanisms to sequester the element. Beneficial microbes have recently become a favored method to promote plant growth through increased uptake of essential micronutrients, like Fe, yet little is known of their mechanisms of action. Functional mutants of the epiphytic bacterium Azospirillum brasilense, a prolific grass-root colonizer, were used to examine mechanisms for promoting iron uptake in Zea mays. Mutants included HM053, FP10, and ipdC, which have varying capacities for biological nitrogen fixation and production of the plant hormone auxin. Using radioactive iron-59 tracing and inductively coupled plasma mass spectrometry, we documented significant differences in host uptake of Fe2+/3+ correlating with mutant biological function. Radioactive carbon-11, administered to plants as 11CO2, provided insights into shifts in host usage of ‘new’ carbon resources in the presence of these beneficial microbes. Of the mutants examined, HM053 exhibited the greatest influence on host Fe uptake with increased plant allocation of 11C-resources to roots where they were transformed and exuded as 11C-acidic substrates to aid in Fe-chelation, and increased C-11 partitioning into citric acid, nicotianamine and histidine to aid in the in situ translocation of Fe once assimilated.
Similar content being viewed by others
Introduction
The role of plant growth promoting bacteria in agriculture
A survey of the literature establishes that many plant growth promoting bacteria (PGPB) can strongly influence plant growth and increase crop yields. Mechanisms have been suggested that support these actions, which include: antagonism toward phytopathogens and induction of plant resistance pathways [1]; phytostimulation through microbial production and secretion of plant relevant hormones like auxin, cytokinins, and gibberellins, as well as nitric oxide [2, 3]; improvements in host nitrogen uptake via biological nitrogen fixation (BNF) [4]; and improvements in host micronutrient uptake [5, 6].
Azospirillum brasilense, a Gram-negative bacterium, is perhaps the best studied of the PGPB. It is known to be a prolific grass root colonizer preferring to grow on the outer surface of its host’s roots. As a diazotroph this bacteria has the ability to fix N2 and, in some circumstances, transfer that source of nitrogen to its host [4]. A. brasilense can also produce auxin which can affect plant development [3]. Over the years, commercial inoculants of various strains of this bacterium have been developed and tested in the field for their influence on growth performance in a variety of grain crops, including maize, under various environmental conditions [7,8,9]. In fact, recently we showed that maize inoculated with the functional mutant HM053 A. brasilense exhibited 12% larger stem diameters, 23% increased leaf thickness and 58% higher chlorophyll content relative to non-inoculated control plants when surveyed during 2018 and 2019 growing seasons [10]. Furthermore, plant growth promotion resulted in significantly higher crop yields compared to non-inoculated control plants where 34% and 53% increases were documented in the number of corn kernels per cob from inoculated plants during the 2018 and 2019 growing seasons, respectively. These phenotypic attributes were ascribed to increased host iron uptake in the presence of the beneficial microbe, which not only improved plant growth, but also had the effect of increasing seed iron content thus improving the nutritional value of the crop product.
The importance of iron in plant growth
Iron is the third most limiting nutrient for healthy plant growth and development; however, this limitation is not attributable to its abundance. Quite the contrary, iron is the fourth most abundant element in the lithosphere. Even so, its bioavailability is limited especially in aerobic, neutral pH soils because it exists predominantly as insoluble Fe3+ oxyhydroxides that are unusable to plants [11, 12]. A deficiency of iron can decrease plant growth affecting crop yield and/or nutritional quality [13,14,15], while excess iron can lead to elevated Fe3+/Fe2+ redox reactions causing cellular damage.
Iron uptake in higher plants follows two mechanisms (Fig. S1), which are distinct between non-graminaceous plants and graminaceous plants. Non-graminaceous plants utilize a reduction-based strategy (Strategy I) through which protons are exuded from the roots via the actions of membrane-bound H+-ATPases [16,17,18,19,20,21,22,23,24,25,26,27,28,29]. These protons acidify the soil, solubilizing Fe3+. Additionally, membrane-bound Fe3+ reductase oxidases [20] reduce this solubilized iron to Fe2+ enabling its transport across the plasma membrane of the root epidermal cells through the actions of specialized ion transport proteins that are part of the larger ZIP (zinc-regulated transporter and iron-regulated transporter) encoding gene family [16, 17, 21,22,23,24]. Graminaceous plants like rice, barley and maize utilize a Fe3+ chelation mechanism (Strategy II) whereby phytosiderophores (PS) are excreted from the roots [25] where they chelate with Fe3+ in the soil. These PS-Fe3+ chelates are then taken into the root cells through either yellow stripe (YS) or yellow stripe-like (YSL) oligopeptide transporters first characterized in maize [26]. Even with these class distinctions nine ZIP-encoding genes have been identified in the maize genome with little known of their function, regarding Fe2+ uptake [24].
Once assimilated, iron will travel through the apoplastic space to reach the endodermis of the root. The endodermal ring can be a site rich in ion transporters facilitating the trafficking of several metal nutrients to the stele and xylem [27]. It also manifests as a natural barrier to ion trafficking in the form of the Casparian band and suberin lamellae. Here, a layer of waterproof lignin and suberin polymer exists, which can force ion nutrients to pass into the symplast [28]. Once in the vascular core, iron transport in the xylem occurs predominantly as the Fe3+–citrate complex [29,30,31,32]. However, iron is also capable of translocation in other forms including nicotianamine chelates (Fe-NA), mugineic acid chelates (Fe-MA) and dihydroxy mugineic acid chelates (Fe-DMA). NA is the direct amino acid precursor to all MAs and is produced from S-adenoysl methionine by nicotianamine synthase [33, 34]. Additionally, Fe2+ is capable of xylem translocation in the form of a histidine chelate [35].
Beneficial microbes as the path forward to promoting plant growth
While evidence is compelling that certain beneficial microbes can influence plant growth through improved micronutrient uptake, we know little about the mechanisms of action, which, if fundamentally understood, could lay the foundation for translating this approach to general farming practices perhaps with greater acceptance by the public.
In the present work, we provide an in-depth analysis into the mechanisms of action for the beneficial effects of A. brasilense in maize. Our work relies on A. brasilense functional mutants providing a spectrum of biological functions spanning auxin biosynthesis to BNF. The use of functional mutants to examine plant-microbial interactions is not new to the field, and has been used extensively in other PGPB systems to examine the effects of associated microorganisms on plant physiology [36]. The functional mutants of A. brasilense used in the present study include HM053, a Nif + constitutively expressed strain hyper-fixing N2 and producing high levels of auxin; FP10, a Nif – strain deficient in N2-fixation, but still produces auxin albeit at a slightly lower capacity than HM053; and ipdC, a strain significantly inhibited in its capacity to produce auxin.
Materials and methods
Plant growth
Maize kernels from Elk Mound Seed Co. (Hybrid 8100) were dark germinated at room temperature for two days. Seeds were inoculated with bacteria culture as appropriate and transplanted to a growth pouch wetted with sterile Hoagland’s basal salt solution (PhytoTechnology Laboratories, Shawnee Mission, KS 66215, USA) for ~1 week. Seedlings were transferred to an aeroponics system (Fig. S2), and nutrients were replenished every five days. Two week old plants were used for all the 59Fe radiotracer and mass spectrometry studies.
For 11C physiology and metabolism studies, including root emission measurements, plants were grown in eight inch plastic cones filled with Turface™ (expanded clay matrix) where the bottom portion of the cone was immersed in de-ionized water (Fig. S3). Nutrient was introduced as Hoagland’s solution every three days. Growth conditions for both set-ups consisted of 12-h photoperiods, 500 μmol m−2 s−1 light intensity, and temperatures of 25 °C/ 20 °C (light/dark) with humidity at 70–80% for two weeks.
Bacteria growth and root inoculation
Functional mutants of A. brasilense were obtained via a material transfer agreement between the corresponding author’s institution and the Federal University of Paraná (UFPR, Curitiba, PR CEP 81531-980, Brazil). HM053 originated as a natural mutant of the wild-type strain of A. brasilense FP2 (Sp7 ATCC 29145 Nif+ Smr Nalr) screened through its resistance to ethylenediamine (EDAr) according to the original work of Machado et al. [37, 38]. The FP10 mutant was obtained through the original work of Pedrosa and Yates [39] by N-nitrosoguanidine mutagenesis of the FP2 wild-type strain of A. brasilense and isolated by growth on glutamate medium. The indole-3-pyruvate decarboxylase gene (ipdC), coding for a key enzyme of the indole-3-pyruvic acid pathway of auxin (indole-3-acetic acid) biosynthesis in A. brasilense, was functionally disrupted in a site-specific manner using a SacB-cassette insertion into the ipdC gene of wild-type FP2 (Sp7 ATCC 29145) followed by homologous recombination. The method allowed for the construction of the ipdC mutation strain without unwanted sequence changes and relied on the λ Red recombineering (Direct and Inverted Repeat stimulated excision; DIRex) a method that works well for generating single point mutations, small insertions or replacements as well as deletions of any size, in a bacterial gene [40]. The resultant knock-out strain exhibited significant reduction in auxin biosynthesis to a level of 10% that of the wild-type strain [41].
Functional mutants were grown in liquid NFbHP-lactate medium following published procedures [4]. Cultures were washed with sterile water and diluted to ~106–108 colony forming units (CFU) per mL volume. Root inoculation involved adding 1 mL of inoculum to a Petri dish containing 10 maize seedlings (2 days after germination) and rocking in the shaking incubator for 2 h. Seeds were placed into plastic seed germination pouches (PhytoAB, Inc., San Jose, CA 95131, USA) for five days before transplanting to aeroponics or Turface™.
Radioactive Fe-59 tracer studies
Two oxidation states of radioactive 59Fe (t½ 44.5 d) were used in the study, ferrous (Fe2+) and ferric (Fe3+), purchased from Perkin-Elmer Life Sciences (Akron, OH USA). One hour before administration of tracer, plants were removed from aeroponics growth cells and placed in 600-mL beakers with 100 mL of 5 μM Fe3+-EDTA to prohibit induction of water- or iron-stress responses during the study. In auxin treatment studies, 50 mL of 30 μM indole-3-acetic acid (auxin) was added to 50 mL of the iron solution for a total of 100 mL. Iron radiotracer of 0.74 MBq was introduced. After 3 h, the roots were cut from the shoots and washed in water followed by an EDTA solution, then blotted dry. The root and shoot tissues were counted in a three inch NaI (PMT: Photomultiplier Tube) gamma well-type detector for percent assimilation and allocation calculations.
Production and administration of radioactive 11CO2
11CO2 (t½ 20.4 min) was produced on the GE PETrace Cyclotron located at the Missouri Research Reactor Center using high-pressure research grade N2 gas target irradiated with a 16.4 MeV proton beam to generate 11C via the 14N(p,α)11C nuclear transformation [42, 43] using our published procedures where details can be found in the Supplementary Methods [44].
Whole-plant 11C-physiology measurements
After pulse labeling using 11CO2, plants were incubated for 3 h before separating the load leaf, shoots, roots and growth media. Roots were washed in 100 mL de-ionized water to remove surface root exudates. Measurement of 11C-activity was performed using gamma counting and data was decay corrected to end-of-bombardment. The individual components (plant tissues, growth media and root wash) were summed for total plant 11C-activity. Individual components were used to calculate leaf export, root allocation and root exudation fractions. Aliquots of the root washings were processed through a 1 mL column packed using 200 mesh AG1 X-8 anion exchange resin (Bio-Rad Laboratories, Inc., Hercules, CA, USA). Afterward, the column and breakthrough rinse were counted for radioactivity providing a measure of the acidic and non-acidic fractions of root exudates.
[11/12C]metabolite analyses
In separate studies, leaves were exposed to 11CO2 for 20 min before metabolite analyses following published procedures [45]. Tissues were flash frozen in liquid nitrogen, ground to a fine powder and extracted in methanol: water (60:40 v/v) in Eppendorf™ tubes. After centrifugation the insoluble and soluble portions were separated and counted for 11C-activity using a NaI (PMT) gamma counter. The insoluble portion contained mostly cell-wall polymers and starch. The soluble portion contained small water-soluble compounds, including sugars, amino acids and non-nitrogen containing organic acids. All data was decay corrected back to the end-of-bombardment. [11C]-Sugars were analyzed by radio thin layer chromatography (TLC) using glass backed NH2-silica HPTLC-plates (200 µm, w/UV254) purchased from Sorbent Technologies (Atlanta, GA, 30071, USA) according to published procedures [46, 47] (see Supplementary Methods).
[11C]Amino acids were analyzed following published procedures [45] using pre-column OPA derivatization and quantified by gradient radio HPLC (see Supplementary Methods).
For [11C]organic acid analysis, leaf extract was processed through a QMA Sep-Pak™ then rinsed with 10 mL of deionized (DI) water followed by 1 mL of 20 mM sodium phosphate buffer (pH = 1.5). Aliquots of the phosphate rinse were analyzed using a gradient radio HPLC equipped with a UV detector (210 nm) and an on-line NaI (PMT) radiation detector. The acids were separated using a Phenomenex Gemini 5 µm C18 (250 × 4.6 mm inner diameter) column with a solvent system of solvent A (20 mM potassium phosphate monobasic pH of 2.5) and solvent B (60:40 methanol: acetonitrile: (v/v)) starting at 100% A at injection and ramping to 75:25 A:B within 10 min at a flow rate of 1 mL min−1. Metabolite mass and radioactivity were recorded and quantified as described above for amino acids.
[11C]Nicotianamine was analyzed by reacting a portion of the QMA extract with 10 mM ferrous sulfate heptahydrate (1:1 v/v) for 30 min to chelate the nicotianamine. A radio HPLC system was used in the analysis (see Supplementary Methods) where 20 mM potassium phosphate buffer at pH 2.50 (solvent A) and 60:40 methanol:acetonitrile (solvent B) were pumped through a Phenomenex Gemini C18 110 A 250 × 4.60 mm column. The solvent gradient changed from 100% solvent A to 75%:25% A:B at 10 min, and back to 100%A by 20 min where it was held for 5 min. The chelated sample was measured by UV absorption at 243 nm. The amount of nicotianamine mass per gram fresh weight tissue was assessed for control, HM053, FP10 and ipdC treatments using authentic nicotianamine standards to calibrate the detector response. Radioactivity associated with this metabolite was measured by NaI (PMT) detector connected in series with the UV detector.
Specific activities (SA) of individual metabolites of interest including [11C]histidine, [11C]citric acid and [11C]nicotianamine were calculated using Eq. (1) following procedures described in the Supplementary Methods:
[2-11C]indole and its application to measuring microbial auxin biosynthesis
Radiosynthesis of [2-11C]indole followed published procedures using [11C]hydrogen cyanide [48]. Synthesis took less than 1 h providing product at >98% radiochemical purity, and specific activity of 176 ± 24.8 GBq μmol−1.
Aliquots of purified radiotracer were dispensed into 10 mL Falcon tubes containing thoroughly washed and harvested bacteria re-suspended in sterile water. Bacteria count was measured by sample turbidity, where OD600 = 1.0 (Optical Density at 600 nm, corresponded to 108 cells mL−1). Once tracer was introduced, tubes were gently rocked at ambient temperature for up to 2 h and samples were periodically removed for analysis of [2-11C]indole-3-acetic acid conversion.
[2-11C]indole-3-acetic acid analysis was carried out using radio HPLC using a Phenomenex Gemini 5 µm C18 (250 × 4.6 mm inner diameter) column at 35 °C and a mobile phase composed of acetonitrile:water (50:50, v/v %) at flow rate of 1.0 mL min−1. The fluorescence detector was set with excitation and emission wavelengths of 230 nm and 360 nm, respectively with a NaI (PMT) gamma radiation detector in series.
Inductively coupled plasma-mass spectrometry (ICP-MS)
For ICP-MS analyses, roots and shoots were dried in an oven, weighed into digestion vessels and digested in 3.0 mL of concentrated nitric acid at 190 °C in a Milestone Ethos Plus (Milestone SRL, Sorisole, Italy) microwave digestion system. Digestants were diluted to 50 mL with ultrapure water and gravimetrically diluted by a factor of 10 with 0.45 N nitric acid. Samples were analyzed via Perkin-Elmer NexION ICP-MS in Kinetic Energy Discrimination mode. Total elemental ion counts were measured of 56Fe and normalized to 12C counts. Reference materials included NIST SRM 1570 spinach leaves and NIST SRM 1573 tomato leaves prepared as samples were. Internal Calibration standards of Sc, In, and Tl at known concentrations were used from stock solutions (High Purity Standards, Charleston, SC 29418, USA).
For Laser Ablation-ICP-MS, roots were removed from shoots and a 3-cm section of primary lateral root was further excised above the apical meristem. Roots were sectioned in OCT embedding media to 100 μm thickness (Fisher Scientific Inc., Hampton, NH 03801, USA) and placed on quartz microscope slides for freeze drying in a FreezeZone 1 dryer (Labconco Corp., Kansas City, MO, USA) before ablation.
Transmission electron microscopy
Specimens were collected from control and inoculated groups and processed for transmission electron microscopy (TEM) using standard procedures of the U. Missouri Microscopy Core, details in the Supplementary Methods.
Root ethylene emission measurements
Aeroponically grown plants were harvested at two weeks, roots weighed and placed in air-tight jars equipped with a sampling port. Roots from 2–3 plants were binned generating a single data point. Roots volatilized for 90–130 h and 5-mL of head space was sampled using a gas-tight syringe. The sample was injected into a flame-ionization Hewlett Packard 5890 A gas chromatograph with a 2-m long, 1-mm inner diameter ShinCarbon ST packed column. The program started with a 2 min hold at 40 °C and increased 10 °C min−1 to 250 °C. The injector temperature was 250 °C and the flame ionization detector temperature was 300 °C. Chromatographic peaks for ethylene were measured using PeakSimple™ chromatography software and quantified against ethylene standards. Ethylene emission values were reported as ρmol ethylene gfw−1 tissue hr−1 as a function of bacteria treatment.
Root gravitropism measurements
Maize seeds were placed upright in Gelrite™ solid media in a growth box. The box was rotated 90° after root growth reached half way down the box in the Gelrite™ media. Photographs of the roots were taken after 12 h. Root bending angles were measured with Image-J software. The degree of bending as a function of bacterial treatment was assessed as an indirect measurement of plant-root auxin status.
Root indole measurements
Fresh roots were harvested from two week old plants, weighed, and placed in air-tight jars equipped with two sampling ports. The roots volatilized for 48–90 h. A glass tube trap was loaded with 50 mg Tenax® GR, a composite mix of Tenax® TA (Scientific Instrument Services, Inc., NJ 08551, USA), porous polymer based on 2, 6-diphenylphenol with 30% graphite additive. This trap was affixed to one port equipped with a silicon rubber seal and an aquarium air pump was affixed to the other port. Air was pumped through the chamber for 1 h at a rate of 50 mL min−1 providing three complete air exchanges. Tenax® traps were rinsed with 0.5 mL of methylene chloride and 1–3 μL volumes of this extract were analyzed by capillary gas chromatography using a Hewlett Packard 5890 A gas chromatograph equipped with a 30-m long, 0.53 mm inner diameter, 0.25 µm film thickness, Restek RTX-WAX column for separation and a flame ionization detector. The program was as described for ethylene measurements. Chromatographic peaks for indole eluted at 15 min and were measured using PeakSimple chromatography software and quantified against standards. Root indole volatile emission rates were reported as ρmole indole gfw−1 h−1 as a function of bacteria treatment.
Root DIMBOA measurements
Maize root tissue (700–800 mg) was macerated by mortar and pestle in 2.00 mL of DI water. The resulting extract was centrifuged, combined with 1 mL of ethyl acetate (ETAC) and gently rocked for 1 min. The DIMBOA/ETAC organic layer was removed and the aqueous phase was re-extracted with 1 mL ETAC and combined with the first organic sample. An aliquot of the organic phase was injected into a Hewlett Packard 5890 A gas chromatograph equipped with a 30-m long, 0.53 mm inner diameter, 0.25 µm film thickness, Restek RTX-WAX capillary column and flame ionization detector. The program started with a 2 min hold at 70 °C and increased 10 °C min−1 to 250 °C. The injector temperature was 250 °C and the flame ionization detector temperature was 300 °C. The retention time of DIMBOA was ~24 min. Chromatographic peaks for DIMBOA were measured using PeakSimple chromatography software and quantified against authentic standards (Santa Cruz Biotechnology, Inc., TX, USA). Root DIMBOA values were reported as μmole DIMBOA gfw−1 tissue as a function of bacteria treatment.
In vitro chemotaxis assays to examine effects of DIMBOA on bacteria growth
A standard of DIMBOA was prepared by dissolving 10 mg of compound in 6 mL of deionized sterile water using 200 μL DMSO (3% v/v) to facilitate sample dissolution. Aliquots of this standard were then added to bacteria growth media (as described in Bacteria Growth Methodology section) to generate 0.02 mM, 0.1 mM and 0.5 mM concentrations of DIMBOA. Each dose was replicated in triplicate. The amount of DMSO for each dose was adjusted to match that of the highest DIMBOA concentration. Additionally, 0 mM DIMBOA controls were prepared (also in triplicate) where an equivalent amount of DMSO as that found in the highest 0.5 mM DIMBOA dose was introduced to null effects of this organic solvent on bacteria growth. A. brasilense cultures of the functional mutants HM053, ipdC and FP10 were introduced into these modified media and grown under previously described conditions for 24 h after administration of DIMBOA to the growth media. An additional sample of media containing 0.5 mM DIMBOA, but no bacteria, was set aside for examining DIMBOA stability at 30 °C. OD600 measurements were taken using a ThermoScientific Evolution 201 UV-vis spectrophotometer at 24 h into the bacteria growth cycle. The 0.5 mM DIMBOA cultures were re-measured after 48 h of incubation. DIMBOA stability measurements were conducted over 27 h using capillary gas chromatography and flame ionization detection using the analytical methodology previously described above. Samples were prepared for GC injection by extracting 200 μL of growth media using 500 μL of ETAC. Aliquots of the organic extract were then injected into the gas chromatograph using a 1:20 sample split upon injection.
Statistical analysis
Data was subjected to the Shapiro–Wilk Normality Test to identify outliers so all data groups reflected normal distributions. Data was analyzed using the Student’s t-test for pair-wise comparisons made between non-inoculated controls and bacteria treatment. Statistical significance was set at P < 0.05. The 59Fe3+/2+ allocation data was analyzed by Principal Component Analysis (PCA) using XLSTAT software version 2020.3 (Addinsoft Inc., New York, NY 10001, USA).
Results and discussion
Bacteria influence host uptake and translocation of iron
Radioactive 59Fe3+ and 59Fe2+ tracers were applied to roots inoculated with the three strains (HM053, FP10 and ipdC) of A. brasilense. Comparisons in host iron assimilation were made to non-inoculated control plants and to plants chemically treated with auxin. As expected from the Strategy II iron uptake mechanism of grasses, non-inoculated maize roots showed higher 59Fe3+ uptake than 59Fe2+ (Fig. 1A). Qualitatively, radiographic imaging following tracer administration in non-inoculated control plants revealed high uptake of 59Fe3+ in root meristems and lateral roots (Fig. 2A). 59Fe2+ was taken up to a much lesser extent. Quantitative tissue counting and root radiographic images of roots inoculated with the functional mutants of A. brasilense revealed very different behavior. The effects of bacteria on host iron assimilation (Fig. 1A) and root-to-shoot translocation (Fig. 1B) are striking. Especially with regard to the HM053 functional mutant, such effects on the host plant seem to be tied to biological nitrogen fixing (BNF) and auxin producing capacity of the microorganism. Drop plate assays examining the degree of microbial colonization under our plant growth conditions (Fig. S4) provided evidence that inoculation with all three functional mutants generated between 6.3 × 106 and 3.2 × 107 CFUs gfw−1 of root tissue relative to roots from non-inoculated control plants yielding a substantially lower background microbial count of 6.3 × 105 CFUs gfw−1. HM053 showed the greatest degree of colonization which could account, in part, for its enhanced performance. However, ipdC and FP10 exhibited the same levels of colonization, yet FP10 outperformed ipdC in influencing host iron assimilation. The extent of colonization does not fully explain the actions here, and thus microbial biological functions must play a role.
A Bar graph of biological assimilation of radioactive ferrous 59Fe2+ and ferric 59Fe3+ after 3 h of incubation of maize roots using 0.74 MBq of tracer. Biological assimilation reflects a combination of root and microorganism assimilation of tracer. Data presented as weight normalized percentage of 59Fe radioactivity administered. Asterisks indicate significant differences of treatment relative to control (*P < 0.05; **P < 0.01; ***P < 0.001). B Bar graph showing the extent of root-to-shoot transport of radioactive 59Fe2+ and 59Fe3+ after 3 h of incubation of maize roots using 0.74 MBq of tracer for the same five study conditions. Data presented as weight normalized percentage of the assimilated 59Fe radioactivity. Asterisks indicate significant differences of treatment relative to control (*P < 0.05; **P < 0.01; ***P < 0.001). C Bar graph showing auxin biosynthetic rates in A. brasilense using [11C]indole radiotracer reveals a trend of increasing rate of production with microbial BNF capacity. Correlations were made to BNF data measured using 13NN [4]. Data (±SE) reflects N = 4 biological replicates. D Correlation plot mapping 59Fe2+ allocation against 59Fe3+ allocation for the study conditions reveals systematic trends of clustering with treatments that correlate to auxin producing and N2-fixing capacities of the beneficial microbes. E Principal component analysis correlates 59Fe translocation to biological functions of the beneficial microbes and to chemical treatment using auxin.
Comparisons were made between non-inoculated control roots (A) and roots inoculated with HM053 (B), ipdC (C) and FP10 (D) strains of A. brasilense with 59Fe3+ images are shown in the upper portion of each panel while 59Fe2+ images are shown in the lower portion. ‘Hot’ spots of 59Fe radioactivity (both for 59Fe3+ and 59Fe2+) are seen at lateral root junctions. Roots inoculated with the FP10 strain of A. brasilense only showed 59Fe3+ ‘hot’ spots. Shoot images for HM053 inoculated plants are shown for 59Fe3+ (E) and 59Fe2+ (F). Translocation of 59Fe3+ to shoots resulted in higher tracer accumulation in all of the leaf tips. Translocation of 59Fe2+ to shoots resulted in a higher distribution of tracer in the younger developing leaf tissue.
The spatial patterning of Fe-59 radioactivity in roots associated with bacteria was different from non-inoculated control roots, with increased 59Fe3+ and 59Fe2+ accumulation noted at the lateral root junctions (Fig. 2B–D). FP10 was the exception, which did not exhibit significant 59Fe2+ assimilation, but substantial 59Fe3+ assimilation (Fig. 2D). The spatial patterning of radioactivity is consistent with published confocal microscopy images demonstrating the lateral root junctions typically hold high populations of these bacteria [4]. Differences in spatial patterning exhibited by non-inoculated and inoculated roots suggests bacteria assimilation of iron can contribute to a significant portion of the measured root radioactivity detected in the NaI (PMT) gamma counter even after thorough root washing.
Systematic trends defining in planta translocation (Fig. 2E, F) of iron become apparent in the correlation plot (Fig. 1D) of 59Fe2+ transport versus 59Fe3+ transport and in the PCA biplot (Fig. 1E). Here, the information included in our allocation measurements were represented by feature vectors (F1 and F2) representing 86.1% and 13.9% of the information embedded in the data respectively (for more detail see Supplementary Information). As displayed, each of the treatments clustered together, indicating behavior within a treatment-type that is distinct from other treatments. It shows that ipdC and non-inoculated maize are similar in overall iron allocation behavior, while auxin chemical treatment and HM053 inoculated maize share similar allocation patterns. FP10 is most unique in its allocation patterns. Non-inoculated maize and ipdC inoculated maize clustered away from active variable vectors. This indicates both treatment types do not allocate much iron overall compared to other treatments, and ipdC narrowly allocates more 59Fe2+ than 59Fe3+. FP10 inoculated maize are distinctly apart from other treatments in the negative X- and Y-axis direction. This is more positively correlated with Fe3+ allocation and negatively with Fe2+ allocation - which is observed in the correlation plot (Fig. 1D), as well. Finally, the auxin chemical treatment and HM053 inoculated maize share positions along the active variable vectors, indicating they have the largest allocation percentages across all treatment types for both 59Fe3+ and 59Fe2+. HM053 inoculated maize tend to have the largest allocation values, especially for the 59Fe2+ as noted by the majority of points along that active variable vector while the auxin chemical treatment tends to show more 59Fe3+ than 59Fe2+ allocation as indicated by its position on the biplot.
To summarize, allocation of 59Fe2+ appeared stimulated by BNF while both 59Fe3+ and 59Fe2+ appeared stimulated by the auxin biosynthetic capacity of the inoculated bacterium. This assumes bacteria-borne auxin or indole-like substrates move from the microorganism into their host. Exogenously applied chemical auxin stimulated host iron assimilation and whole-plant iron translocation just as HM053 inoculated plants did (Fig. 1A, B). BNF and auxin biosynthesis are not mutually exclusive microbial functions. BNF capacity seems to impact bacterial ability to produce auxin (Fig. 1C), although it is not essential for that function. FP10, the BNF deficient strain, was able to biosynthesize auxin at a rate of 7.0 ± 0.4 molecules cell−1 s−1, slightly less than the 9.2 ± 0.9 molecules cell−1 s−1 production rate demonstrated by the FP2 wild-type strain and significantly less than the 13.4 ± 0.9 molecules cell−1 s−1 auxin production rate of the HM053 strain. Though not directly measured by our group using tracer analysis as done with other mutants, ipdC’s capacity to produce auxin was reported to be ~10% that of the wild-type strain [41].
Bacteria influence host cellular-scale iron distribution
Laser ablation-inductively coupled-mass spectrometry (LA-ICP-MS) was applied to root sections to corroborate radiotracer data by spatially mapping stable Fe-56 at cellular scales. Significantly, elevated Fe-56 accumulation was observed in the endodermal ring from HM053, but not ipdC or FP10 inoculated plants, nor in non-inoculated control root sections (Fig. 3A, B). Digestive ICP-MS analysis enabling measurement of absolute Fe-56 concentrations in roots and leaves (Fig. 3C) showed that HM053 significantly increased whole-plant tissue iron levels 4-fold in leaves and 10-fold in roots relative to controls, corroborating radiotracer experiments. Plants inoculated with ipdC showed lower levels of Fe-56 than non-inoculated controls. Though not entirely consistent with our radiotracer data, we note that levels of Fe-59 allocation over the short term were very small such that over a longer period of plant growth, iron accumulation could manifest in the same or lower iron signature than controls. Similarly, FP10 showed only slightly lower or same levels of iron as controls.
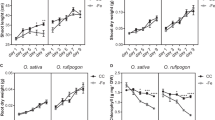
A Representative Fe-56 ion signals (in red) from the laser ablation of 100 μm root sections (taken 1 cm from the root tip) from non-inoculated control plants, and plants inoculated using HM053, ipdC and FP10 bacteria. B Data from ablation tracks was averaged across N = 6 biological replicates, and presented graphically as % distribution of the integrated Fe-56 ion signal for the different regions of the root cross section (epidermis, cortex, endodermis and inner pith region). These regions are identified on the inset anatomical key. HM053 treatment resulted in significantly higher iron levels in the endodermis than controls. C Absolute concentrations of Fe-56 (ppm) in leaves and roots were determined from direct injection ICP-MS of digested tissues and compared against NIST standards. D Transmission electron microscopy images on endodermal root cells reveal morphological differences between cell walls of non-inoculated control roots and those from HM053. HM053 inoculated roots appeared to have a more defined Casparian band in endodermal radial root walls than in control plants (highlighted by black arrows). E Representative Fe-56 signals from 40 μm diameter x 2 mm laser tracks rastered across freeze dried leaf tissues harvested from non-inoculated, HM053, ipdC and FP10 inoculated plants. The periodic patterning of the iron ion signals (red lines) presumably are due to the higher disposition of iron in leaf cell walls after the freeze drying process. The white lines reflect an average ion signal across the laser track. HM053 elicits a unique patterning with lower iron signal in and around vascular tissues that was not seen with the other beneficial microbes. F The ion signal for Fe-56 was integrated across the laser ablation track and plotted as a function of biologic type. HM053 showed a higher integrated iron signal than non-inoculated control, as well as ipdC and FP10 inoculated plants. G Relative percent of the Fe-56 ion signal in vascular tissue was plotted as a function of biologic type. HM053 inoculation decreased the relative percentage of iron in leaf vascular tissue suggesting greater mobility. Data (±SE) reflected N = 4–6 biological replicates. Asterisks indicate significant differences of treatment relative to control (*P < 0.05; **P < 0.01; ***P < 0.001).
To investigate high iron accumulation in the endodermal ring of HM053 inoculated plants, we used transmission electron microscopy to examine the radial walls of root endodermal cells (Fig. 3D). Morphological differences were seen between non-inoculated and HM053-inoculated roots with HM053 showing more prominent development of the Casparian band between cell walls. The Casparian band is a layer of waterproof lignin and suberin located at the endodermal ring. This barrier forces iron nutrients to pass from the apoplast into the symplast as they move to the stele and vascular core [27]. Once iron has entered the symplast, it is bound to various chelators that facilitate its transport as Fe3+-chelate. However, the endodermal ring can be a site harboring a rich density of metal transporters that may transport Fe2+ as well as other metals [27].
High accumulation of iron in the endodermal ring seen with HM053 inoculation implies the bacteria must influence cellular transport of this element, whether as Fe2+ or Fe3+, from the outer epidermal cells to the endodermal ring. Because our LA-ICP-MS analyses were conducted on freeze-dried root sections, likely metal ion distributions across each cell will be affected. Therefore, we are reluctant to draw any conclusions at this time on whether apoplastic transport dominated iron cellular trafficking. In future studies, we hope to re-examine this behavior once a cryostage is constructed enabling laser ablation of fresh tissue sections.
There is evidence, however, that cellular iron transport is promoted by HM053 from leaf tissue analyses using LA-ICP-MS (Fig. 3E–G). Here, HM053 inoculated plants significantly increased Fe-56 transport out of foliar vascular tissues (Fig. 3G) relative to controls while ipdC and FP10 inoculated plants showed no difference in vascular tissue iron levels compared to controls.
Bacteria influence host use of its carbon resources
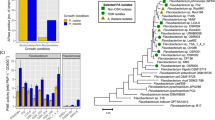
The effect of HM053, ipdC and FP10 inoculants on their host’s physiological and metabolic posture was examined to better understand the mechanism for bacterial induced enrichment of plant iron. Administration of radioactive 11C to live plants as 11CO2 allowed the measurement of leaf carbon fixation, export of 11C-photosynthates from shoots-to-roots, and exudation of 11C-substrates to the surrounding rhizosphere (Fig. 4A–C). These experiments showed that the functional mutants induced significantly different host physiological responses relative to non-inoculated control plants and each other. Increased carbon input via 11CO2 fixation with HM053 and FP10 inoculants correlated with higher root allocation of carbon resources and root exudation (Fig. 4A–C). Exudation of acidic substrates with these functional mutants was 2.8-fold and 2.1-fold higher for HM053 and FP10 inoculated plants, respectively, than non-inoculated control plants (Fig. 4C). This behavior was confirmed with a pH visual assay (Fig. S5). Taken together, these observations could help explain the noted increase in Strategy II iron assimilation using HM053 and FP10 inoculants because certain acidic exudates behave as chelating agents of Fe3+. Additionally, different patterns of metabolic partitioning of ‘new’ carbon (as 11C) into the plant’s metabolic landscape were noted between the functional mutants and non-inoculated control plants (Fig. 4D). In particular, HM053 and FP10 inoculation significantly increased 11C partitioning into the amino acid (AA) pool 4.5-fold and 2.3-fold, respectively, relative to controls. Inoculation with ipdC did not have an effect on the AA pool relative to controls.
A Leaf tissue was normalized to 11CO2 fixation and presented as % 11C-activity in the pulse applied to the leaf cuvette. B Leaf export and root allocation of 11C-photosynthates (measured at 3 h post 11CO2 pulse) presented as % fixed 11C-activity by the plant. C Root exudation of 11C-substrates (A = acidic substrates; N = non-acidic substrates measured at 3 h post 11CO2 pulse) presented as % fixed 11C-activity by the plant. D Metabolic landscape reflecting the partitioning of ‘new’ carbon (as 11C) into different metabolic pools of the load leaf tissue. E [11/12C]-histidine, [11/12C]-nicotianamine and [11/12C]-citric acid specific activities (SA) levels measured in leaf tissue 20 min. after tracer administration and represented as % total 11C-activity/μmol substrate gfw−1 of tissue. Data (±SE) reflects N = 7–9 biological replicates. Asterisks indicate significant differences relative to control (*P < 0.05; **P < 0.01; ***P < 0.001).
Other subtle differences in the metabolic landscape were seen including HM053 influence increasing host ‘new’ carbon partitioning into structural metabolite pools (Fig. 4D). This strongly correlates with our prior work demonstrating that plants inoculated with HM053 had thicker leaves and larger stems than non-inoculated controls [10].
Insight into particular points of metabolic regulation and the bearing these actions may have on host iron transport prompted examination of individual metabolite 11/12C specific activities (SA). Large changes in metabolite SA values reflect high metabolic input of ‘new’ carbon resources while substrate endogenous concentrations are being depleted by the host’s demands for a particular metabolite. SA values for [11/12C]histidine, [11/12C]nicotianamine, and [11/12C]citric acid were significantly elevated with HM053 inoculation relative to control plants. Plants inoculated with FP10 showed significant elevation in [11/12C]nicotianamine SA, but significantly lower values for [11/12C]histidine and [11/12C]citric acid relative to controls. ipdC inoculation had no effect on SA values (Fig. 4E).
Histidine, one of the least abundant AAs in plant metabolism, plays important roles as a proton shuttle substrate in the electron transport train of the plant’s photosynthetic PSII system [49]. HM053 promoted increased 11CO2 fixation (Fig. 4A) suggesting up-regulation of leaf photosynthesis. However, this may also be due in part to observed increases in leaf chlorophyll with HM053 inoculation [10].
We note that histidine is also an important chelating agent in the cell-to-cell transport of divalent metals including Fe2+ [50]. Hence, the significant increase in [11/12C]histidine SA with HM053 inoculation may, in part, contribute to the increased translocation of 59Fe2+ (Fig. 1B). Inoculations with ipdC or FP10 bacteria had no effect or decreased [11/12C]histidine SAs relative to controls, respectively. Recall that ipdC showed slightly elevated levels of 59Fe2+ allocation relative to controls while FP10 showed a significant reduction in 59Fe2+ allocation.
Like histidine, nicotianamine and citric acid are also important chelating agents of iron, facilitating transport through the plant’s vascular architecture [51]. Both HM053 and FP10 inoculates significantly elevated [11/12C]nicotianamine SAs while only HM053 increased [11/12C]citric acid SA. Together, these changes in host metabolic regulation are relevant to explaining why HM053 was the better performer in stimulating host iron assimilation and allocation.
Bacteria-borne indole could influence host root function
Additional influences of A. brasilense on host root performance were examined to better understand the mechanisms for iron enrichment. Ethylene, auxin, DIMBOA, and indole are interrelated in regulation and/or root metabolic and physiological functions. DIMBOA, a benzoxazinoid secondary metabolite in maize, shares a common branch point with auxin biosynthesis from the shikimate pathway at the indole precursor (Fig. 5). Therefore, the bacteria’s own auxin producing capabilities might influence host root auxin biosynthesis and/or sensing.
The biosynthesis of auxin precursors such as indole-3-glycerol phosphate, indole and L-Trp takes place in plastids and are generated via the shikimate pathway. Four putative L-Trp pathways for auxin biosynthesis are shown. Enzymes known to operate these pathways are shown in italics. Solid pathway arrows reflect active processes. Dashed arrows are suggested to exist, but have not been proven. Benzoxazinoid biosynthesis yielding DIMBOA and its conjugate analogs share a common metabolic branch point with auxin as indole. A Gravitropic responses of root tips are associated with auxin biosynthesis and/or sensing. Inoculation of roots using HM053, ipdC and FP10 bacteria reduced root gravitropic responses relative to controls. B Auxin and ethylene biosynthesis are synergistic – inoculation of roots using HM053, ipdC and FP10 bacteria reduced root ethylene emission rates relative to non-inoculated controls. C Root indole emission rates were significantly elevated relative to controls in roots inoculated with HM053, ipdC and FP10 bacteria. D Root DIMBOA concentrations were significantly increased relative to control in roots inoculated with HM053 and FP10 bacteria and unchanged for ipdC bacteria. Data (±SE) reflects N = 6–8 biological replicates. Asterisks indicate significant differences relative to control (*P < 0.05; **P < 0.01; ***P < 0.001).
In all higher plants, indole is produced from indole-3-glycerol phosphate and channeled into different pathways. Indole is transformed into tryptophan, and later auxin, by the tryptophan synthase-α subunit and tryptophan synthase-β subunit [52, 53]. In maize specifically, indole is also produced by the BX1 enzyme as an intermediate in the production of DIMBOA, and other benzoxazinoid secondary metabolites, which play important roles as allelochemicals in maize defense [54, 55]. However, indole and its role in maize metabolism is more complicated than serving as a metabolic intermediate for DIMBOA or auxin biosynthesis. Aside from the previously mentioned routes, it can also be formed by indole-3-glycerol phosphate lyase (IGL), which subsequently releases it as a volatile in response to herbivore attack [56]. Indole release has been shown to occur earlier than other induced biochemical responses where the Igl gene is induced in response to the actions of certain fatty acid derivatives such as N-(17-hydroxylinolenoyl)-L-glutamine isolated from regurgitate of some herbivore-pests [57]. Hence, as a chemical agent, indole can play an important role in maize defense priming, preparing systemic tissues and/or neighboring plants for an impending attack [58].
Ancillary data was acquired supporting the premise that bacteria-borne indole may be influencing host metabolism and root physiology. First, root behavior related to auxin (root gravitropism) was investigated due to the genotypic differences in the bacterial mutant strains’ ability to produce auxin. It was noted that FP10, ipdC and HM053 strains all exerted similar reductions in root gravitropism (Fig. 5A), a phenotype highly dependent on auxin biosynthesis and/or sensing in the root tip [59]. Because this was elicited by all bacteria strains, it is suspected that infusion of bacteria-borne indole, rather than auxin, may offset auxin homeostasis at the root tip. All functional mutants used in the present study should produce indole, while solely ipdC is deficient in the major pathway of auxin formation from indole.
Auxin and ethylene biosynthesis are mutually intertwined [60] with auxin exerting control over 1-aminocyclopropane-1-carboxylate oxidase (ACC oxidase), the controlling enzyme in ethylene biosynthesis. ACC oxidase is a member of the non-heme Fe2+ dependent family of oxygenases and oxidases in higher plants [61]. Hence, there could be a strong connection between plant auxin regulation and iron assimilation through the auxin-ethylene relationship. Indeed, it was noted that treatment of plants with exogenous auxin increased uptake and allocation of Fe3+ and Fe2+ (Fig. 1A, B). In root volatile emissions measurements all functional mutants caused significant reductions in host root ethylene emissions relative to non-inoculated controls (Fig. 5B). This behavior is similar to previously reported results using the endophyte Herbaspirillum seropedicae (strain SmR1) in rice [62]. Past published work suggests that PGPB harboring ACC deaminase, the bacteria enzyme that breaks down ACC, lower host ethylene emissions [36, 63], yet physiology studies consistently predict that lower ethylene levels increase plant sensitivity to environmental stress [64, 65]. While there may be some truth to both sides of this paradox, we believe the story is more complex than a simple relationship between microbial ACC deaminase, plant ethylene regulation and plant growth. For instance, it has been previously documented that some members of the PGPB family Azospirillum sp. do not possess the ACC deaminase gene [66, 67]. This is true of A. brasilense, yet our work demonstrates that root ethylene emissions are significantly reduced with these A. brasilense mutants. Furthermore, we contend that the relationship between microbial ACC deaminase and plant growth promotion may not be generalizable across all PGPB interactions since HM053 A. brasilense was noted to promote plant growth and, in fact, increase crop yield in maize [10]. Instead, we suggest host ethylene regulation could be coupled to microbial indole synthesis, the precursor for auxin. In the present work, measurements of root indole levels showed that all three functional mutants of A. brasilense significantly elevated indole emission rates relative to non-inoculated control plants (Fig. 5C). We suspect this effect could be attributed to the trafficking of bacteria-borne indole into the host root noting that while ipdC is deficient in producing auxin, its upstream metabolic machinery for producing indole should be intact. In fact, ipdC elicited a higher elevation in root indole emission than HM053 or FP10 inoculants.
There are numerous examples in the literature attesting to the complex interaction between ethylene and auxin [49, 68,69,70]. Studies have shown that ethylene stimulates auxin biosynthesis and upregulates the transcription of several auxin transporters including, PIN1, PIN2, AUX1 [71,72,73]. Ethylene-induced auxin production is localized in the root tip [73] and the auxin signal is subsequently redistributed upward through polar auxin transport. We propose that microbial indole infusion in the upper proximal zones of the roots where the bacteria tend to colonize [4] could locally boost auxin biosynthesis in these tissues thereby impairing auxin biosynthesis in the root tip by disrupting its natural redistribution gradient, and thereby causing a reduction in ethylene emission. This theory is supported by the fact that all three functional mutants caused a significant decrease in root gravitropic response – an action that is linked to root tip auxin regulation.
Finally, HM053 and FP10 inoculants significantly elevated levels of root DIMBOA (Fig. 5D) while ipdC did not. The data presented in this figure represents the sum of the endogenous root pool and surface-bound exudate. The influence of HM053 and FP10 on root DIMBOA may, in part be due to the increased indole pool caused by the bacteria, but then the lack of a response from ipdC cannot be explained. A theory here is these functional mutants may influence the partitioning of host carbon resources between DIMBOA and auxin biosynthesis, but that remains to be tested in future radiotracer studies. We note that while DIMBOA is typically considered part of maize’s natural defense mechanism, it can also play a role as a Fe3+ chelating agent for promoting plant uptake of iron [56]. The observed increase in root DIMBOA levels with HM053 and FP10 inoculation may, in part, contribute to observed elevations in 59Fe3+ assimilation with these inoculants.
One question that remains is why A. brasilense would promote production of a bactericide that could potentially be harmful to the microorganism’s health. We note that past work has shown that Pseudomonas putida, another PGPB, was tolerant of DIMBOA [74]. To test the theory that certain functional mutants of A. brasilense were also tolerant of DIMBOA, we conducted a chemotaxis bioassay examining microbial growth at 0 mM, 0.02 mM, 0.1 mM and 0.5 mM doses of DIMBOA added to the growth media. These doses were selected based on earlier work reported with P. putida [74]. In parallel with these assays we conducted a DIMBOA stability test over a 27 hr window (Fig. 6A). Spanning 27 h, DIMBOA underwent only 15% degradation. Results of the chemotaxis assay (Fig. 6B–D) revealed that all three functional mutants tolerated DIMBOA doses of 0.02 mM, but their growth was significantly reduced at higher doses. Using 11C data from the present work (Fig. 4B), we used ~30% as an upper limit of root allocation of fixed 11C over a 3 h period in the presence of either HM053 or FP10 bacteria. These same bacteria resulted in ~1% fixed 11C as acidic exudates over the same 3 h period. Assuming that DIMBOA was the dominant acidic exudate, we estimate that it could account for 0.3% of the total root DIMBOA that was measured in the present work (Fig. 5D) which for HM053 and FP10, could be as high as 10 μg gfw−1 or 0.150 μM accumulated over 3 h. Adjusting for 24 h of accumulation and for 15% loss due to degradation, we estimate DIMBOA exudation in the present system could be as high as 1 μM. That said, all three functional mutants tolerated DIMBOA at ~20-times that level. We also conducted a temporal profile of bacteria growth testing the effects of an acute 0.05 mM dose of DIMBOA over 48 h (Fig. 6E–G). HM053 showed a significant increase in bacteria content even at this acute dose over 48 h while ipdC showed a significant loss in content, and FP10 remained constant (suggesting that its rate of growth matched the rate of cell death). Hence both HM053 and FP10 functional mutants, which were shown to promote DIMBOA levels within the maize roots, appeared to tolerate the substrate.
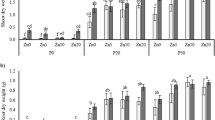
A DIMBOA stability in the bacteria growth medium was examined using 0.5 mM DIMBOA. Gas chromatography peak area units (PAU) reflecting DIMBOA levels at different time points spanning 27 h shows a ~15% loss of DIMBOA integrity. B–D DIMBOA dose response effects on bacteria growth at 24 h as measured spectrophotometrically where OD600 = 1 corresponds to 108 CFU mL−1. E–G Influence of an acute 0.5 mM DIMBOA dose on bacteria growth after 48 h as measured spectrophotometrically where OD600 = 1 corresponds to 108 CFU mL−1. Data (±SE) reflected N = 3 biological replicates at each DIMBOA dose. Asterisks indicate significant differences of bacteria growth with different DIMBOA doses relative to 0 mM DIMBOA control (*P < 0.05; **P < 0.01; ***P < 0.001).
Future studies will examine A. brasilense growth patterns and demand for plant-borne carbon resources as a function of different DIMBOA doses in the growth media. Recently, we published a method that describes using a combination of C-11 radiographic imaging, gamma counting and optical fluorescence imaging with RAM10 Herbaspirillum seropedicae, a green fluorescence reporting strain of an endophytic PGPB, to enable for the first time precise quantification of plant-borne carbon assimilation by a root-associating microorganism [75]. Efforts are underway to generate a stable fluorescent reporting strain of A. brasilense that will enable us to translate this method to this bacterium and test its tolerance for DIMBOA.
Altogether, the results reported herein show that inoculation of maize seedlings with HM053 A. brasilense bacteria boosts host iron assimilation and translocation to aerial portions during early stages of development. This behavior was seen to persist throughout the growing season having beneficial effects on plant growth that resulted in significant increases in crop yield and seed iron content [10]. Thus, the use of beneficial microbes not only influence plant growth, but also can promote crop nutritional content.
Regarding the mechanisms of action, the data presented indicates there are strong influences of bacterial BNF and auxin producing capacities on Fe3+ and Fe2+ assimilation and in planta translocation. While it is enticing to suggest HM053 endows Strategy I iron capabilities in maize, the possibility cannot be ruled out that bacteria metabolism may influence the supplied iron oxidation state. However, we note that in the radiographic images taken of aerial portions from 59Fe3+ and 59Fe2+ exposures in HM053 inoculated plants (Fig. 2E, F) that there was a different spatial patterning of radioisotope across the leaf surfaces. This observation suggests that the iron oxidation states were not altered by the microorganisms. Future studies, however, will examine this aspect more thoroughly using 59Fe3+ and 59Fe2+ coupled with ion chromatography to enable measurement of bacterial uptake and/or manipulation of the different oxidation states of iron as a function of their BNF and auxin biological capabilities.
Supplementary information is available at ISME’s website.
Data availability
All data needed to evaluate the conclusions in the paper are present in the main text or the supplementary materials.
References
Verhagen BW, Glazebrook J, Zhu T, Chang HS, van Loon LC, Pieterse CM. The transcriptome of rhizobacteria-induced systemic resistance in Arabidopsis. Mol Plant Microbe Inter. 2004;17:895–908.
Richardson A, Barea J-M, McNeill A, Prigent-Combaret C. Acquisition of phosphorus and nitrogen in the rhizosphere and plant growth promotion by microorganisms. Plant Soil. 2009;321:305–39.
Fibach-Paldi S, Burdman S, Okon Y. Key physiological properties contributing to rhizosphere adaptation and plant growth promotion abilities of Azospirillum brasilense. FEMS Microbiol Lett. 2012;326:99e108.
Pankievicz VCS, Amaral FP, Santos KFD, Agtuca B, Xu Y, Schueller MJ, et al. Robust biological nitrogen fixation in a model grass-bacterial association. Plant J. 2015;81:907–19.
Brusamarello-Santos LCC, Alberton D, Valdameri G, Camilios-Neto D, Covre R, Lopes K, et al. Modulation of defence and iron homeostasis genes in rice roots by the diazotrophic endophyte Herbaspirillum seropedicae. Sci Rep. 2018;9:10573.
Devendra SD, Prasanna R. Potential of microbes in the biofortification of Zn and Fe in dietary food grains. A review. Agron Sustain Dev. 2020;40:15–36.
Bashan Y, Holguin G, de-Bashan LE. Azospirillum-plant relationships: physiological, molecular, agricultural, and environmental advances. Can J Microbiol. 2004;50:521–77.
Dobbelaere S, Croonenborghs A, Thys A, Ptacek D, Vanderleyden J, Dutto P, et al. Responses of agronomically important crops to inoculation with Azospirillum. Funct Plant Biol. 2001;28:871–9.
Hungria M, Campo R, de Souza E, Pedrosa F. Inoculation with selected strains of Azospirillum brasilense and A. lipoferum; improves yields of maize and wheat in Brazil. Plant Soil. 2010;331:413–25.
Scott S, Housh A, Powell G, Benoit M, Anstaett A, Gerheart A, et al. Azospirillum brasilense (HM053) boosts corn yield and ferritin content. MDPI-Agron J. 2020;10:394–405.
Zuo Y, Zhang F. Soil and crop management strategies to prevent iron deficiency in crops. Plant Soil. 2011;339:83–95.
Samaranayake P, Peiris BD, Dssanayake S. Effect of excessive ferrous (Fe2+) on growth and iron content in rice (Oryza sativa). Int J Agric Biol. 2012;14:296–8.
Mahender A, Swamy BPM, Anandan A, Ali J. Tolerance of iron-deficient and – toxic soil conditions in rice. Plants. 2019;8:31–35.
Curie C, Briat JF. Iron transport and signaling in plants. Annu Rev Plant Biol 2003;54:183–206.
Fageria NK, Baligar VC, Li YC. The role of nutrient efficient plants in improving crop yields in the twenty first century. J Plant Nutr. 2008;31:1121–57.
Römheld V, Marschner H. Mechanism of iron uptake in peanut plants I. Fe reduction, chelate splitting and release of phenolics. Plant Physiol. 1983;71:949–54.
Marschner H, Römheld V. Strategies of plant for acquisition of iron. Plant Soil. 1994;165:261–74.
Jeong J, Guerinot ML. Homing in on iron homeostasis in plants. Trends Plant Sci. 2009;14:280–5.
Cesco S, Neumann G, Tomasi N, Pinton R, Weisskopf L. Release of plant borne flavanoids into the rhizosphere and their role in plant nutrition. Plant Soil. 2010;329:1–25.
Jeong J, Connolly EL. Iron uptake mechanisms in plants: functions of FRO family of ferric reductases. Plant Sci. 2009;176:709–14.
Römheld V, Marschner H. Evidence for a specific uptake system for iron phytosiderophore in roots of grasses. Plant Physiol. 1986;80:175–80.
Römheld V. Different strategies for iron acquisition in higher plants. Plant Physiol. 1987;70:231–4.
Lin YF, Liang HM, Yang SY, Boch A, Clemens S, Chen CC, et al. Arabidopsis IRT3 is a zinc-regulated and plasma membrane localized zinc/iron transporter. N. Phytol. 2009;182:392–404.
Li S, Zhou X, Huang Y, Zhu L, Zhang S, Zhao Y, et al. Identification and characterization of the zinc-regulated transporters, iron-regulated transporter-like protein (ZIP) gene family in maize. BMC Plant Biol. 2013;13:114–28.
Ishimaru Y, Suzuki M, Tsukamoto T, Suzuki K, Nakazono M, Kobayashi T. Rice plants take up iron as an Fe+3-phytosiderophre and as Fe. Plant J. 2006;45:335–46.
Curie C, Panaviene Z, Loulergue C, Dellporta SL, Briat JF, Walker EL. Maize yellow stripe1 encodes a membrane protein directly involved in Fe(III) uptake. Nature. 2001;409:346–9.
Bao Z, Bai J, Cui H, Gong C. A missing link in radial ion transport: Ion transporters in the endodermis. Front Plant Sci. 2019;10:1–8. Article 713.
Barberon M. The endodermis as a checkpoint for nutrients. N. Phytol. 2017;213:1604–10.
Von Wiren N, Klair S, Bansal S, Briat JF, Khodr H, Shioiri T, et al. Nicotianamine chelates both Fe(III) and Fe(II). Implications for metal transport in plants. Plant Physiol. 1999;119:1107–14.
Durrett TP, Gassmann W, Rogers EE. The FRD3-mediated efflux of citrate into the root vasculature is necessary for efficient iron translocation. Plant Physiol. 2007;144:197–205.
Rellán-Álvarez R, Abadía J, Álvarez-Fernández A. Formation of metal-nicotianamine complexes as affected by pH, ligand exchange with citrate and metal exchange. A study by electrospray ionization time-of-flight mass spectrometry. Rapid Commun Mass Sp. 2008;22:1553–62.
Rellán-Álvarez R, Giner Martínez Sierra J, Orduna J, Orera I, Rodríguez Castrillón JÁ, García-Alonso JI, et al. Identification of a tri-iron(III), tri-citrate complex in the xylem sap of iron-deficient tomato resupplied with iron: New insights into plant iron long-distance transport. Plant Cell Physiol. 2009;51:91–102.
Inoue H, Higuchi K, Takahashi M, Nakanishi H, Mori S, Nishizawa NK. Three rice nicotianamine synthase genes, OsNAS1, OsNAS2, and OsNAS3 are expressed in cells involved in long-distance transport of iron and differentially regulated by iron. Plant J. 2003;36:366–81.
Bonneau J, Baumann U, Beasley J, Li Y, Johnson AAT. Identification and molecular characterization of the nicotianamine synthase gene family in bread wheat. Plant Biotechnol J. 2016;14:2228–39.
Kerkeb L, Krämer U. The role of free histidine in xylem loading of nickel in Alyssum lesbiacum and Brassica juncea. Plant Physiol. 2003;131:716–24.
Ravanbakhsh M, Kowalchuk GA, Jousset A. Root-associated microorganisms reprogram plant life history along the growth–stress resistance trade-off. ISME J. 2019;13:3093–101.
Machado HB, Funayama S, Rigo LU, Pedrosa FO. Excretion of ammonium by Azospirillum brasilense mutants resistant to ethylenediamine. Can J Microbiol. 1991;37:549e553.
Schenberger Santos AR, Etto RM, Furmam RW, de Freitas DL, d’Eça Nogueira Santos KF, de Souza EM, et al. Labeled Azospirillum brasilense wild type and excretion-ammonium strains in association with barley roots. Plant Physiol Biochem. 2017;118:422e426.
Pedrosa FO, Yates MG. Regulation of nitrogen fixation (nif) genes of Azospirillum brasilense by nifA and ntr (gln) type gene products (Azospirillum brasilense; Nif- mutants; nifA; ntrC; MoFe protein). FEMS Microbiol Lett. 1984;23:95–101.
Näsvall J, Knöppel A, Andersson DI. Duplication-Insertion Recombineering: a fast and scar-free method for efficient transfer of multiple mutations in bacteria. Nucleic Acids Res. 2017;45:e33.
de Souza EM, private communication.
Ferrieri RA, Wolf AP. The chemistry of positron-emitting nucleogenic (hot) atoms with regard to preparation of labeled compounds of practical utility. Radiochim Acta. 1983;34:69–83.
Ferrieri RA. Production and application of synthetic precursors labeled with carbon-11 and fluorine-18. In Welch MJ, Redvanly CS, editors. Handbook of radiopharmaceuticals: radiochemistry and applications. Chichester, England: John Wiley & Sons, Ltd.; 2003.
Ferrieri RA, Gray DW, Babst BA, Schueller MJ, Schlyer DJ, Thorpe MR, et al. Use of carbon-11 in Populus shows that exogenous jasmonic acid increases biosynthesis of isoprene from recently fixed carbon. Plant Cell Environ. 2005;25:591–602.
Qu W, Robert CAM, Erb M, Hibbard BE, Paven M, Gleede T, et al. Dynamic precision phenotyping reveals mechanisms of crop tolerance to root herbivory. Plant Physiol. 2016;172:776–88.
Klaus R, Fisher W, Hauck HE. Application of a thermal in situ reaction for fluorometric detection of carbhhydrates on NH2-layers. Chromatographia. 1990;29:467–72.
Babst BA, Karve A, Judt T. Radio-metabolite analysis of carbon-11 biochemical partitioning to nonstructural carbohydrates for integrated metabolism and transport studies. Plant Cell Physiol. 2013;54:1016–25.
Lee SJ, Fowler JS, Alexoff DL, Schueller MJ, Kim D, Nauth AA. et al. An efficient and practical synthesis of [2-11C]indole via superfast nucleophilic [11C]cyanation and RANEY® Nickel catalyzed reductive cyclization. Org. Biomol. Chem.2015;13:11235–43.
Onidasa D, Stachnik JM, Brucker S, Kratzig S, Gerwert K. Histidine is involved in coupling proton uptake to electron transfer in photosynthetic proteins. Eur J Cell Biol. 2010;89:983–9.
Ingle RA. Histidine biosynthesis. In the Arabidopsis book. The American Society of Plant Biologists; Washington, D.C.: 2011. https://doi.org/10.1199/tab.0141.
Connorton JM, Balk J, Rodríguez-Celma J. Iron homeostasis in plants - a brief overview. Metallomics. 2017;9:813–23.
Frey M, Spiteller D, Boland W, Gierl A. Transcriptional activation of Igl, the gene for indole formation in Zea mays: a structure-activity study with elicitor-active N-acyl glutamines from insects. Phytochemistry. 2004;65:1047–55.
Frey M, Chomet P, Glawischnig E, Stettner C, Grün S, Winklmair A, et al. Analysis of a chemical plant defense mechanism in grasses. Science. 1997;277:696–9.
Frey M, Schullehner K, Dick R, Fiesselmann A, Gierl A. Benzoxazinoid biosynthesis, a model for evolution of secondary metabolic pathways in plants. Phytochemistry. 2009;70:1645–51.
Zhou S, Richter A, Jander G. Beyond defense: multiple functions of benzoxazinoids in maize metabolism. Plant Cell Physiol. 2018;59:1528–37.
Hu L, Mateo P, Ye M, Zhang X, Berset JD, Handrick V, et al. Plant iron acquisition strategy exploited by an insect herbivore. Science. 2018;361:694–7.
Frey M, Stettner C, Paré PW, Schmelz EA, Tumlinson JH, Gierl A. An herbivore elicitor activates the gene for indole emission in maize. Proc Nat’l Acad Sci USA. 2000;97:14801–6.
Erb M, Veyrat N, Robert CAM, Xu H, Frey M, Ton J, et al. Indole is an essential herbivore-induced volatile priming signal in maize. Nat Commun. 2015;16:6273–83.
Band LR, Wells DM, Larrieu A, Sun J, Middleton AM, French AP, et al. Root gravitropism is regulated by a transient lateral auxin gradient controlled by a tipping-point mechanism. Proc Nat Acad Sci USA. 2012;109:4668–73.
Madhaiyan M, Poonguzhali S, Sa T. Characterization of 1-aminocyclopropane-1-carboxylate (ACC) deaminase containing Methylobacterium oryzae and interactions with auxins and ACC regulation of ethylene in canola (Brassica campestris). Planta. 2007;226:867–76.
Zhang Z, Barlow JN, Baldwin JE, Schofield CJ. Metal-catalyzed oxidation and mutagenesis studies on the iron(II) binding site of 1-aminocyclopropane-1-carboxylate oxidase. Biochemistry. 1997;36:15999–6007.
Alberton D, Müller-Santos M, Brusamarello-Santos LCC, Valdameri G, Cordeiro FA, Yates MG, et al. Comparative proteomics analysis of the rice roots colonized by Herbaspirillum seropedicae strain SmR1 reveals induction of the methionine recycling in the plant host. J Proteome Res. 2013;12:4757–4768.
Glick BR. Modulation of plant ethylene levels by the bacterial enzyme ACC deaminase. FEMS Microbiol Lett. 2005;251:1–7.
Thao NP, Khan MIR, Thu NBA, Hoang XLT, Asgher M, Khan NA, et al. Role of ethylene and its cross talk with other signaling molecules in plant responses to heavy metal stress. Plant Physiol. 2015;169:73–84.
Keunen E, Schellingen K, Vangronsveld J, Cuypers A. Ethylene and metal stress: small molecule, big impact. Front Plant Sci. 2016;7:23.
Kende H. Ethylene biosynthesis. Annu Rev Plant Physiol Plant Mol Biol. 1993;44:283–307.
Holguin G, Glick BR. Expression of the ACC deaminase gene from Enterobacter cloacae UW4 in Azospirillum brasilense. Micro Ecol. 2001;41:281–8.
Muday GK, Rahman A, Binder BM. Auxin and ethylene: collaborators or competitors? Trends Plant Sci. 2012;17:181–95.
Van de Poel B, Smet D, Van Der Straeten D. Ethylene and hormonal crosstalk in vegetative growth and development. Plant Physiol. 2015;169:61–72.
Hu Y, Vandenbussche F, Van Der Straeten D. Regulation of seedling growth by ethylene and the ethylene-auxin crosstalk. Planta. 2017;245:467–89.
Ruzicka K, Ljung K, Vanneste S, Podhorská R, Beeckman T, Friml J, et al. Ethylene regulates root growth through effects on auxin biosynthesis and transport-dependent auxin distribution. Plant Cell. 2007;19:2197–212.
Stepanova AN, Yun J, Likhacheva AV, Alonso JM. Multilevel interactions between ethylene and auxin in Arabidopsis roots. Plant Cell. 2007;19:2169–85.
Swarup R, Perry P, Hangenbeek D, Van Der Straeten D, Beemster GT, Sandberg G, et al. Ethylene upregulates auxin biosynthesis in Arabidopsis seedlings to enhance inhibition of root cell elongation. Plant Cell. 2007;19:2186–96.
Neal AL, Ahmad S, Gordon-Weeks R, Ton J. Benzoxazinoids in root exudates of maize attract pseudomonas putida to the rhizosphere. PLoS ONE. 2012;7:e35498.
Waller S, Wilder SL, Schueller MJ, Housh AB, Ferrieri RA. Quantifying plant-borne carbon assimilation by root-associating bacteria. Microorganisms. 2020;8:700–10.
Acknowledgements
We would like to thank DeAna Grant (University of Missouri, Electron Microscopy Core) for assisting in sample preparation and transmission electron microscopy analysis. We would also like to thank Prof. Emanuel de Souza (U. Paraná, Brazil) whose institution provided the functional mutants of A. brasilense in this work.
Funding:
This research was supported by Agriculture and Food Research Initiative Award No. 2017-67013-26216 from the USDA National Institute of Food and Agriculture. We also acknowledge the National Science Foundation for the award BCS-0922374 to the University of Missouri, which funded the ICP-MS instrument used in this research.
Author information
Authors and Affiliations
Contributions
Experimental design and supervision - RAF, MJS; data collection and analysis - ABH, GP, AP, SS, AA, AG, SW, MB, SLW, JMG, BH; initial draft - RAF, ABH, SLW, MJS.; final draft - all authors.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Housh, A.B., Powell, G., Scott, S. et al. Functional mutants of Azospirillum brasilense elicit beneficial physiological and metabolic responses in Zea mays contributing to increased host iron assimilation. ISME J 15, 1505–1522 (2021). https://doi.org/10.1038/s41396-020-00866-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41396-020-00866-x
This article is cited by
-
Enhancement of beneficial microbiomes in plant–soil continuums through organic fertilization: Insights into the composition and multifunctionality
Soil Ecology Letters (2024)
-
Development of ACCd producer A. brasilense mutant and the effect of inoculation on red pepper plants
3 Biotech (2022)
-
Effect of Plant growth promoting rhizobacteria (PGPR) and mycorrhizal fungi inoculations on essential oil in Melissa officinalis L. under drought stress
Biologia (2022)
-
Insights on plant–microbe interactions in soil in relation to iron dynamics
Vegetos (2022)
-
Zinc use efficiency of maize-wheat cropping after inoculation with Azospirillum brasilense
Nutrient Cycling in Agroecosystems (2021)