Abstract
Background
Prostate cancer (PCa) shows racial disparity in clinical and genomic characteristics, and Asian patients with PCa often present with more aggressive phenotypes at diagnosis. The ability of TP53 to serve as a prognostic biomarker of PCa has been well studied in Western populations. However, no studies to date have examined the role of TP53 in the disparities of primary hormone-naïve prostate cancer (HNPC) between Chinese and Western populations.
Methods
We collected prostate tumors and matched normal tissues or blood samples to perform targeted next-generation sequencing of 94 Chinese primary localized HNPC samples, and correlated these genomic profiles with clinical outcomes. The OncoKB knowledge database was used to identify and classify actionable alterations.
Results
The aberrations of PTEN, CDK12, and SPOP in Chinese HNPC samples were similar to those in the Western samples. However, we demonstrated an association of a high frequency of TP53 alterations (21/94) with a relatively higher percentage of alterations in the Wnt signaling pathway (15/94) in Chinese HNPC. Additionally, we highlighted alterations of LRP1B as accounting for a high proportion of PCa and found more frequent alterations in CDH1 in Chinese PCa. Of these, only CDH1 alteration was associated with rapid biochemical recurrence (BCR). However, we verified that TP53 status was at the core of the genomic alteration landscape in Chinese HNPC with putative driver mutations because of the strong connections with other signaling pathways. The mutually exclusive relationship between alterations in TP53 and Wnt/CTNNB1 further molecularly characterizes subsets of prostate cancers. Moreover, the alteration of KMT2C was more likely to co-occur with TP53 alteration, indicating a more aggressive phenotype of PCa, which was associated with sensitivity to treatment with poly ADT–ribose polymerase (PARP) inhibitors.
Conclusions
Detection of TP53 alterations has clinical utility for guiding precision cancer therapy for HNPC, especially in the Chinese population.
Similar content being viewed by others
Introduction
Prostate cancer (PCa) is the most common cancer followed by lung cancer in men in 2020, accounting for more than 20% of new male cancer diagnoses in the US, and there will be 33,330 PCa-related deaths in the US [1]. Although the majority of new cases are diagnosed with localized PCa, 4% of patients are diagnosed with metastatic PCa at their first presentation, especially in unscreened populations [2].
With dramatic economic growth and westernized lifestyle changes, the incidence rate of PCa has increased rapidly in China with an annual percentage change of 12.6% since 2000 [3]. Thirty percent of Chinese patients are diagnosed with metastatic PCa, and this phenomenon that an aggressive phenotype of PCa at diagnosis is more common in Asian than in Western population was confirmed in other Asian countries [4, 5]. Currently, second-generation androgen receptor signaling inhibitors (e.g., abiraterone and enzalutamide) and chemotherapy (e.g., cabazitaxel and docetaxel) are mainstays of treatment for advanced PCa [6,7,8]. Unfortunately, after the initial response, most patients develop secondary resistance, along with 10–40% patients who exhibit primary resistance [2].
As a key tumor suppressor gene (TSG), TP53 plays a pivotal role in genomic stability, cell cycle arrest, and other important signaling pathways [9]. TP53 mutation is one of the most common alternations, affecting 50% of metastatic PCa cases [10]. Many studies have demonstrated that TP53 status has the prognostic clinical significance in castration-resistant prostate cancer (CRPC), and acts as a biomarker of poor response to novel hormonal therapies (abiraterone and enzalutamide) [11, 12]. However, studies of TP53 status in hormone-naïve prostate cancer (HNPC) are rare, especially in the Chinese population.
We performed targeted next-generation sequencing of 94 prostate cancer samples to molecularly characterize prostate cancer in the Chinese population, especially the role of TP53. Together, these data provide insights into the genomic landscape of Chinese primary PCa and the important role of TP53 in HNPC. We also highlight molecular aberrations that may help to triage patients for precision prostate cancer therapy.
Materials and methods
Patients and samples
We included a cohort of 293 patients with PCa who had undergone radical prostatectomy or prostatic biopsy from January 2018 to March 2019 at Qi Lu Hospital of Shandong University. Prostate tumor and matched normal tissues or blood samples were collected under the experimental protocols approved by the Institutional Ethics Review Committee of Shandong University. We excluded patients who received neoadjuvant androgen deprivation therapy (ADT) and those who were lost to follow-up. We ultimately included 101 patients (94 localized HNPC, 7 metastatic HNPC) in our study, and all patients had complete clinical and follow-up records, including age, PSA, Gleason score, pathological stage, and National Comprehensive Cancer Network (NCCN) risk group. Informed consent was obtained from all participants. All tumor tissues were reviewed by two independent pathologists, and samples with estimated tumor purity >20% on histopathologic assessment were subjected to genomic profiling.
Targeted next-generation sequencing and genetic analysis
At least 50 ng of genomic DNA from patient samples in our cohort were subjected to the hybridization capture-based next-generation sequencing panel, which contained all exons of 450 cancer-related genes and selected introns of 36 genes (Supplementary Table S1). The captured libraries were sequenced on an Illumina NextSeq-500 Platform (Illumina Incorporated) and the genomic alterations were analyzed. The genomic alteration profile is shown in Supplementary Tables S2 and S3.
Identification and classification of actionable alterations
The actionability of genetic alterations was determined according to the OncoKB knowledge database (https://www.oncokb.org/), in which the actionable alterations were classified as level 1, 2A/B, 3A/B, and 4. In our study, we defined alterations within levels 1–4 as translational targets, while levels 1-2B as actionable targets.
Statistical analysis
All analyses were performed using R version 3.4.2. Continuous variables are expressed as the mean ± SD if they were normally distributed, otherwise as the median with interquartile ranges are presented. The R package “rcompanion” was used to conduct Fisher’s exact test or the χ2 test and post hoc tests for comparisons of co-mutation and exclusion analysis. The rapid biomedical recurrence is biomedical failure within 18 months, and Fisher’s exact test or the χ2 test was performed to examine the association of gene aberrations with rapid BCR. All reported P values were two-tailed, and P < 0.05 was considered statistically significant.
Results
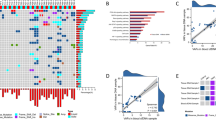
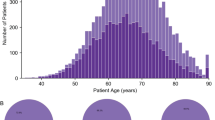
A total of 101 HNPC patients were enrolled in our study, including 94 patients who underwent radical prostatectomy (RP) with primary localized PCa and 7 metastatic tumor biopsies (bone (N = 3), lung (N = 2), liver (N = 1) and lymph node (N = 1)). We ultimately included 94 localized HNPCs for analysis given the heterogeneity between primary and metastatic PCa. All 94 paired samples were received from patients without ADT who were pathologically diagnosed with prostate adenocarcinoma, with a median follow-up of 20 months. The median age of 94 patients at diagnosis was 68 years old, and the median prostate-specific antigen (PSA) level was 16.94 ng/ml. Other clinical and pathological characteristics are summarized in Table 1. To illustrate the genomics of Chinese HNPC, we performed a targeted next-generation sequencing panel that captured mutations in coding regions of 450 cancer-related genes, including the key TSGs: TP53, PTEN, RB1, and other prostate cancer-related genes, such as AR and DNA damage repair (DDR) genes (Supplementary Table S1). The most frequently aberrant genes in our cohort included TP53 (22.3%), ERG (18.1%), SPOP (17.0%), BRCA2 (7.4%), APC (6.4%), CDH1 (5.3%) and LRP1B (5.3%) (Fig. 1a).
a Frequency of genomic alterations. All the type of ERG fusions were TMPRSS2-ERG fusions, the other TMPRSS2 fusion occurred between TMPRSS2 and KLF12, PDE9A, MIR99AHG, and SLC5A4. b Mutational signatures of TP53 in prostate cancer. TAD Transactivation domain, DBD DNA-binding domain, OD Oligomerization domain.
TP53 alterations
In our study, 21/94 (22.3%) of cases harbored TP53 alterations, and the TP53 alteration rate was higher than that reported in other studies (3–12.5%) [13,14,15]. This difference in TP53 alteration between Chinese and Western populations corresponds with the phenomenon that advanced PCa at diagnosis is more common in Asians [5]. In our study, the TP53 alteration types were substitutions/indels (73.9%) and truncation (26.1%) (Fig. 1a). As shown in Fig. 1b, the majority of TP53 alterations occurred in the region of the central core sequence-specific DNA-binding domain (DBD), which was similar to the results reported in prior studies [16]. There was one patient who carried two different TP53 alterations (Patient 023: V157F, E287Rfs*58) in our study, and the most common mutation types were R175H (2/22), R273C/H (2/22), and R337L (2/22), which were also the most commonly explored substitution type of TP53 mutation, showing oncogenic gain of function(GOF) in the initiation and progression of cancer [9].
AR alterations
In our study, only 1/94 localized HNPC sample harbored AR alteration, and in aggregate, 2/7 of cases harbored AR alterations in metastatic HNPCs (Supplementary Table 3). All three AR alterations were AR amplifications. This situation related to AR aberrations in our study further demonstrates that AR alterations are uncommon in primary hormone-naïve disease but have a higher rate in patients with metastatic prostate cancer [17].
Wnt signaling pathway
In total, 15/94 (16%) of cases harbored aberrations in the Wnt signaling pathway, including 6 cases with APC inactivating mutations and 4 cases with activating mutations in CTNNB1 (Fig. 3a). Thus, patients with HNPC in our cohort had a higher alteration frequency for the Wnt signaling pathway than observed in a Western study (7%) [18]. Moreover, the alteration frequency for the Wnt signaling pathway in our primary localized HNPC cases (15/94) was similar to that reported in metastatic castration-resistant prostate cancer (mCRPC, 27/150) [10], further demonstrating that Asian patients have more aggressive phenotypes of PCa at diagnosis than Western patients.
Interestingly, we found that alterations in the Wnt signaling pathway were rarely observed in conjunction with TP53 mutation, and there was a mutually exclusive relationship between TP53 alteration and CTNNB1 mutation (Fig. 2a, b).
DNA damage repair pathway
In total, 21/94 (22.3%) of cases harbored at least one alteration in the DNA damage repair (DDR) pathway. The most affected DDR subtypes were homologous recombination (HR) and mismatch repair (MMR) (Fig. 2a). The most frequently mutated DDR gene was BRCA2, followed by ATM (Fig. 3b). Furthermore, the mutation rate of BRCA2 was 7/94 of HNPC in our cohorts, which corresponded with prior Western studies [14, 18]. Most importantly, 13/94 (BRCA2 N = 7, BRCA1 N = 3, ATM N = 4, one case harboring both BRCA2 alteration and ATM alteration) of cases would benefit from treatment with poly ADT–ribose polymerase (PARP) inhibitors [19]. It is well established that patients with mismatch-repair deficiency (dMMR) or microsatellite instability-high (MSI-H) have a good response to immune checkpoint–inhibiting therapies [20]. In total, 5/94 (5.3%) of cases harbored at least one alteration in MMR, including two patients harboring MSH6 alterations, two patients harboring MLH1 alterations and one patient harboring alterations of both MSH6 and MSH2 (Fig. 2a). The prevalence of dMMR in our localized HNPC was higher than that in the Western population (9/333) and appears to be similar to that in metastatic prostate cancers (3.1–8.1%) [15, 21, 22]. Through integrative analysis, we found co-occurrence of BRCA2 mutation and LIMK1 alternation (Fig. 2b), and high expression of LIMK1 in prostate cancer indicates a poor outcome [23].
a Mutational signatures of APC/CTNNB1. ARM Armadillo/beta-catenin-like repeat. b Mutational signatures of BRCA 1/2 and ATM. HELC helical domain, OB oligosaccharide-binding domain, zf-C3HC4 Zinc finger, C3HC4 type, BRCT_assoc Serine-rich domain associated with BRCT, EIN3 Ethylene insensitive 3 domain, BRCT BRCA1 c-terminus domain, TAN Telomere-length maintenance and DNA damage repair, FAT FRAP-ATM-TRRAP domain, PI3_I4K phosphatidylinositol 3- and 4-kinase, FATC FAT c-terminal domain. c Mutational signatures of SPOP. MATH MATH domain, BTB Broad Complex, Tramtrack and Bric a brac/poxvirus and zinc finger domain. d Mutational signatures of CDH1. Cadherin_pro Caderin prodomain like, Cadherin_C Cadherin cytoplasmic region. e Mutational signatures of LRP1B.
Germline DDR mutations, especially loss-of function BRCA2 mutations, are found primarily in prostate cancer presenting a more aggressive phenotype and a poor prognosis [24]. In our study, we found germline mutations in BRCA2, ATM, CHEK2, RAD50, and RAD50D, among which BRCA2 (5/7) harbored the most frequent germline mutation.
Other key gene alterations in prostate cancer
The mutation in SPOP, the substrate-binding pocket of the E3 ubiquitin ligase adapter, was present in 6–15% of primary prostate cancers as previously reported in Western cohorts [25]. In total, 16/94 (17.0%) of cases harbored SPOP mutations in our cohort, which was similar to report of other cohort [25]. Moreover, as shown in Fig. 3c, all these SPOP mutations were substitutions, and no SPOP mutations were found in metastatic HNPC in our cohort (Supplementary Table 3), which corresponded with the interesting finding that SPOP mutations were less frequent in metastatic PCa than in primary cases [15].
Strikingly, we found a mutually exclusive relationship between SPOP mutation and TP53 alteration (Fig. 2b). We also found strong mutual exclusivity between SPOP mutation and alterations in TMPRSS2-ERG fusion as in a prior study (Fig. 2b) [25]. This further demonstrates that the SPOP mutation represents a molecular subtype of PCa [15].
CDH1, encoding the protein E-cadherin, was one of the top 15 genes harboring alterations in our cohort, and the most alteration subtype was truncation (Figs. 1a and 3d). Moreover, we found more frequent CDH1 alterations than that in prior studies (5.3% in our localized HNPC, 0.9–2.9% in Western primary PCa) [25, 26].
Low-density lipoprotein receptor-related protein 1B (LRP1B), a tumor suppressor gene encoding an endocytic LDL-family receptor, received our attention because it had a relatively high mutation rate in our cohort. There were 5/94 cases harboring LRP1B alterations which were similar to the 2–12% of primary PCa cases reported in other studies [13, 15]. As shown in Fig. 3e, the alteration subtypes were substitutions and gene homozygous deletions. Furthermore, LRP1B alterations are more commonly presented in mCRPC, with ~9–18% patients affected [10, 27].
Actionable alterations
In our study, as shown in Fig. 4a, 48/94 (51.1%) cases harbored at least one targeted alteration with any level of OncoKB recommendations [28], including 8 cases in the TP53 mutant group and 40 cases in the TP53 wild-type group. More importantly, 29/94 (30.9%) cases harbored actionable alterations including OncoKB recommendations of level 1-level 2B. The proportion of actionable alterations in the TP53 mutant group (6/21) was similar to that in the TP53 wild-type group (23/73) (Fig. 4a). However, most actionable alterations (5/8) were recommended as level 1 in the TP53 mutant group but only (13/40) in the TP53 wild-type group. As shown in Fig. 4b, c, the actionable alterations were oncogenic mutations of BRCA2, ATM, CHEK2, and PIK3CA in the TP53 mutant group, while the TP53 wild-type group harbored more other actionable alterations (e.g., BRCA1, RAD51D) except for these four genes in the TP53 mutant group.
a The upper pie-plot indicates the frequency of patients with TP53mut PCa (N = 21) or TP53wt PCa (N = 73) who were identified with translational targets in our cohort. The lower pie-plot shows the distribution of OncoKB levels for translational targets in patients with TP53mut or TP53wt PCa. b The flow diagram in the left part shows the list of translational targets for each OncoKB recommendation level in TP53 mutant group, and the right part presents for TP53 wild-type PCa. The colors of the curving belts represent different signaling pathways, and the widths of the belts indicate different frequencies for each target at every level. c The panel shows the comparison of actionable alteration frequencies between TP53mut and TP53wt PCa.
Association of genomic alterations with clinical outcomes
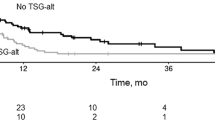
Although our study included patients registered in our center from January 2018 to March 2019, we still obtained a median follow-up time of 20 months. There were 29 cases with biochemical recurrence (BCR, PSA nadir + 0.2 ng/mL), including bone metastasis in 4 cases. Rapid biochemical recurrence is biochemical failure within 18 months of primary local therapy, and it is a marker of poor prognosis of prostate cancer [29, 30]. There were 29/94 (30.9%) cases suffering rapid biochemical recurrence in our study, which is higher than reported in the Western population (~20%) [29, 30]. We then examined the association of gene aberrations with rapid BCR (Table 2). In our study, we found an association of CDH1 alteration with rapid BCR. However, there were no associations of aberrations of TP53, PTEN, BRCA2 with rapid BCR, although these aberrations were risk factors for BCR in prior studies [31,32,33]. These different conclusions between our study and prior studies may be attributed to the small sample size in our study, and further studies are needed to validate the association of genomics with clinical outcomes of HNPC.
Discussion
PCa is a typical cancer harboring not only intratumor heterogeneity but also intertumor heterogeneity, especially in patients with different racial and ethnic backgrounds [34]. The heterogeneity of PCa could explain the phenomenon that Asians have a more aggressive phenotype of PCa than individuals from Western countries at diagnosis. However, the underlying molecular mechanism is unclear. Genomic molecular alterations in Asians should be explored in detail to find actionable alterations, thus guiding precision cancer therapy.
In our study, the most frequent alteration was TP53, with 21/94 of individuals affected, strongly implying a pivotal role in the carcinogenesis and progression of PCa [35]. The most commonly mutated region of TP53 was DBD, and R175H, R273C/H, and R337L were the most frequently mutated subtypes. Unlike to R175H and R273C/H, which were localized at DBD and their oncogenic GOF were comprehensively explored, studies on R337L mutation of TP53 are rare. Kunimasa et al. discovered that lung cancers harboring the TP53 R337L mutation have poor outcomes [36]. However, there are no studies focusing on the underlying mechanism of TP53 R337L. As shown in Fig. 1b, TP53 R337L occurred at the region of the oligomerization domain, which is mainly involved in ubiquitination of TP53 [37]. Therefore, we hypothesized that TP53 R337L plays an important role in the stability and subcellular localization of TP53, as the ubiquitination status of TP53 (monoubiquitination or polyubiquitination) is a sure sign of its distinct biological function [38]. Thus, the biological function of TP53 R337L warrants investigation in the cancer model systems.
Moreover, unlike the primary HNPCs in the West, which harbor infrequent alterations in the Wnt pathway, our data identified a subset of localized HNPCs (16%) in Chinese patients that harbored aberrations in the Wnt signaling pathway. The Wnt alteration prevalence in our cohort appears similar to that of metastatic prostate cancer in Western populations [10, 39]. Activated Wnt signaling is more common in high-risk PCa and promotes cell proliferation in an androgen-independent manner [40]. These genomic differences in the Wnt signaling pathway could partly explain the phenomenon that Asian patients have more aggressive phenotypes of PCa at diagnosis than Western patients. Furthermore, the mutation of CTNNB1 had a mutually exclusive relationship with alterations of TP53, representing two distinct molecular subsets of PCa in the Chinese population. More interestingly, aberrations in the Wnt pathway act as biomarkers of the response to immunotherapy in cancers. Pinyol et al. proved that alterations in the Wnt signaling pathway, especially mutations in CTNNB1, represent a “cold tumor” for immunotherapy, which means that these cases harboring Wnt pathway alterations develop resistance to immunotherapy [41]. Linch et al. found that patients with activated Wnt/CTNNB1 mutations in PCa have a low CD8+/FOXP3+ ratio in the tumor microenvironment, suggesting immune evasion [42].
Interestingly, we found a high frequency of LRP1B alterations in our cohort, which was confirmed as the top mutated gene by other studies not only in primary PCa but also in mCRPC [10, 15]. In addition to PCa, LRP1B, as the significant TSG, has a high frequency of mutations in lung cancer and colorectal cancer [43, 44]. However, few studies have elucidated the biological function of LRP1B mutations. Only a few studies have proposed that LRP1B mutations, always co-occurring with a high tumor mutation burden, may indicate a good response to immunotherapy [45, 46]. The significant prognostic value of LRP1B mutations will need to be prospectively assessed in clinical trials.
Through integrative analysis, we found that TP53 status was at the core of the genomic alteration landscape in Chinese HNPC with putative driver mutations. First, TP53 alterations (21/94) were the most frequent alteration in our Chines e HNPC cohort. Second, TP53 alteration was highly connected with other signaling pathways. There was a mutually exclusive relationship between alterations in TP53 and SPOP (Fig. 2a, b). The highest level of AR activity is present in the subtype of SPOP mutation, indicating a good response to ARSI, while TP53 alterations, relating to reduced tumor dependency on AR signaling, are associated with resistance to the ARSI [47]. Furthermore, there was also a mutually exclusive relationship between alterations in TP53 and Wnt/CTNNB1, which promoted PCa progression in an androgen-independent manner [40]. Additionally, there was a co-occurring relationship between alterations in KMT2C and TP53, and the mutation of KMT2C is common in PCa [48]. Rampias et al. demonstrated that KMT2C could regulate DDR genes, and patients with KMT2C mutation are more sensitive to treatment with PARP inhibitors [49].
We noticed that the prevalence of gene aberrations in our Chinese cohort was different from that in the Ren et al. study [5]. The commonly mutated genes in the Ren et al. Chinese cohort were SPOP (9.52%), TP53 (3.33%), ATM (2.86%), and CTNNB1 (0.95%), and the mutation frequencies were lower than those in our cohort, especially for TP53 (22.3%) and CTNNB1 (4.3%). Moreover, the frequency of TMPRESS2-ERG fusion in our cohort is two times that in the Ren et al. cohort (18.1% vs 9.2%), both of which are much lower than those in the Western population (~50%) [50]. Through analysis of the characterizations of enrolled cases, we found that the proportion of the NCCN high/very high-risk group in our cohort was much higher than that in Ren et al. study (75.72% vs 66.52%). Another plausible explanation for the genomic differences between these two studies is the different geographic distributions of the study population. Patients in our study come from Shandong Province, northern China, while patients in the Ren et al. study come from Shanghai, southern China. Shanghai is the most developed city in China, in which people have a more westernized lifestyle, and it is truly different from Shandong Province. Moreover, we also found this heterogeneity in prostate cancer caused by geographic distribution between mainland China and Hong Kong [4].
There were several limitations in our study that need to be considered. First, the patients enrolled in our cohort came from a single center, which is not representative of patients nationally. Furthermore, due to the relatively small sample size in our study, bias could not be eliminated completely and further studies enrolling a large sample size from different centers in China should be performed to validate the genomic characteristics of HNPC in Chinese men. In addition, our next-generation sequencing panel contained only exons of 450 cancer-related genes and selected introns of 36 genes, which led to data loss of other gene aberrations and other types of variants, such as genomic structural variants (SVs) involved in the introns.
Overall, this study provides new insight into the genomic alterations that characterize HNPC in the Chinese population. More importantly, our data on HNPC indicate that it is essential to perform further targeted NGS for precision treatment after finding significant TP53 alterations in PCa.
References
Rebecca LS, Kimberly DM, Ahmedin J. Cancer statistics, 2020. CA Cancer J Clin. 2020;70:7–30.
Buttigliero C, Tucci M, Bertaglia V, Vignani F, Bironzo P, Di Maio M, et al. Understanding and overcoming the mechanisms of primary and acquired resistance to abiraterone and enzalutamide in castration resistant prostate cancer. Cancer Treat Rev. 2015;41:884–92.
Chen W, Zheng R, Baade PD, Zhang S, Zeng H, Bray F, et al. Cancer statistics in China, 2015. CA Cancer J Clin. 2016;66:115–32.
Chen R, Ren S, Yiu MK, Fai NC, Cheng WS, Ian LH. Chinese Prostate Cancer Consortium et al. Prostate cancer in Asia: a collaborative report. Asian J Urol. 2014;1:15–29.
Ren S, Wei GH, Liu D, Wang L, Hou Y, Zhu S, et al. Whole-genome and transcriptome sequencing of prostate cancer identify new genetic alterations driving disease progression. Eur Urol. 2018;73:322–39.
Beer TM, Armstrong AJ, Rathkopf DE, Loriot Y, Sternberg CN, Higano CS, et al. Enzalutamide in metastatic prostate cancer before chemotherapy. N. Engl J Med. 2014;371:424–33.
de Bono JS, Logothetis CJ, Molina A, Fizazi K, North S, Chu L, et al. Abiraterone and increased survival in metastatic prostate cancer. N. Engl J Med. 2011;364:1995–2005.
Oudard S, Fizazi K, Sengeløv L, Daugaard G, Saad F, Hansen S, et al. Cabazitaxel versus docetaxel as first-line therapy for patients with metastatic castration-resistant prostate cancer: a randomized phase III trial-FIRSTANA. J Clin Oncol. 2017;35:3189–97.
Muller PA, Vousden KH. Mutant p53 in cancer: new functions and therapeutic opportunities. Cancer Cell. 2014;25:304–17.
Robinson D, Van Allen EM, Wu YM, Schultz N, Lonigro RJ, Mosquera JM, et al. Integrative clinical genomics of advanced prostate cancer. Cell. 2015;161:1215–28.
Hamid AA, Gray KP, Shaw G, MacConaill LE, Evan C, Bernard B, et al. Compound genomic alterations of TP53, PTEN, and RB1 tumor suppressors in localized and metastatic prostate cancer. Eur Urol. 2019;76:89–97.
De Laere B, Oeyen S, Mayrhofer M, Whitington T, van Dam PJ, Van Oyen P, et al. TP53 outperforms other androgen receptor biomarkers to predict abiraterone or enzalutamide outcome in metastatic castration-resistant prostate cancer. Clin Cancer Res. 2019;25:1766–73.
Fraser M, Sabelnykova VY, Yamaguchi TN, Heisler LE, Livingstone J, Huang V, et al. Genomic hallmarks of localized, non-indolent prostate cancer. Nature. 2017;541:359–64.
Beltran H, Yelensky R, Frampton GM, Park K, Downing SR, MacDonald TY, et al. Targeted next-generation sequencing of advanced prostate cancer identifies potential therapeutic targets and disease heterogeneity. Eur Urol. 2013;63:920–6.
Cancer Genome Atlas Research Network. The molecular taxonomy of primary prostate cancer. Cell. 2015;163:1011–25.
Bode AM, Dong Z. Post-translational modification of p53 in tumorigenesis. Nat Rev Cancer. 2004;4:793–805.
Taplin ME, Bubley GJ, Shuster TD, Frantz ME, Spooner AE, Ogata GK, et al. Mutation of the androgen-receptor gene in metastatic androgen-independent prostate cancer. N. Engl J Med. 1995;332:1393–8.
Mateo J, Seed G, Bertan C, Rescigno P, Dolling D, Figueiredo I, et al. Genomics of lethal prostate cancer at diagnosis and castration resistance. J Clin Invest. 2020;130:1743–51.
de Bono J, Mateo J, Fizazi K, Saad F, Shore N, Sandhu S, et al. Olaparib for metastatic castration-resistant prostate cancer. N. Engl J Med. 2020;382:2091–102.
Le DT, Uram JN, Wang H, Bartlett BR, Kemberling H, Eyring AD, et al. PD-1 blockade in tumors with mismatch-repair deficiency. N. Engl J Med. 2015;372:2509–20.
Abida W, Cheng ML, Armenia J, Middha S, Autio KA, Vargas HA, et al. Analysis of the prevalence of microsatellite instability in prostate cancer and response to immune checkpoint blockade. JAMA Oncol. 2019;5:471–8.
Nava Rodrigues D, Rescigno P, Liu D, Yuan W, Carreira S, Lambros MB, et al. Immunogenomic analyses associate immunological alterations with mismatch repair defects in prostate cancer. J Clin Invest. 2018;128:4441–53.
Huang JB, Wu YP, Lin YZ, Cai H, Chen SH, Sun XL, et al. Up-regulation of LIMK1 expression in prostate cancer is correlated with poor pathological features, lymph node metastases and biochemical recurrence. J Cell Mol Med. 2020;24:4698–706.
Gleicher S, Kauffman EC, Kotula L, Bratslavsky G, Vourganti S. Implications of high rates of metastatic prostate cancer in BRCA2 mutation carriers. Prostate. 2016;76:1135–45.
Barbieri CE, Baca SC, Lawrence MS, Demichelis F, Blattner M, Theurillat JP, et al. Exome sequencing identifies recurrent SPOP, FOXA1 and MED12 mutations in prostate cancer. Nat Genet. 2012;44:685–9.
Taylor BS, Schultz N, Hieronymus H, Gopalan A, Xiao Y, Carver BS, et al. Integrative genomic profiling of human prostate cancer. Cancer Cell. 2010;18:11–22.
Kumar A, Coleman I, Morrissey C, Zhang X, True LD, Gulati R, et al. Substantial interindividual and limited intraindividual genomic diversity among tumors from men with metastatic prostate cancer. Nat Med. 2016;22:369–78.
Chakravarty D, Gao J, Phillips SM, Kundra R, Zhang H, Wang J, et al. OncoKB: a precision oncology knowledge base. JCO Precis Oncol. 2017;1:1–16.
Lalonde E, Ishkanian AS, Sykes J, Fraser M, Ross-Adams H, Erho N, et al. Tumour genomic and microenvironmental heterogeneity for integrated prediction of 5-year biochemical recurrence of prostate cancer: a retrospective cohort study. Lancet Oncol. 2014;15:1521–32.
Buyyounouski MK, Pickles T, Kestin LL, Allison R, Williams SG. Validating the interval to biochemical failure for the identification of potentially lethal prostate cancer. J Clin Oncol. 2012;30:1857–63.
Ecke TH, Schlechte HH, Schiemenz K, Sachs MD, Lenk SV, Rudolph BD, et al. TP53 gene mutations in prostate cancer progression. Anticancer Res. 2010;30:1579–86.
Tosoian JJ, Almutairi F, Morais CL, Glavaris S, Hicks J, Sundi D, et al. Prevalence and prognostic significance of PTEN loss in African-American and European-American men undergoing radical prostatectomy. Eur Urol. 2017;71:697–700.
Kim SH, Park WS, Yun SI, Joo J, Joung JY, Seo HK, et al. Overexpression of BRCA1 or BRCA2 in prostatectomy specimens is predictive of biochemical recurrence after radical prostatectomy. Histopathology. 2016;68:673–9.
Ateeq B, Bhatia V, Goel S. Molecular discriminators of racial disparities in prostate cancer. Trends Cancer. 2016;2:116–20.
Donehower LA, Soussi T, Korkut A, Liu Y, Schultz A, Cardenas M, et al. Integrated analysis of TP53 gene and pathway alterations in the cancer genome atlas. Cell Rep. 2019;28:1370–84.e5.
Kunimasa K, Hirotsu Y, Nakamura H, Tamiya M, Iijima Y, Ishida H, et al. Rapid progressive lung cancers harbouring multiple clonal driver mutations with big bang evolution model. Cancer Genet. 2020;241:51–6.
Yamamoto S, Iwakuma T. Regulators of oncogenic mutant TP53 gain of function. Cancers (Basel). 2018;11:4.
Li M, Brooks CL, Wu-Baer F, Chen D, Baer R, Gu W. Mono- versus polyubiquitination: differential control of p53 fate by Mdm2. Science. 2003;302:1972–5.
Stopsack KH, Nandakumar S, Wibmer AG, Haywood S, Weg ES, Barnett ES, et al. Oncogenic genomic alterations, clinical phenotypes, and outcomes in metastatic castration-sensitive prostate cancer. Clin Cancer Res. 2020;26:3230–8.
Rajan P, Sudbery IM, Villasevil ME, Mui E, Fleming J, Davis M, et al. Next-generation sequencing of advanced prostate cancer treated with androgen-deprivation therapy. Eur Urol. 2014;66:32–9.
Pinyol R, Sia D, Llovet JM. Immune exclusion-Wnt/CTNNB1 class predicts resistance to immunotherapies in HCC. Clin Cancer Res. 2019;25:2021–3.
Linch M, Goh G, Hiley C, Shanmugabavan Y, McGranahan N, Rowan A, et al. Intratumoural evolutionary landscape of high-risk prostate cancer: the PROGENY study of genomic and immune parameters. Ann Oncol. 2017;28:2472–80.
Ding L, Getz G, Wheeler DA, Mardis ER, McLellan MD, Cibulskis K, et al. Somatic mutations affect key pathways in lung adenocarcinoma. Nature. 2008;455:1069–75.
Cao H, Liu X, Chen Y, Yang P, Huang T, Song L, et al. Circulating tumor DNA is capable of monitoring the therapeutic response and resistance in advanced colorectal cancer patients undergoing combined target and chemotherapy. Front Oncol. 2020;10:466.
Chen H, Chong W, Wu Q, Yao Y, Mao M, Wang X. Association of LRP1B mutation with tumor mutation burden and outcomes in melanoma and non-small cell lung cancer patients treated with immune check-point blockades. Front Immunol. 2019;10:1113.
Tucker MD, Zhu J, Marin D, Gupta RT, Gupta S, Berry WR, et al. Pembrolizumab in men with heavily treated metastatic castrate-resistant prostate cancer. Cancer Med. 2019;8:4644–55.
Annala M, Vandekerkhove G, Khalaf D, Taavitsainen S, Beja K, Warner EW, et al. Circulating tumor DNA genomics correlate with resistance to abiraterone and enzalutamide in prostate cancer. Cancer Discov. 2018;8:444–57.
Lindberg J, Mills IG, Klevebring D, Liu W, Neiman M, Xu J, et al. The mitochondrial and autosomal mutation landscapes of prostate cancer. Eur Urol. 2013;63:702–8.
Rampias T, Karagiannis D, Avgeris M, Polyzos A, Kokkalis A, Kanaki Z, et al. The lysine-specific methyltransferase KMT2C/MLL3 regulates DNA repair components in cancer. EMBO Rep. 2019;20:e46821.
Rubin MA, Maher CA, Chinnaiyan AM. Common gene rearrangements in prostate cancer. J Clin Oncol. 2011;29:3659–68.
Acknowledgements
This study was supported by the National Natural Science Foundation of China (NO.81800672) and the Key Research and Development Program of Shandong Province (NO.2017GSF18105 and NO.2019GSF108123).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Liu, Z., Guo, H., Zhu, Y. et al. TP53 alterations of hormone-naïve prostate cancer in the Chinese population. Prostate Cancer Prostatic Dis 24, 482–491 (2021). https://doi.org/10.1038/s41391-020-00302-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41391-020-00302-3
This article is cited by
-
Integrated analysis of single-cell and bulk RNA sequencing identifies a signature based on macrophage marker genes involved in prostate cancer prognosis and treatment responsiveness
Functional & Integrative Genomics (2023)
-
Cuproptosis-related lncRNA predict prognosis and immune response of lung adenocarcinoma
World Journal of Surgical Oncology (2022)
-
MAPK8IP2 is a potential prognostic biomarker and promote tumor progression in prostate cancer
BMC Cancer (2022)
-
Impact of TP53 mutations in Triple Negative Breast Cancer
npj Precision Oncology (2022)