Abstract
Fecal microbiota transfer (FMT) is a very efficient approach for the treatment of severe and recurring C. difficile infections. However, the beneficial effect of FMT in other disorders such as ulcerative colitis (UC) or Crohn’s disease remains unclear. Furthermore, it is currently unknown how disease-associated genetic variants in donors or recipients influence the effect of FMT. We found that bacteria-transfer from wild-type (WT) donors via cohousing was efficient in inducing recovery from colitis in WT mice, but not in mice deficient in protein-tyrosine phosphatase non-receptor type 22 (PTPN22), a known risk gene for several chronic inflammatory diseases. Also cohousing of PTPN22-deficient mice with diseased WT mice failed to induce faster recovery. Our data indicate that the genetic background of the donor and the recipient influences the outcome of microbiota transfer, and offers a potential explanation why transfer of fecal microbes from some, but not all donors is efficient in UC patients.
Similar content being viewed by others
Introduction
The human intestine is populated with up to 1014 bacteria that form a complex ecosystem and have a tremendous impact on our health.1,2 For several diseases, including obesity, diabetes, rheumatoid arthritis, liver diseases, and inflammatory bowel disease (IBD), shifts in the intestinal microbiota composition and a reduction in overall bacterial diversity as compared with healthy individuals have been described.3,4 However, it is still unclear whether those changes are the result of the underlying condition or if they functionally contribute to disease development/progression.5
Therapeutic transfer of fecal microbiota from healthy subjects to patients suffering from mainly intestinal diseases is increasingly attempted. In severe cases of recurring Clostridium difficile infections, fecal microbiota transplantation (FMT) has been shown to be very efficient,6,7,8 but the therapeutic value of FMT in other intestinal and non-intestinal disorders is still unresolved,9 and conflicting data from clinical trials using FMT in IBD have been reported.10,11,12,13,14 There is evidence that in the IBD subform ulcerative colitis (UC), the successful outcome of FMT might depend on the microbial composition of the donor,15 leading to the phenomenon of so-called “super-donors” that induce remission in more than 90% of transplanted cases, while most donors have success rates below 50%.10,11,12,13,14,16 Thus, it is important to define what constitutes a “successful” donor for the treatment of UC. Given the impact of genetic variants on IBD pathogenesis, the genetic background of the host as well as of the donor might exert a prominent role in therapeutic success.
Host genetic factors are known to influence the susceptibility to develop disease, and to date, 240 genetic loci have been identified that contribute to the relative risk to develop IBD.17,18,19,20 There is clear evidence that some of these genetic variants affect microbiota composition,21 but it is not known how genetic variants affect the outcome of FMT. A variant in the gene locus encoding PTPN22 results in an altered-function protein product and has been shown to protect from CD,22 but at the same time enhances the risk of developing auto-immune disorders such as rheumatoid arthritis or type-I-diabetes.23,24,25,26 PTPN22 mRNA and protein expression is reduced in IBD patients,27 and PTPN22-deficient mice are more susceptible to dextran sodium sulfate (DSS)-induced colitis,28 while mice carrying the autoimmunity-associated variant are protected from colitis onset,29 effects dependent on changes in the intestinal microbiota.30 Further, we recently found that presence of the autoimmunity-associated variant in IBD patients is associated with altered intestinal microbiota composition.31
Assessing the role of genetic factors for the successful outcome of FMT in UC in humans is difficult: FMT is still an experimental treatment for colitis, hence patient cohorts in FMT trials are typically not large enough to stratify according to genotypes in risk genes, and the genetic makeup of a given patient is often complex including several risk genes. Furthermore, donors are usually not genotyped for IBD risk genes. To overcome these drawbacks, we performed a study using cohoused mice to systematically address the role of the risk-gene PTPN22 on success of microbiota transfer. Specifically, we used the DSS-induced and naïve T-cell transfer models of colitis in mice carrying the autoimmunity-associated variant in PTPN22 as well as PTPN22-deficient mice.
Results
Cohousing of DSS-treated mice with healthy littermates promotes recovery
First, we addressed whether transfer of microbiota from healthy mice can promote recovery from acute colitis. For this aim, we treated C57BL/6 mice with 2% DSS in the drinking water for 7 days. At day 8, DSS was removed and one group of mice was cohoused with healthy littermates. Due to coprophagy, cohousing results in fast transfer of microbiota between mice in the same cage.32 The cohoused group recovered earlier (Fig. 1a) and 11 days after DSS removal, cohoused animals showed reduced intestinal inflammation (less pronounced shortening of the colon, reduced levels of myeloperoxidase, and reduced histological colitis scores in the terminal colon when compared with non-cohoused DSS-treated mice; Fig. 1b–d).
Cohousing promotes recovery in acute DSS colitis. Acute colitis was induced in WT mice via administration of 2.5% DSS in the drinking water for 7 days. At day 8, half of the DSS-treated mice were cohoused with healthy WT littermates. a Weight development (mean ± SEM, n = 8 per group), b colon length at the end of the experiment (day 17), c myeloperoxidase (MPO) levels in colon tissue (day 17), and d representative pictures and histological scoring from H&E-stained sections from the terminal colon (day 17). Indents in the pictures show infiltration (I) and epithelial damage (E) scores of the image. Each dot represents one individual mouse; asterisks denote significant differences (*p < 0.05, **p < 0.01, ***p < 0.001, Mann–Whitney). Representative results from one out of four independent experiments
Cohousing results in faster normalization of the intestinal microbiota during recovery from DSS
To analyze alterations in the intestinal microbiota upon DSS induction and during recovery, we collected stool before start of the experiment (day 0), just before cohousing/DSS removal (day 8), 3 days after DSS removal (day 11) and at the end of the experiment (day 17). As expected, DSS treatment resulted in reduced bacterial diversity (Fig. 2a) and a clear shift in bacterial composition (Fig. 2b, c; MANOVA of the first two PC on day 8, DSS vs. H2O: F(2,14) = 13.8, p = 2.4 × 10−6). In particular, we observed an increase in Akkermansiaceae (Akkermansia muciniphila) in DSS-treated mice (ANOVA of log relative abundances: F(1,14) = 143, p = 9.6 × 10−9). A more detailed investigation on the individual amplicon sequence variants (ASV) level revealed additional ASVs that were differentially abundant in DSS-treated mice relative to control mice (Supplementary Fig. 1a, b). These changes were transient, and bacterial composition normalized upon recovery from colitis. Of note, however, normalization of the microbiota was faster in cohoused DSS-treated mice (Fig. 2; MANOVA of the first two PC on day 11, cohoused vs. alone: F(2,6) = 3.71, p = 0.0346). In particular, five ASVs that significantly changed as a result of DSS treatment reverted already on day 11 in cohoused mice (Supplementary Fig. 1c). Interestingly, we also observed changes in the microbiota of the healthy mice that were cohoused with DSS-treated animals: on day 11 the cohoused healthy animals converged with their DSS cohoused counterparts (Fig. 2b, c), in particular through a decrease in the abundance of Akkermansiaceae in the cohoused DSS mice and an increase in the cohoused healthy mice. Taken together, this suggests that cohousing results in transfer of microbiota, which might crucially contribute to the observed faster recovery.
Cohousing induces faster normalization of microbiota composition. WT mice were treated as in Fig. 1 and fecal samples analyzed using 16S sequencing at the beginning of the experiment (day 0), directly before start of cohousing (day 8), 3 days after start of cohousing (day 11) and at the end of the experiment (day 17). a Bacterial diversity as indicated by Shannon index, b relative abundance of ASVs from indicated bacterial families (each column represents one mouse), and c principal component analysis. Color of the symbol indicates treatment (black = H2O, red = DSS); circles around the symbol indicate cohousing. Asterisks denote significant differences (*p < 0.05, Mann–Whitney)
Though segmental filamentous bacteria (SFB) might influence immune responses in the intestine, in our mouse colony abundance of SFB was rather low and did not differ in cohoused vs. non-cohoused mice (Supplementary Fig. 2a–c), suggesting that SFB do not play a prominent role in the observed effects. In line with changes in microbiota composition, we also observed a shift in the fecal abundance of the short chain fatty acid butyrate, which was decreased in DSS-treated mice, an effect less pronounced in DSS-treated mice cohoused with healthy individuals (Supplementary Fig. 2d). Other prominent short chain fatty acids were not affected by cohousing (Supplementary Fig. 2e, f).
To assess whether the beneficial effect of cohousing was due to either microbiota transfer or other factors, we next gavaged DSS-treated mice with the cecum content from either healthy control mice, or with cecum content from DSS-treated mice. Cecum content was used for gavages since the cecum is the main fermentation site in the mouse digestive system, and DSS-induced changes were similar in cecum content and feces (own unpublished observation). DSS-treated mice receiving cecum content from healthy mice gained weight faster than mice that did not receive cecum content or cecum content from DSS-treated mice, and showed reduced signs of intestinal inflammation at the end of the experiment (Supplementary Fig. 3). This indicates that the beneficial effect of cohousing is indeed mediated via transfer of microbiota.
Cohousing ameliorates T-cell-mediated colitis
Having shown that cohousing promotes recovery in chemically induced colitis, we next investigated whether cohousing also promotes health in an immune-cell-mediated colitis model. For this aim, we transferred naïve T cells into RAG1−/− hosts, which resulted in first symptoms of colitis 20 days later. One group of RAG1−/− was then cohoused with healthy wild-type (WT) mice. Of note, in cohoused RAG1−/− mice, disease severity stabilized (Fig. 3a), and 2 weeks after start of the cohousing, determination of spleen weight, colon length, and histological assessment of the colon revealed less severe colitis in cohoused RAG1−/− mice when compared with non-cohoused RAG1−/− mice (Fig. 3b–e). Again, 16S sequencing revealed that cohousing induced a pronounced microbiota shift (Fig. 3f; ANOVA of PC1: F(3,15) = 363, p = 3.18 × 10−14, Tukey’s HSD: RAG−/− T-cells non-cohoused/no T cells > RAG−/− T-cells cohoused, p = 10−11, RAG−/− T-cells cohoused > healthy WT donors, p = 0.00131) toward the microbiota of healthy donors. This indicates that cohousing-induced microbiota transfer not only reverts DSS-induced microbiota alterations, but might also modulate host defense and/or immune mechanisms.
Cohousing induces faster recovery in T-cell transfer colitis. 0.25 × 106 naïve T cells were transferred into RAG2−/− mice. After 18 days, half of the mice were cohoused with healthy C57BL/6 mice. a Weight development (mean ± SEM, n = 6–7 per group), b spleen weight, c colon length, d representative pictures, and e histological scoring of H&E-stained sections from the terminal colon, and f principal component (PCo) plots of 16S sequencing. Each dot represents one individual mouse; asterisks denote significant differences (*p < 0.05, Mann–Whitney). Depicted are results from one out of two independent experiments
Cohousing promotes recovery of the epithelial barrier and suppresses induction of Th1 cells in the lamina propria
Next, we investigated the host mechanisms underlying the enhanced recovery from colitis in cohoused mice. As expected, DSS treatment resulted in pronounced infiltration of macrophages and granulocytes into the lamina propria (Supplementary Fig. 4a, b) and there was accumulation of T cells, including CD4 + T helper (Th) cells, CD8+ cytotoxic T cells as well as IFN-γ+ Th1 and FoxP3+ regulatory T cells in the lamina propria from day 13 onward. In contrast, a significant accumulation of Th17 cells was only observed on day 15, a time point when CD8+ cells decreased to normal levels (Fig. 4a, b, Supplementary Fig. 4c–f). Cohousing did not affect numbers of infiltrating granulocytes and macrophages (Supplementary Fig. 4a, b). However, the accumulation of CD3+ cells, particularly of CD4+ Th1 cells at day 13 and 15 was almost completely prevented (Fig. 4a, b). Since we observed a reduction of myeloperoxidase (MPO) levels in cohoused mice, we further investigated whether cohousing affects granulocyte function. There was a slightly enhanced level of Mpo, Lyz2, and Nos2 mRNA in granulocytes from cohoused vs. non-cohoused mice (Fig. 4c), while degranulation capacity was decreased (Fig. 4d), indicating that cohousing modulates granulocyte function.
Reduced induction of Th1 cells, altered granulocyte function, and more proliferating epithelial cells in cohoused mice. Acute DSS colitis was induced in WT mice via administration of 2.5% DSS in the drinking water for 7 days. At day 8, half of the DSS-treated mice were cohoused with healthy littermates and mice sacrificed just before cohousing (day 8), 3 days after start of cohousing (day 11), 5 days after start of cohousing (day 13), or 7 days after start of cohousing (day 15). a IHC staining for CD3 of terminal colon sections, b flow cytometric quantification of IFN-γ-producing CD3+CD4+ cells; c Mpo, Lyz1, and Nos2 mRNA expression levels normalized to Actb and d degranulation capacity in colonic lamina propria granulocytes at day 15; e IHC staining for the proliferation marker Ki67 on terminal colon sections. f Mice were starved for 6 h prior to gavage with FITC-Dextran (4 kDa). 1 h later, level of FITC-Dextran was quantified in the serum. Each dot represents one individual mouse; asterisks denote significant differences (*p < 0.05, **p < 0.01, Mann–Whitney). Depicted are results from one out of two independent experiments
While in non-cohoused DSS-treated mice, proliferation of intestinal epithelial cells (IEC) (indicated by Ki67 IHC staining) was maximal at day 13 and 15, IEC proliferation was already very high in cohoused mice at day 11 (Fig. 4e), and was not restricted to base of the crypts, suggesting that cohousing promotes IEC proliferation and mucosal regeneration.
As a marker for intestinal barrier disruption, after gavage with fluorescein isothiocyanate (FITC)-dextran, FITC-dextran levels in the serum were high at day 8 in all DSS-treated mice, but further increased in non-cohoused mice at day 11 and stayed at that level until day 13 (Fig. 4f). In cohoused mice, in contrast, FITC-dextran levels did not further increase at day 11 and normalized much faster (Fig. 4f). Taken together, this indicates that cohousing promotes IEC proliferation, epithelial barrier reconstitution, suppresses granulocyte activation, and prevents the accumulation of potentially pathogenic Th1 cells. To assess whether Th1 cells indeed play a role in delaying recovery from acute DSS-induced colitis, we treated mice with an inhibitory anti-IFN-γ antibody. Of note, this treatment resulted in faster recovery (Supplementary Fig. 5), indicating that Th1 cells indeed play a role in maintaining colitis after acute DSS treatment. Nevertheless, cohoused mice recovered even faster than mice undergoing anti-IFN-γ treatment, indicating that this is not the only factor contributing to faster recovery in cohoused mice.
The efficacy of cohousing depends on the genetic makeup of the healthy “donor” mice
When addressing the expression of a series of IBD-associated genes (not shown) in cohoused vs. non-cohoused DSS-treated mice, we observed profound changes in Ptpn22 mRNA expression. Ptpn22 mRNA levels were only moderately enhanced in non-cohoused DSS-treated animals, but highly increased in cohoused counterparts (Fig. 5a), which prompted us to further investigate the role of PTPN22 in mediating the beneficial effect of cohousing. Further, we have previously shown that loss of PTPN2 or presence of a disease-associated variant in the gene encoding PTPN22 results in changes in microbiota composition.30 For this aim, we cohoused WT DSS-treated mice with healthy PTPN22 knockout (PTPN22−/−) mice or mice that express an altered-function PTPN22 variant that protects from CD onset (619W mice). PTPN22−/− mice have previously been reported to be more susceptible to acute DSS-induced colitis, while 619W mice are protected.28,29 Interestingly, when WT mice were cohoused with PTPN22−/− mice they did not recover faster than non-cohoused mice, while presence of the 619W variant did not alter the beneficial effect of cohousing (Fig. 5b, c). Also in the naïve T-cell transfer model of colitis, cohousing with PTPN22−/− mice did not promote recovery nor inhibited disease progression (Fig. 5d, e). This indicates that the genetic background of the healthy donor affects the beneficial effect of cohousing.
Cohousing of WT mice with PTPN22−/− mice does not promote recovery. a Acute DSS colitis was induced in WT mice via administration of 2.5% DSS in the drinking water for 7 days. At day 8, half of the DSS-treated mice were cohoused with healthy littermates and mice killed just before cohousing (day 8), 3 days after start of cohousing (day 11), 5 days after start of cohousing (day 13), or 7 days after start of cohousing (day 15) and colon tissue analyzed for Ptpn22 mRNA expression normalized to Atcb and H2O mice from day 8. b, c Acute colitis was induced in WT mice by administration of 2.5% DSS in the drinking water for 7 days. At day 8, mice were cohoused either with WT, PTPN22-619W (TG), or PTPN22−/− (KO) mice. Depicted are (b) weight development (mean ± SEM, n = 4 per group) and c representative pictures and histological scoring of H&E-stained sections from the terminal colon. d, e Transfer colitis was induced via transfer of 0.25 × 106 naïve T cells into RAG2−/− mice. After 18 days, mice were cohoused with WT or PTPN22−/− mice. d Weight development (mean ± SEM, n = 6 per group), and e representative pictures and histological scoring of H&E-stained sections from the terminal colon. (f) Principal component analysis of 16S sequencing from the stool from the mice in a and b. Each dot represents one individual mouse; asterisks denote significant differences (*p < 0.05, *p < 0.01, Mann–Whitney). Representative results from one out of two independent experiments
Changes in microbiota composition of PTPN22−/− mice result in failure to promote microbiota normalization
To address why cohousing with PTPN22−/− mice was unable to promote recovery, we analyzed the composition of the intestinal microbiota in WT, PTPN22−/− and 619W mice. In agreement with the previous experiment, DSS-treated mice showed an increase in Akkermansiaceae, but additionally also increases in the relative abundance of Prevotellaceae, Muribaculaceae, Erysipelotrichaceae on day 8 (Supplementary Fig. 6). Consistent with previous studies,30 PTPN22−/− mice harbored a different microbiota with decreased abundances of Helicobacteraceae, Deferribacteraceae, and trends for increased Akkermansiaceae but decreased Desulfovibrionaceae (Supplementary Fig. 6). Many individual ASVs were differentially abundant in PTPN22−/− mice compared with WT mice, but not in 619W mice (Supplementary Fig. 7). Both cohoused WT and 619W mice showed a slight decrease in Bacteroidaceae on day 12, while non-cohoused WT DSS mice and the cohoused PTPN22−/− mice showed an increase in Bacteroidaceae (Supplementary Figs. 6 and 7). While these changes in cohoused DSS mice were subtle, all cohoused untreated mice transiently converged toward a microbial composition that was more similar to their cohoused DSS counterparts (Fig. 5f and Supplementary Fig. 6). Similar effects were observed in the transfer colitis model (Supplementary Fig. 8). The effect of PTPN22-depletion on the intestinal microbiota seems to be dominant and gene-dose dependent, since PTPN22−/− mice showed a distinct microbiota composition from their WT littermates and heterozygous siblings showed an intermediate microbiota (Supplementary Fig. 9).
Genetic background of the diseased “recipient” determines success of cohousing
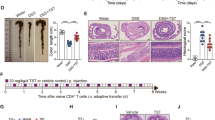
When PTPN22−/− mice were subjected to DSS treatment, neither cohousing with WT nor with PTPN22−/− mice promoted recovery (Fig. 6a, b). In contrast, DSS-treated 619W mice recovered faster when cohoused either with WT or 619W donors, and there was no difference between 619W and WT donors, indicating that presence of the 619W variant does not further improve the beneficial effect of cohousing (Fig. 6c, d). These results were fully confirmed in histological assessment of colitis severity (Fig. 6e, f). Of note, the failure to respond to the beneficial effect of cohousing seemed not to be due to more severe colitis in PTPN22−/− mice, since cohousing also failed to promote recovery in PTPN22−/− mice subjected to a lower dose of DSS, which showed mild colitis (Supplementary Fig. 10).
Cohousing fails to induce recovery in PTPN22−/− mice. Acute colitis was induced in PTPN22−/− (a, b) or 619W (c, d) mice by administration of 2.5% DSS in the drinking water for 7 days. At day 8, mice were cohoused either with WT or PTPN22−/− (a, b) or WT or 619W (c, d) mice. Depicted are (a, c) weight development (mean ± SEM, n = 4 per group) and (b, d) colon length. Show representative pictures (e) and histological scoring (f) of H&E-stained sections from the terminal colon from the same mice as in a–d. g Principal component analysis of 16S Sequencing from the stool from the mice in a. Each dot represents one individual mouse; asterisks denote significant differences (*p < 0.05, Mann–Whitney). Depicted are results from one out of two independent experiments
As reported previously,30 DSS-treated PTPN22−/− mice presented a strong shift in the intestinal microbiota composition on day 8 (Fig. 6g, Supplementary Fig. 11), primarily characterized by increased relative abundances of Erysipelotrichaceae (Supplementary Fig. 12a). The microbiota further changed from day 8 to day 12 and to day 16, though without any indication of convergence towards the healthy mice (Fig. 6e and Supplementary Fig. 12a).
In line with our previous findings,30 DSS-treated 619W mice also presented a strongly shifted microbial composition on day 8, equivalently characterized by an increase in Erysipelotrichaceae (Supplementary Figs. 11 and 12b, c). Upon cohousing, the microbiota converged more closely toward their cohoused counterparts. Importantly, 619W displayed a baseline microbial community that differed from WT mice (increased Erysipelotrichaceae and reduced Akkermansiaceae), and those DSS mice that were cohoused with WT mice converged toward the WT mice, despite their own 619W genotype (Fig. 6g and Supplementary Fig. 12b, c). This shift was partly reduced on day 16, with the microbiota returning to a composition that resembled that of healthy 619W mice.
Th1 responses are not suppressed in WT mice cohoused with PTPN22−/− mice
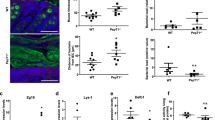
Having observed that cohousing promotes barrier recovery and suppresses Th1 cell induction in the inflamed intestine in WT mice, we next analyzed whether these effects are also present in WT mice upon cohousing with PTPN22−/− or 619W mice. In line with a failure to induce recovery, cohousing of DSS-treated WT mice with healthy PTPN22−/− mice was not able to suppress the accumulation of Th1 cells at day 15, while cohousing with WT or 619W mice clearly prevented Th1 accumulation (Fig. 7a). Further, in WT mice cohoused with PTPN22−/− mice, proliferation of IEC was reduced at day 11 when compared with DSS-treated WT mice that were cohoused with either healthy WT or 619W mice (Fig. 7b). In line with a role for granulocytes in mediating the observed effects of cohousing, granulocytes from mice cohoused with PTPN22−/− donors did not show differences in Mpo mRNA expression or decreased degranulation capacity (Fig. 7d, e). In addition, PTPN22−/− granulocytes showed reduced MPO expression (Fig. 7f). This clearly indicates that cohousing with PTPN22−/− mice fails to induce recovery via failure to suppress granulocyte activation, pro-inflammatory Th cell accumulation, and inability to promote barrier reconstitution.
a Acute colitis was induced in WT mice by administration of 2.5% DSS in the drinking water for 7 days. At day 8, mice were cohoused with WT or PTPN22−/− (KO) or 619W (TG) mice and 7 days later (day 15) colon tissue was analyzed by flow cytometry for the abundance of IFN-γ+ cells among CD4 + T cells. b, c Mice were treated as in a and killed 3 days after start of cohousing (day 11). Depicted are (b) representative pictures from IHC of the proliferation marker Ki67. c Mice were starved for 6 h prior to gavage with FITC-dextran (4 kDa). 1 h after gavages, level of FITC-dextran was quantified in the serum. Mice were treated as in (a) and colonic lamina propria granulocytes analyzed for (d) mRNA expression of Mpo normalized to Actb, and (e) degranulation capacity. f Coloic lamina propria granulocytes were isolated from WT, KO, and TG mice and analyzed for mRNA expression of Mpo normalized to Actb. Each dot represents one individual mouse; asterisks denote significant differences (*p < 0.05, **p < 0.01, Mann–Whitney). Depicted are results from one out of two independent studies
Discussion
In this study, we investigated the therapeutic concept of microbiota transfer via cohousing using mouse colitis models and assessed how transfer of healthy microbiota affects recovery from intestinal inflammation. We found that the beneficial effect of cohousing critically depends on the genetic background of the donor and the recipient, an observation that might be of great relevance in the clinical setting.
As a mode of action, we demonstrated that cohousing promotes recovery from colitis via induction of earlier epithelial cell proliferation and restoration of a functional epithelial barrier, while suppressing (excessive) granulocyte activation and induction of Th1 cells. Deletion of the IBD-associated gene PTPN22 in donor or recipient mice abrogated these beneficial effects. This clearly indicates that the genetic makeup of the donor, as well as of the recipient crucially determines the beneficial effect of microbiota transfer, which might be of high clinical relevance for the selection of suitable donors and for prediction of treatment success.
Most experimental studies addressing the effect of certain microbes on intestinal health used either germ-free or antibiotic-treated mice. However, these models poorly reflect the real-life situation in IBD patients receiving FMT, since those patients are usually not premedicated with antibiotics. Furthermore, germ-free mice have severe defects/alterations in the architecture of the intestinal immune system (reviewed in ref. 33), a factor that obviously plays a central role in the development of intestinal disorders in general, and in IBD in particular. In our model, microbiota is transferred via cohousing, which—due to coprophagy in mice—results in efficient transfer of the intestinal microbiota between mice in the same cage. From a microbe’s point of view, this closely reflects the situation in IBD patients, since in both models and in IBD patients the transferred microbiota encounters an already colonized host. Nevertheless, cohousing has the disadvantage that the amount of transferred microbiota and the direction of transfer cannot be controlled. However, since transfer of microbiota via oral gavages with a defined amount of donor material was equally efficient in terms of disease outcome, we assume that cohousing is an adequate way to study (fecal) microbiota transfer in already colonized hosts.
Previous studies have assessed the effect of (genotype-mediated) changes in the micobiota composition on colitis severity/susceptibility.32,34,35 However, these studies assessed how changes in the microbiota before colitis induction affect disease course, while our study assessed how the genetic makeup of microbiota donors affects the beneficial effects of microbiota transfer/cohousing after disease induction when colitis is already present.
In cases of recurrent C. difficile infections it is suggested that FMT alleviates dysbiosis and restores a normal, healthy microbiota in the gut that is resilient to C. difficile overgrowth.7,36 In contrast, the mode of action of how FMT promotes induction/maintenance of remission in colitis is less clear. In our model, cohousing promoted normalization of the microbiota composition, and in those mice where normalization of the microbiota was not observed (i.e., in recipients that were cohoused with PTPN22−/− donors), no induction of faster recovery was observed. This indicates that the beneficial effect of cohousing might be caused via alleviation of dysbiosis. Of note, cohousing was efficient in both a model of epithelial injury and in immune-cell-mediated colitis. In line with this, we observed that cohousing promoted growth of IEC and barrier reconstitution, but also affected granulocyte function and Th cell subsets. Recent publications showed that the composition of the microbiota might crucially affect the outcome of FMT, and in general, higher bacteria richness positively influenced the effect of FMT.13,14,15 Likewise, increased levels of Ruminococcaceae and Lachnospiraceae were associated with better response to FMT.13,15,37 However, neither this present study, nor our recently published report on microbial changes in PTPN22−/− mice30 indicated a specific bacterial strain or group that was clearly associated with recovery from colitis upon cohousing, or specific strains that were missing in PTPN22−/− mice that would explain why cohousing with PTPN22−/− mice did not promote recovery. Another explanation for the failure of cohousing to induce recovery in PTPN22−/− mice might be the fact that those mice suffer from a more severe disease than their WT littermates. Nevertheless, when PTPN22−/− mice were subjected to mild DSS colitis, cohousing still failed to promote recovery, indicating that disease severity is not the only factor preventing response to cohousing in those mice.
Regarding immune cells, Th cells showed interesting patterns in response to cohousing. While cohousing suppressed DSS-induced Th1 accumulation, frequencies of Th17 cells were not affected. Although connoted with inflammation,38,39 Th17 are abundant in healthy colons40 and exert important functions for immunity against extracellular bacteria and fungi,41 and normal production of IL17 seems to be important to prevent systemic dissemination of pathogens.42 Th1 and their hallmark cytokine IFN-γ, in contrast, have been implicated in driving intestinal inflammation and tissue damage in murine colitis models.43,44 Furthermore, genome-wide association studies linked SNPs around the IFNG locus and enhanced IFNG expression with IBD.45,46 Certain bacteria promote specific T-cell responses, e.g., SFB drive the development of Th17 cells.47 Likewise, specific bacteria in healthy mice might suppress the induction of Th1 cells. This implicates that in colitis microbiota transfer via cohousing might act in a dual mode, on one hand via alleviating dysbiosis, and on the other hand via promotion of an intact epithelial barrier and suppression of exacerbated immune responses.
Aside from T cells, we also found differences in granulocyte function upon cohousing, an effect not observed upon cohousing with PTPN22−/− mice. These findings are partially in line with reports that show regulation of neutrophil migration and activation by PTPN22.48,49 This indicates that granulocytes might be importantly involved in determining the beneficial effects of cohousing. However, additional functional studies would be necessary to clarify the role of granulocytes in FMT, and especially studies in humans receiving FMT are needed to understand whether granulocytes are important in the human setting.
Of high interest for future studies on possible therapeutic approaches using FMT are our findings that the genetic background of donor and recipient affected the outcome of microbiota transfer upon colitis induction. In particular, loss of PTPN22 abrogated the beneficial effect of cohousing. PTPN22 is an important regulatory molecule involved in T- and B-cell biology50,51 as well as in controlling monocyte/macrophage responses to interferons,52 autophagy,53 and inflammasome activation.29 Alterations in all of these factors can influence the composition of the intestinal microbiota,54,55,56,57 and indeed PTPN22−/− donor mice as well as PTPN22−/− recipients showed changes in the composition of the intestinal microbiota. Consistent with reluctance to faster recovery upon cohousing, Helicobacteraceae, which contain potential pathobionts,58 were elevated in PTPN22−/− mice. However, Deferribacteraceae and Desulfovibronacea (both reduced in PTPN22−/− mice) are described to increase in murine colitis models, and it has been suggested that they might be an indicator for disease activity.59,60 Hence, it appears counterintuitive that these bacterial families are decreased in PTPN22−/− mice. However, increased levels of Akkermansiaceae at baseline might contribute to enhanced susceptibility for colitis, non-response to cohousing, and the lack of ability to induce recovery via cohousing. Akkermansiaceae are mucus-degrading bacteria that increase during colitis induction in animal models.59 Nevertheless, there is a study showing that in human IBD Akkermanciaceae levels are clearly decreased, while other mucin-degrading bacteria are increased.61 Although these changes are interesting, we did not find specific bacteria that predicted the success of cohousing, and additional studies are required to address which bacteria are causatively involved in promoting recovery in mice cohoused with WT donors.
Taken together, we have demonstrated that microbiota transfer via cohousing is an efficient method to induce recovery from colitis in mouse models of intestinal inflammation. This effect is mediated via promotion of a faster normalization of the microbiota in diseased mice, induction of earlier barrier reconstitution, and suppression of potentially pathogenic T-cell populations. In addition, our data demonstrate that genetic factors in donors and hosts crucially affect the outcome of microbiota transfer. This is of great importance for clinical practice, since it lays the groundwork for future FMT studies with the aim of defining and selecting more efficient donors. Nevertheless, it will be of importance to validate the findings of our study in the human setting.
Methods
Mice, colitis models, and cohousing
PTPN22-deficient (PTPN22−/−) mice in a C57B/6 background were obtained from Andrew C. Chan from Genentech (San Francisco, California)62 and crossed with WT mice. A local colony of heterozygous mice was maintained for breeding of PTPN22−/− and WT littermates. No heterozygous mice were used for the studies except for Supplementary Fig. S9, where the difference in microbiota composition between WT, PTPN22+/−, and PTPN22−/− littermates was assessed. Mice carrying the 619W variant in the PTPN22 gene in a C57BL/6 background were obtained from D. Rawlings at the Children’s Hospital in Seattle (University of Washington School of Medicine, Seattle, Washington),63 bred with WT mice, and a local colony of heterozygous mice was maintained in our facility for breeding of 619W and WT littermates. In experiments involving PTPN22−/− and 619W mice, a mix of WT littermates from both breeding colonies were used. All animal experiments were conducted according to Swiss animal welfare legislation and were approved by the local animal welfare commission (Tierschutzkommission des Kantons Zürich). DSS colitis64 was induced by administration of 2.5% DSS in the drinking water for 7 days. At day 8, water was exchanged to normal drinking water to allow recovery from colitis. For transfer colitis,65 naïve T cells (CD4+, CD62Lhigh, CD44low single cells) were sorted from the spleen of donor mice and 5 × 105 cells injected intraperitoneally into RAG2 deficient hosts. At start of cohousing (day 8 of acute DSS-induced colitis, day 18 for transfer colitis), all mice were transferred into fresh cages. For IFN-γ inhibition, mice were injected at experimental days 8, 11, and 14 with 1 mg/kg anti-IFN-γ (clone XMG1.2; BioXCell West Lebanon, NH).
Analysis of colitis severity
Colitis severity was monitored daily by assessing body weight development, stool consistency, general appearance, and activity of each mouse according to the disease activity index previously described for colitis experiments.27 At the end of the experiment, animals were euthanized and colon length and spleen weight analyzed as indirect measures of colitis severity. MPO activity was measured in 0.5 cm long colon pieces as described,27 and sections from paraffin-embedded terminal colon pieces were hematoxylin/eosin stained using standard protocols27 to determine the extent of colitis by histology. A well-established and previously described scoring system was used to determine the severity of colitis in histological sections.27
16S sequencing and analysis of microbiota
16S sequencing and analysis was performed according to standard methods and described in detail in Supplementary methods. In brief, stool was collected at indicated time points and stored at −80 °C until isolation of genomic DNA using the MoBio PowerLyzer Soil Kit from Qiagen (Thermo Fisher Scientific). Samples were then sent to Microsynth (Balgach, Switzerland) for sequencing of the V4 hypervariable domain in the 16S ribosomal DNA using the primers pair 515F (GTGYCAGCMGCCGCGGTAA) and 806R (GGACTACNVGGGTWTCTAAT) prior to 350 × 350 paired end sequencing on the Illumina MiSeq platform. After trimming, merging of forward and reverse reads and removal of chimeric reads, taxonomic assignment of ASVs using SILVA 132, differential abundance was calculated using DESeq2 using local regression to estimate dispersion.
Flow cytometry and isolation of lamina propria granulocytes
Single cells suspensions from mesenteric lymph nodes and lamina propria were prepared as described previously,29 and cell analysis performed according to standard procedures as described in Supplementary methods. All analyses were performed on an LSR Fortessa analyzer (BD Pharmingen). For granulocyte isolation, lamina propria cells were isolated as described previously29 and live, CD45 + Ly6G + CD3-B220-NK1.1-cells sorted on a FACSAria (BD).
Degranulation assay
To assess degranulation, granulocytes were treated with formylated peptide (fMLP, 100 nM) for 10 min and lactoferrin levels in the cell culture supernatant, as well as within the cells, were measured using an ELISA kit (Hycult Biotech, Uden, Netherlands). Percent degranulation was calculated as proportion of lactoferrin in the supernatant divided by total lactoferrin levels.
Immunhistochemistry
Immunhistochemistry was performed according to standard procedures and as described previously.66 Details are given in Supplementary methods.
In vivo barrier permeability
Mice were starved for 6 h prior to gavage with FITC-dextran (4 kDa, 600 mg/kg body weight). After 1 h, 120 µL blood was collected from the sublingual vein into serum collection tubes. The blood was centrifuged at 4 °C, 8000 g, for 3 min, serum diluted in an equal volume of PBS (pH 7.4), and analyzed for FITC-dextran concentration with a fluorescence spectrophotometer (SynergyII plate-reader with Gen5 software; BioTek Instruments, Winooski, VT) using an excitation wavelength of 485 nm and an emission wavelength of 535 nm. Standard curves for calculating FITC-dextran concentration in the samples were obtained by diluting FITC-dextran in serum from non-gavaged mice.
Statistical analysis
Unless otherwise stated, data are representative for one of at least two independent experiments with n replicates each; unless otherwise stated, data are represented as average and standard error of the mean. Statistical significances were determined using ANOVA followed by Wilcoxon–Mann–Whitney test for significance. P values below 0.05 are considered significant. Microbiota read counts were first grouped and summed at a given taxonomic level. Principal component analysis was performed on centered log-ratio transformed data after adding a pseudo count equal to the mean number of read counts to each sample.
Study approval
All animal experiments have been conducted according to Swiss animal welfare legislation. The local animal welfare commission (Veterinary Office of the Canton of Zurich) has approved all procedures involving mice (approval numbers ZH2014/255 and ZH2017/20).
Data availability
All datasets generated and analyzed during this study are available from the corresponding author upon request.
References
Ley, R. E., Peterson, D. A. & Gordon, J. I. Ecological and evolutionary forces shaping microbial diversity in the human intestine. Cell 124, 837–848 (2006).
Sender, R., Fuchs, S. & Milo, R. Revised estimates for the number of human and bacteria cells in the body. PLoS Biol. 14, e1002533 (2016).
De Cruz, P. et al. Characterization of the gastrointestinal microbiota in health and inflammatory bowel disease. Inflamm. Bowel Dis. 18, 372–390 (2012).
Kostic, A. D., Xavier, R. J. & Gevers, D. The microbiome in inflammatory bowel disease: current status and the future ahead. Gastroenterology 146, 1489–1499 (2014).
Sartor, R. B. & Wu, G. D. Roles for intestinal bacteria, viruses, and fungi in pathogenesis of inflammatory bowel diseases and therapeutic approaches. Gastroenterology 152, 327–339 (2017).
Hamilton, M. J., Weingarden, A. R., Sadowsky, M. J. & Khoruts, A. Standardized frozen preparation for transplantation of fecal microbiota for recurrent Clostridium difficile infection. Am. J. Gastroenterol. 107, 761–767 (2012).
van Nood, E. et al. Duodenal infusion of donor feces for recurrent Clostridium difficile. N. Engl. J. Med. 368, 407–415 (2013).
Khoruts, A. & Weingarden, A. R. Emergence of fecal microbiota transplantation as an approach to repair disrupted microbial gut ecology. Immunol. Lett. 162(2 Pt A), 77–81 (2014).
Sha, S. et al. Systematic review: faecal microbiota transplantation therapy for digestive and nondigestive disorders in adults and children. Aliment Pharm. Ther. 39, 1003–1032 (2014).
Colman, R. J. & Rubin, D. T. Fecal microbiota transplantation as therapy for inflammatory bowel disease: a systematic review and meta-analysis. J. Crohns Colitis 8, 1569–1581 (2014).
Goyal, A. et al. Safety, clinical response, and microbiome findings following fecal microbiota transplant in children with inflammatory bowel disease. Inflamm. Bowel Dis. 24, 410–421 (2018).
Browne, A. S. & Kelly, C. R. Fecal transplant in inflammatory bowel disease. Gastroenterol. Clin. North Am. 46, 825–837 (2017).
Rossen, N. G. et al. Findings from a randomized controlled trial of fecal transplantation for patients with ulcerative colitis. Gastroenterology 149, 110–118 e114 (2015).
Paramsothy, S. et al. Multidonor intensive faecal microbiota transplantation for active ulcerative colitis: a randomised placebo-controlled trial. Lancet 389, 1218–1228 (2017).
Moayyedi, P. et al. Fecal microbiota transplantation induces remission in patients with active ulcerative colitis in a randomized controlled trial. Gastroenterology 149, 102–109 (2015).
Fuentes, S. et al. Microbial shifts and signatures of long-term remission in ulcerative colitis after faecal microbiota transplantation. ISME J. 11, 1877–1889 (2017).
Wellcome Trust Case Control C. Genome-wide association study of 14,000 cases of seven common diseases and 3,000 shared controls. Nature 447, 661–678 (2007).
Anderson, C. A. et al. Meta-analysis identifies 29 additional ulcerative colitis risk loci, increasing the number of confirmed associations to 47. Nat. Genet. 43, 246–252 (2011).
Liu, J. Z. et al. Association analyses identify 38 susceptibility loci for inflammatory bowel disease and highlight shared genetic risk across populations. Nat. Genet. 47, 979–986 (2015).
de Lange, K. M. et al. Genome-wide association study implicates immune activation of multiple integrin genes in inflammatory bowel disease. Nat. Genet. 49, 256–261 (2017).
Chu, H. et al. Gene-microbiota interactions contribute to the pathogenesis of inflammatory bowel disease. Science 352, 1116–1120 (2016).
Diaz-Gallo, L. M. et al. Differential association of two PTPN22 coding variants with Crohn’s disease and ulcerative colitis. Inflamm. Bowel Dis. 17, 2287–2294 (2011).
Begovich, A. B. et al. A missense single-nucleotide polymorphism in a gene encoding a protein tyrosine phosphatase (PTPN22) is associated with rheumatoid arthritis. Am. J. Hum. Genet. 75, 330–337 (2004).
Kyogoku, C. et al. Genetic association of the R620W polymorphism of protein tyrosine phosphatase PTPN22 with human SLE. Am. J. Hum. Genet. 75, 504–507 (2004).
Ramirez, M. et al. The PTPN22 C1858T variant as a risk factor for rheumatoid arthritis and systemic lupus erythematosus but not for systemic sclerosis in the Colombian population. Clin. Exp. Rheuma. 30, 520–524 (2012).
Stanford, S. M. & Bottini, N. PTPN22: the archetypal non-HLA autoimmunity gene. Nat. Rev. Rheuma. 10, 602–611 (2014).
Spalinger, M. R. et al. Loss of protein tyrosine phosphatase nonreceptor type 22 regulates interferon-gamma-induced signaling in human monocytes. Gastroenterology 144, 978–988 e910 (2013).
Wang, Y. et al. The autoimmunity-associated gene PTPN22 potentiates toll-like receptor-driven, Type 1 interferon-dependent immunity. Immunity 39, 111–122 (2013).
Spalinger, M. R. et al. NLRP3 tyrosine phosphorylation is controlled by protein tyrosine phosphatase PTPN22. J. Clin. Invest. 126, 1783–1800 (2016).
Spalinger, M. R. et al. Protein tyrosine phosphatase non-receptor type 22 modulates colitis in a microbiota-dependent manner. J. Clin. Invest. 130, 2527–2541 (2019).
Yilmaz, B. et al. The presence of genetic risk variants within PTPN2 and PTPN22 is associated with intestinal microbiota alterations in Swiss IBD cohort patients. PLoS ONE 13, e0199664 (2018).
Couturier-Maillard, A. et al. NOD2-mediated dysbiosis predisposes mice to transmissible colitis and colorectal cancer. J. Clin. Invest. 123, 700–711 (2013).
Round, J. L. & Mazmanian, S. K. The gut microbiome shapes intestinal immune responses during health and disease. Nat. Rev. Immunol. 9, 313–323 (2009).
Elinav, E. et al. NLRP6 inflammasome regulates colonic microbial ecology and risk for colitis. Cell 145, 745–757 (2011).
Britton, G. J. et al. Microbiotas from humans with inflammatory bowel disease alter the balance of gut Th17 and RORgammat(+) regulatory t cells and exacerbate colitis in mice. Immunity 50, 212–224 e214 (2019).
Kelly, C. R. et al. Effect of fecal microbiota transplantation on recurrence in multiply recurrent clostridium difficile infection: a randomized trial. Ann. Intern. Med. 165, 609–616 (2016).
Kump, P. et al. The taxonomic composition of the donor intestinal microbiota is a major factor influencing the efficacy of faecal microbiota transplantation in therapy refractory ulcerative colitis. Aliment Pharm. Ther. 47, 67–77 (2018).
Korn, T., Bettelli, E., Oukka, M. & Kuchroo, V. K. IL-17 and Th17 cells. Annu. Rev. Immunol. 27, 485–517 (2009).
Miossec, P. & Kolls, J. K. Targeting IL-17 and TH17 cells in chronic inflammation. Nat. Rev. Drug Discov. 11, 763–776 (2012).
Kempski, J., Brockmann, L., Gagliani, N. & Huber, S. T(H)17Cell and epithelial cell crosstalk during inflammatory bowel disease and carcinogenesis. Front Immunol. 8, 1373 (2017).
Huang, W., Na, L., Fidel, P. L. & Schwarzenberger, P. Requirement of interleukin-17A for systemic anti-Candida albicans host defense in mice. J. Infect. Dis. 190, 624–631 (2004).
Ouyang, W., Kolls, J. K. & Zheng, Y. The biological functions of T Helper 17 cell effector cytokines in inflammation. Immunity 28, 454–467 (2008).
Powrie, F. et al. Inhibition of Th1 responses prevents inflammatory bowel disease in scid mice reconstituted with CD45RBhi CD4+ T cells. Immunity 1, 553–562 (1994).
Ito, R. et al. Interferon-gamma is causatively involved in experimental inflammatory bowel disease in mice. Clin. Exp. Immunol. 146, 330–338 (2006).
Jostins, L. et al. Host-microbe interactions have shaped the genetic architecture of inflammatory bowel disease. Nature 491, 119–124 (2012).
Gonsky, R. et al. IFNG rs1861494 polymorphism is associated with IBD disease severity and functional changes in both IFNG methylation and protein secretion. Inflamm. Bowel Dis. 20, 1794–1801 (2014).
Ivanov, I. I. et al. Induction of intestinal Th17 cells by segmented filamentous bacteria. Cell 139, 485–498 (2009).
Bayley, R. et al. The autoimmune-associated genetic variant PTPN22 R620W enhances neutrophil activation and function in patients with rheumatoid arthritis and healthy individuals. Ann. Rheum. Dis. 74, 1588–1595 (2015).
Vermeren, S. et al. PTPN22 is a critical regulator of fcgamma receptor-mediated neutrophil activation. J. Immunol. 197, 4771–4779 (2016).
Wu, J. et al. Identification of substrates of human protein-tyrosine phosphatase PTPN22. J. Biol. Chem. 281, 11002–11010 (2006).
Dai, X. et al. A disease-associated PTPN22 variant promotes systemic autoimmunity in murine models. J. Clin. Invest. 123, 2024–2036 (2013).
Wang, Y. et al. PTPN22 variant R620W is associated with reduced toll-like receptor 7-induced type I interferon in systemic lupus erythematosus. Arthritis Rheuma. 67, 2403–2414 (2015).
Spalinger, M. R. et al. Protein tyrosine phosphatase non-receptor type 22 modulates NOD2-induced cytokine release and autophagy. PLoS ONE 8, e72384 (2013).
Levy, M. et al. Microbiota-modulated metabolites shape the intestinal microenvironment by regulating NLRP6 inflammasome signaling. Cell 163, 1428–1443 (2015).
Lee, N. & Kim, W.-U. Microbiota in T-cell homeostasis and inflammatory diseases. Exp. Mol. Med. 49, e340 (2017).
Rogler, G. The importance of gut microbiota in mediating the effect of NOD2 defects in inflammatory bowel disease. Gut 59, 153–154 (2010).
Hooper, L. V., Littman, D. R. & Macpherson, A. J. Interactions between the microbiota and the immune system. Science 336, 1268–1273 (2012).
Mazmanian, S. K., Round, J. L. & Kasper, D. L. A microbial symbiosis factor prevents intestinal inflammatory disease. Nature 453, 620–625 (2008).
Rooks, M. G. et al. Gut microbiome composition and function in experimental colitis during active disease and treatment-induced remission. ISME J. 8, 1403–1417 (2014).
Berry, D. et al. Intestinal microbiota signatures associated with inflammation history in mice experiencing recurring colitis. Front Microbiol. 6, 1408 (2015).
Png, C. W. et al. Mucolytic bacteria with increased prevalence in IBD mucosa augment in vitro utilization of mucin by other bacteria. Am. J. Gastroenterol. 105, 2420–2428 (2010).
Hasegawa, K. et al. PEST domain-enriched tyrosine phosphatase (PEP) regulation of effector/memory T cells. Science 303, 685–689 (2004).
Dai, Y. & Hu, S. Recent insights into the role of autophagy in the pathogenesis of rheumatoid arthritis. Rheumatol. (Oxf.) 55, 403–410 (2016).
Chassaing, B., Aitken, J. D., Malleshappa, M. & Vijay-Kumar, M. Dextran sulfate sodium (DSS)-induced colitis in mice. Curr. Protoc. Immunol. 2014; 104 : Unit 15.25.
Ostanin, D. V. et al. T cell transfer model of chronic colitis: concepts, considerations, and tricks of the trade. Am. J. Physiol. Gastrointest. Liver Physiol. 296, G135–G146 (2009).
Spalinger, M. R. et al. PTPN2 controls differentiation of CD4 T cells and limits intestinal inflammation and intestinal dysbiosis. Mucosal Immunol. 8, 918–929 2015.
Acknowledgements
This work was supported by research grants from the Swiss National Science Foundation to MiS (Grant No. 314730-146204, Grant No. 314730_166381 and Grant No. CRSII3_154488/1) and AG (Grant No. 35150) and the Stiftung Experimentelle Biomedizin to MiS. The funding institutions had no role in study design and data interpretation.
Author information
Authors and Affiliations
Contributions
M.R.S: study design, performing experiments, data analysis, and interpretation, writing the manuscript; Ma.S., L.H., C.G., S.L., K.B., An.S., Al.Sa. and Al.Sh.: performing experiment and data analysis; A.G., C.L., G.E.L.: data analysis and interpretation, critical input on microbiota data; D.J.R., X.D., A.A.C.: generation of PTPN22 KO and TG mice; G.R. and D.F.M.: critical intellectual input; Mi.S.: conceived, designed and supervised the study, funding. All authors wrote, corrected, and approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementaty Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Spalinger, M.R., Schwarzfischer, M., Hering, L. et al. Loss of PTPN22 abrogates the beneficial effect of cohousing-mediated fecal microbiota transfer in murine colitis. Mucosal Immunol 12, 1336–1347 (2019). https://doi.org/10.1038/s41385-019-0201-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41385-019-0201-1
This article is cited by
-
Temperature-triggered in situ forming lipid mesophase gel for local treatment of ulcerative colitis
Nature Communications (2023)
-
Gut microbiota involved in leptospiral infections
The ISME Journal (2022)