Abstract
Mantle cell lymphoma (MCL) is a lethal hematological malignancy with a median survival of 4 years. Its lethality is mainly attributed to a limited understanding of clinical tumor progression and resistance to current therapeutic regimes. Intrinsic, prolonged drug treatment and tumor-microenvironment (TME) facilitated factors impart pro-tumorigenic and drug-insensitivity properties to MCL cells. Hence, elucidating neoteric pharmacotherapeutic molecular targets involved in MCL progression utilizing a global “unified” analysis for improved disease prevention is an earnest need. Using integrated transcriptomic analyses in MCL patients, we identified a Fibroblast Growth Factor Receptor-1 (FGFR1), and analyses of MCL patient samples showed that high FGFR1 expression was associated with shorter overall survival in MCL patient cohorts. Functional studies using pharmacological intervention and loss of function identify a novel MYC-EZH2-CDKN1C axis-driven proliferation in MCL. Further, pharmacological targeting with erdafitinib, a selective small molecule targeting FGFRs, induced cell-cycle arrest and cell death in-vitro, inhibited tumor progression, and improved overall survival in-vivo. We performed extensive pre-clinical assessments in multiple in-vivo model systems to confirm the therapeutic potential of erdafitinib in MCL and demonstrated FGFR1 as a viable therapeutic target in MCL.

Similar content being viewed by others
Introduction
Mantle cell lymphoma (MCL) is an aggressive mature B-cell neoplasm [1, 2], with 60% of patients showing bone marrow involvement at diagnosis [3, 4]. Despite intensive therapeutic approaches, the median progression-free survival after first-line treatment is four years [5,6,7]. The emergence of chemoresistance is rapid, durable responses to second and third-line therapies are rare, and relapse is virtually universal in MCL [8,9,10]. However, for most patients who progress on targeted agents like ibrutinib, survival is only 6-10 months [8, 9, 11]. The interaction of MCL with the tumor microenvironment (TME) provides specific niches for lymphoma cells to communicate and promote growth and resistance to chemotherapeutic agents [12]. Therefore, the identification of clinically relevant targets and the development of new therapies are urgently needed for MCL patients.
Cell cycle dysregulation is a hallmark of MCL, resulting from a t(11;14) chromosomal translocation that drives unrestrained proliferation of cancer cells [13, 14]. While the Eμ-Cyclin D1 mouse model does not develop MCL [15], Sox11 and CyclinD1 are essential MCL biomarkers and upregulated in 80% of MCL cases. Eµ-SOX11:CCND1 double transgenic (DT) mice exhibit cell cycle programming and significant enrichment of E2F1 target genes [16]. Rb, a tumor suppressor protein often mutated in MCL, binds to and blocks the activating E2F transcription factors [17, 18]. CDK4/6 may form a complex with cyclin D1 to phosphorylate Rb [19], resulting in E2F transactivation leading to transition of G1 cells to the S phase [19]. However, clinical trials of CDK inhibitors in MCL, such as Palbociclib [20] (CDK4/6 inhibitor), showed an overall response rate (ORR) of 18% and complete response (CR) of 8% in previously treated MCL [20].
Fibroblast growth factor receptors (FGFRs), transmembrane receptor tyrosine kinases, play an essential role in development, differentiation, cell survival, migration, angiogenesis, and carcinogenesis [21,22,23,24,25,26,27,28,29,30]. The FGFR pathway is the third most frequently altered pathway in cancer, with the most common abnormality being FGFR1 amplification [31]. Fusion of the B-cell receptor with FGFR1, utilizing the kinase activity of FGFR1, has been reported in ALL but not in MCL [32,33,34]. Clinical trials of CDK inhibitors in MCL that block RB phosphorylation to regulate E2F1 mediated transactivation, such as palbociclib [20] (CDK4/6 inhibitor), showed promise alone or in combination with ibrutinib in relapsed MCL [20]. Interestingly, FGFR1 signaling is shown to mediate resistance to Palbociclib in other disease models [35].
In summary, we identified a novel role of FGFR1 in MCL survival, regulating cell cycle-dependent processes primarily by activating E2F1-mediated transactivation through epigenetic repression of CDKN1C. We show that high FGFR1 expression is associated with a poor prognosis in MCL patients. Functionally, genetic knockdown of FGFR1 or pharmacological targeting with erdafitinib, a selective small molecule targeting FGFRs, induced cell cycle arrest, cell death in-vitro, reduced tumor formation, and improved overall survival in-vivo. Our findings point toward the FGFR1 signaling pathway as a critical modulator of cell survival and represent a novel and promising candidate for targeted therapy for patients with relapsed MCL.
Materials and Methods
Patient samples
Peripheral blood from healthy donors and peripheral blood, bone marrow samples, and lymph nodes from MCL patients were obtained from Leukemia Tissue Bank, OSU, following written informed consent under a protocol approved by the Institutional Review Board of OSU per the Declaration of Helsinki. MCL PDX cells were obtained from PROxe and were used for ex-vivo analysis.
Further detailed information on experimental procedures, methods, and materials is available in the Supplementary file.
Results
Elevated FGFR1 is associated with poor survival in MCL patients
Previously it has been shown that MCL–TME physical interactions with the stromal cells confer MCL survival and drug resistance [36]. To confirm whether physical cell contact was indispensable for MCL cell growth, we cultured patient-derived MCL cell lines in the presence or absence of stromal-conditioned media (CM) and investigated the impact on MCL cell proliferation and drug resistance. As shown in Fig. S1A, the culture of MCL cell lines, MCL PDX ex-vivo, and mouse MCL cells with bone marrow (BM) stromal CM (HS-5CM) significantly increased growth and abrogated response to drugs (doxorubicin, ibrutinib, and acalabrutinib) (Fig. S1B-C). To identify critical up-regulated genes driving MCL proliferation, bulk transcriptome analyses on RNA-seq data (p < 0.01; FC > 1.6) from (a) MCL patient vs. healthy-donor, (b) MCL ibrutinib-relapsed vs. treatment-naïve patients, (c) MCL with/without BM, and, (d) a chronic ibrutinib resistance MCL model described previously [36], were overlapped to identify up-regulated genes. The integrated analysis identified four genes (FGFR1, ABRACL, IFI27, and GEN) (Fig. S1D; Supplementary Table 1-2). Fibroblast Growth Factor Receptor-1 (FGFR1) showed minimal variation (coefficient of variation<6%) among MCL patients (n = 122) (Fig. S1E). However, FGFR1 is overexpressed at the transcript level in MCL Patients (n = 122) compared to naïve or activated-B cells from healthy donors (Fig. 1A) as analyzed in publicly available datasets (details in supplementary methods). In addition, FGFR1 levels were higher in MCL primary patient samples and MCL cell lines compared to healthy donor PBMC or B cells (Fig. 1B), primary MCL or MCL cell lines cultured with HS5CM, and resistant MCL cell lines described previously [36] (Fig. S1F). High FGFR1 expression in Rosenwald Cohort [37] treated with multiagent therapy(n = 92) or in Morin Cohort treated with R/CHOP (n = 30) show significant adverse effects on overall survival (OS) (Fig. 1C-D; Supplementary Table 3). Immunohistochemical staining of FGFR1 and Ki67 in MCL patients (n = 31) (Our cohort; Supplementary Table 3) uniformly treated with front-line R/CHOP and normal lymph node controls were performed and analyzed by a hematopathologist (Fig. 1E) for validation. Using the X-tile approach, the cut-off for FGFR1 expression (FGFR1+) was set at ≥10%; 18 (58%) of the 31 patients had high expression (FGFR1+), and 13 (42%) had low levels of FGFR1(FGFR1-). Retrospective analysis of MCL patients with FGFR1 expression showed significantly poorer OS (p = 0.001; Fig. 1F) and high Ki67 expression (61%) in MCL patients correlated with poor OS (p = 0.0007; Fig. 1G). Analyses of staining for FGFR1 and Ki67 in MCL patients revealed that MCL patient tumors with high FGFR1 expression significantly expressed higher Ki67 (Fig. S2A). To understand whether FGFR1’s prognostic effects depended on Ki67, we integrated FGFR1 and Ki67 protein data. In Ki67+ cases, FGFR1 continued to show significant adverse effects on overall survival (OS) with median overall survival of 1.75 years, compared to 6.4 years in Ki67+ FGFR1- patients (Fig. S2B). Further validation in Morin cohort [38] shows that MKI67 was associated with significant adverse prognostic effects (Fig. S2C), and in MKI67High cases, FGFR1 continued to show adverse effects on overall survival (OS) (Fig. S2D). Genomic analysis of the available 26 patients in the Morin cohort did not show any mutation in FGFR1 (supplementary table 3). Moreover, an extension of our analysis to other available studies [38, 39] in MCL (n = 162) did not reveal any activating mutation in the FGFR1 gene (Fig. S2E). Proliferation gene signature (PSG) in MCL is an average of the proliferative genes defined by Rosenwald, and we found that both FGFR1 and PSG continued to show significant adverse effects on overall survival in MCL Patients (n = 92) (Fig. 1C and Fig. S2F). Moreover, PSGHighFGFR1High MCL Patients show significant adverse effects on overall survival (OS) with median overall survival of 0.89 years, compared to 2.2 years in PSGHighFGFR1Low patients (Fig. S2G). Our findings validated in multiple patient cohorts suggest that FGFR1 has independent adverse prognostic effects on overall survival in MCL patients.
A FGFR1 expression is elevated in MCL patients (n = 122) compared to naïve or activated B-cells. Ordinary one-way ANOVA. B Expression of FGFR1 protein is elevated in MCL primary MCL (n = 4) three MCL PDX compared to two healthy donor Peripheral blood mononuclear cells (PBMC). Expression of FGFR1 protein is elevated in MCL cell lines compared to healthy donor CD19 + B-cells. Unpaired T-test. C Rosenwald cohort (LLMP) was analyzed for FGFR1 expression in MCL Patients (n = 92), and the Kaplan Meier plot for overall survival was generated and analyzed with the Log-rank Mantel-Cox test. D Morin Cohort was analyzed for FGFR1 expression MCL Patients who received CHOP/R (≤6 cycles) (n = 30), and Kaplan Meier plots for overall survival were generated and analyzed with the Log-rank Mantel-Cox test. E Representative images of Hematoxylin-eosin (H&E) staining and immunohistochemical (IHC) staining of Ki-67 and FGFR1 in MCL patient tumor samples from Young Cohort (n = 31). F FGFR1 or (G) Ki67 expression in MCL Patients (Young Cohort; n = 31) accessed by hematopathologists and Kaplan Meier plots for overall survival were generated and analyzed with the Log-rank Mantel-Cox test. H Tumor volume in NSG mice injected with FGFR1 VC vs. knockdown Z-138 (VC n = 5, shFGFR1 n = 6), Granta-519 (VC n = 9, shFGFR1 n = 9), and Jeko-1R (VC n = 5, shFGFR1 n = 5) cells, measured at ERC of first VC mouse. Unpaired T-test (I) Kaplan-Meier graphs to show the overall survival of NSG mice injected with either FGFR1 VC or knockdown Z-138, Granta-519, and Jeko-1R cells. p-values were calculated by the Log-rank Mantel-Cox test.
Loss of FGFR1 reduces tumorigenicity in-vitro and in-vivo
To test if FGFR1 contributes to MCL’s aggressive cell growth and survival, we generated knockdown clones in MCL cell lines (Z-138, Granta-519, Jeko-1R, and SP49-R) using different shRNA constructs (Fig. S2H-I). Next, to determine the role of FGFR1 in neoplastic progression in-vivo, we injected vector control (Vec Ctrl; VC) or the shFGFR1 clones of three representative MCL cell lines (Z-138, Granta-519, and Jeko-1R) into the flanks of NSG mice and measured tumor volume until the ERC of any experimental mice was reached. Consistent with our in-vitro studies, mice transplanted with shFGFR1 clones showed reduced tumor volume compared to vector controls (Fig. 1H) and exhibited a significantly prolonged median overall survival compared to control mice (shFGFR1 vs. Vec: Z138 [35 days vs. 28 days]; Granta-519 [32 days vs. 25]; Jeko-1-R [23 days vs. 15 days]) (Fig. 1I). However, despite this improvement, the engrafted mice eventually reached ERC (end-point of the study), in addition, the survival advantage for Jeko-1R was modestly significant. Our results demonstrate that FGFR1 is up-regulated in MCL, and depletion of FGFR1 alleviates tumor burden and improves survival in-vivo in MCL cell-derived xenograft models.
FGFR1 can be therapeutically targeted using a selective inhibitor of FGFRs
To explore the therapeutic potential of targeting FGFR1 in MCL, we first confirmed the expression of other FGFR family members (FGFR2-4) in MCL patients. Notably, other FGFR homologs (FGFR2-4) were not significantly up-regulated in MCL as compared to B cells (Fig. S3A-C), and FGFR1 was the only FGFR homolog significantly up-regulated in both MCL primary patient samples and cell lines (Fig. S3D-E). We could not detect the expression of FGFR2 using either Taqman or SYBR green Real-time PCR in MCL cell lines. Erdafitinib (JNJ-42756493) is an oral selective pan-FGFR kinase inhibitor [40, 41], with potent antitumor activity in pre-clinical studies exhibiting limited off-target effects on other kinases [40]. In-vitro anti-MCL activity of erdafitinib was demonstrated in MCL cell lines, which showed dose-dependent cellular death as determined by annexin-PI staining (Fig. S3F); further addition of HS5-CM failed to promote the proliferation of MCL cells in the presence of erdafitinib in-vitro (Fig. S3G-H). Since our in-vitro data suggested adding HS5-CM to MCL cells up-regulated FGFR1, we questioned a) whether stroma can promote MCL tumor progression in-vivo and b) if treatment with erdafitinib can abrogate stromal-mediated tumor progression in-vivo. To answer this, we utilized cell-line-derived xenograft (CDX) models of MCL cells (Jeko-1) injected subcutaneously in NSG mice with a) Jeko-1 cells alone, b) HS5 cells alone, and c) Jeko-1 and HS5 cells together, and measured tumor volume until the ERC of any experimental mice was reached. No tumor formation was observed in HS5 cells alone, confirming that normal human stromal cells do not exhibit neoplastic properties in-vivo. However, Jeko-1 cells injected with HS5 cells showed significantly higher tumor volume (p < 0.001; Fig. 2A) and significantly shorter overall survival (34 days) compared to control (41 days; Fig. 2B), suggesting the involvement of stromal cells in facilitating the growth of Jeko-1 cells in-vivo. Conversely, treatment with erdafitinib increased the overall survival of Jeko-1 cells injected subcutaneously in NSG mice with HS5 cells, and these mice had a statistically significant decrease in tumor volume (Fig. 2C-D). Next, the representative MCL cell lines Z-138, Granta-519, and Jeko-1 cells were independently injected subcutaneously into NSG mice flanks, followed by subsequent treatment with erdafitinib. We found erdafitinib treatment to limit tumor progression significantly (p < 0.001) and increase the overall survival in-vivo (p < 0.001; Fig. 2E-H). Next, we found that Jeko-1R cells reached ERC faster than Jeko-1 cells when implanted subcutaneously in NSG mice and that Jeko-1R have increased tumor volume compared to Jeko-1 injected mice (Fig. S3I-J). Furthermore, the tumor volume of the mice implanted with Jeko-1R treated with erdafitinib showed a modest reduction in tumor volume (Fig. 2G) and improved overall survival (Fig. 2H). Our results demonstrate that erdafitinib treatment in MCL alleviates tumor burden and improves survival in-vivo in MCL cell-derived xenograft models.
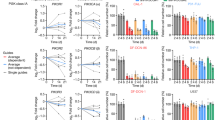
A Tumor volume at ERC of the first mouse was measured when Jeko-1 cells alone (n = 5) or combined with HS-5 cells (n = 5) were injected subcutaneously in NSG mice and subsequently treated with erdafitinib. HS-5 cells alone (n = 5) were injected in a separate group of mice as a control. B The Kaplan-Meier graph shows the survival time for mice described in (A). C Tumor volume at ERC of the first mouse was measured when NSG mice were injected subcutaneously with Jeko-1 cells in combination with HS-5 cells and subsequently treated with vehicle (n = 7) or erdafitinib (n = 7). D The Kaplan-Meier graph shows the survival time for mice described in (C). E Tumor volume at ERC of the first mouse was measured when NSG mice were injected subcutaneously with Z-138 and Granta-519 and subsequently treated with vehicle (Z138 n = 5, Granta-519 n = 6) or erdafitinib (Z138 n = 5, Granta-519 n = 7). F The Kaplan-Meier graph shows the survival time for mice described in (E). G Tumor volume at ERC of the first mouse was measured when NSG mice were injected subcutaneously with Jeko-1 and Jeko-1R and subsequently treated with vehicle (Jeko-1 n = 7, Jeko-1R n = 5) or erdafitinib (Jeko-1 n = 8, Jeko-1R n = 5). H The Kaplan-Meier graph shows the survival time for mice described in (G). Unpaired T-tests were used for tumor volume analysis, and p-values for survival analysis were calculated using the Log-rank (Mantel-Cox) test.
Loss of FGFR1 affects cell cycle progression and downregulates the expression of EZH2
Next, we performed gene expression profiling with knockdown of FGFR1 in two representative MCL cell lines (Z-138 and Granta-519) to identify common differentially expressed genes (DEG) (Fig. 3A; Supplementary Table 4). Further, KEGG 2021 human pathway analysis on DEG using enricher [42] identified the cell cycle and DNA replication as significantly altered pathways (Fig. 3B; Supplementary Table 5). Cell cycle dysregulation [14] and a defined PSG are MCL hallmarks [37, 43]. Furthermore, we found that FGFR1 loss in MCL down-regulates the expression of genes included in PSG (Fig. 3C) and show a higher number of cells in the G1 phase in shFGFR1 and erdafitinib MCL cells (Fig. 3D-E). Next, enricher-enabled ARCHS analysis [44] revealed EZH2, a histone methyl transferase and core member of the PRC2 complex, to be the most significant transcription factor co-expressed with DEG (Fig. 3F; Supplementary Table 6). Previously we have shown that EZH2 expression is up-regulated in MCL and is associated with poor prognosis [45]; hence we asked if FGFR1 regulates the expression of EZH2. We found EZH2 downregulated at both the RNA (Fig. S4A) and protein levels upon FGFR1 knockdown (Fig. 3G) or treatment with erdafitinib (Fig. 3H). Moreover, inhibiting EZH2 activity using the EZH2 inhibitor tazemetostat or with EZH1/2 dual inhibitor valemetostat increased the percentage of cells in the G1-phase (Fig. S4B). Notably, the expression of SUZ12 (Fig. S4C-D) and no other PRC1/2 members tested were consistently altered upon FGFR inhibition in MCL cell lines. Next, we confirmed that EZH2 forms a complex in MCL cells with PRC2(SUZ12)-PRC1(PCGF4) and PRC1.1 protein(KDM2B) (Fig. 3I). Moreover, we also found the expression of KDM2B to be downregulated in shFGFR1 MCL cells (Fig. S4E), and ectopic expression of KDM2B rescued the expression of EZH2 (Fig. S4F), confirming FGFR1 mediated regulation of KDM2B and EZH2 in MCL.
(A) Venn Diagram showing the differentially expressed gene in shFGFR1 vs. Vec ctrl analysis (p < 0.05, FC > 2) (B) Enrichment of the DEG obtained in panel A using Enrichr (C) Heatmap to show expression of MCL proliferation signature genes (PSG) in FGFR1 VC vs. knockdown cells of MCL cell lines Z-138 and Granta-519. (D) Cell cycle analysis determined the percentage of the G1 population in VC vs. shFGFR1 cells of Z-138 and Granta-519. (E) Cell cycle analysis determined the percentage of the G1 population in control vs. erdafitinib treated (16 h) cells of Z-138 and Granta-519. 2-way ANOVA (a = 0.05). (F) The hockey plot for the TF co-expressed with DEG was determined by ARCHS analysis. (G) Western blots show EZH2 expression in Z-138, Granta-519, Jeko-1R, and SP-49R control vs. shFGFR1 cells; EZH2 expression in Z138 was probed on the same blot from Fig. S2H. (H) Western blots show EZH2 expression in Z-138, Granta-519, Mino, and Jeko-1R cells upon erdafitinib treatment. (I) Co-immunoprecipitation of EZH2 and KDM2B (PRC1.1), SUZ12 (PRC2) or PCGF4 (PRC1) followed by western blot in Jeko-1R and Z-138 cells. 2% input was loaded with IP with appropriate IgG isotype control, followed by an IP-EZH2 sample.
EZH2 binds to the noncanonical PRC1.1 KDM2B and regulates the expression of the Cip/Kip2 protein CDKN1C
To identify genomic regions co-occupied by EZH2 and KDM2B, we performed Cut and Run chromatin immunoprecipitation (ChIP) followed by Next-gen sequencing using validated antibodies (Supplementary Table 7). As shown in Fig. S5A, EZH2, and KDM2B bind primarily to promoter regions. Overlay of both KDM2B and EZH2 peaks identified 1950 common peaks in the promoter regions ( ± 2KBTSS) (p < 0.0001) in MCL (Fig. 4A). Further enrichment analysis on the co-occupied peaks showed enrichment in cell cycle pathways (Fig. 4B; Supplementary Table 8), of which 14 peaks pertain to the promoters of genes involved in G1 to S cell cycle control (Fig. 4C), including a cell-cycle dependent kinase inhibitor 1C (CDKN1C) (Fig. 4D). In addition, we found that MCL patients that express high FGFR1 protein levels have higher expression of both KDM2B and EZH2 and lower levels of CDKN1C, as opposed to MCL patients expressing low FGFR1 (Fig. 4E). This observation, combined with the co-occupied binding of KDM2B and EZH2 on CDKN1C, led us to hypothesize that FGFR1 negatively regulates CDKN1C, primarily through epigenetic repression. Next, using ChIP with an H3K27me3-validated antibody followed by qRT-PCR, we show a repressive mark on the promoter of CDKN1C (Fig. S5B). Further, treatment of MCL cells with either tazemetostat or valemetostat restored CDKN1C expression(Fig. 4F). Since there are no specific KDM2B inhibitors, we performed transient knockdown of both KDM2B and EZH2 and found restoration of CDKN1C (Fig. S5C), similar to inhibition of EZH2, confirming a role of KDM2B and EZH2 to repress the expression of CDKN1C in MCL cells. Since FGFR1 loss downregulates the expression of both KDM2B and EZH2, we asked a) if loss or inhibition of FGFR1 in MCL can restore the expression of CDKN1C and b) if restoration of CDKN1C is needed for a high percentage of G1 Phase cells in MCL. Compared to controls, we found upregulation of CDKN1C protein expression in shRNA FGFR1 MCL cells (Fig. 4G) and upon treatment with erdafitinib (Fig. 4H). Further, erdafitinib treatment in MCL cells shows a higher percentage of cells in the G1-Phase, similar to shFGFR1 clones(Fig. 3D-E). To determine if the observed increase in G1 percentage in MCL cells is due to the restoration of CDKN1C, we knocked down CDKN1C (siRNA) in the presence of erdafitinib; as expected, CDKN1C loss significantly reduced the percent change of G1 cells in response to treatment (Fig. 4I, J). This role of CDKN1C in regulating G1 arrest in MCL agrees with previously published studies [46,47,48] in other disease models, strengthening our observation.
A Co-occupied peaks of KDM2B and EZH2 in Z138 cells (within 2KB of TSS. B Enriched pathways analysis using Enrichr for the co-occupied 1950 peaks from panel A showing enrichment of cell cycle pathways. C Cell cycle-related loci (129) were functionally distinguished using enricher to identify 14 genes with functional relation to G1/S transition, including CDKN1C. D Genomic browser (hg19) for enrichment KDM2B and EZH2 on the target CDKN1C Loci with an overlay of ATAC in Z138 cells. E Heatmap showing Relative Densitometric Unit (RDU) calculated from Western Blot expression of FGFR1, KDM2B, EZH2, and CDKN1C in FGFR1 high vs. low MCL patient samples. F Western blot depicting CDKN1C increase in expression upon tazemetostat and valemetostat treatment in Z-138, Jeko-1R, and SP-49R cells. Veh: vehicle control. G Western blots to show CDKN1C expression in Z-138, Granta-519, Jeko-1R, and SP-49R control vs. shFGFR1 cells. H Western blots to show EZH2 expression in Z-138, Granta-519, Jeko-1R, and SP-49R cells upon erdafitinib treatment. I Percent increase in G1 cell percentage in Z-138 and (J) Granta-519 cells upon transfection with scrambled si-RNA (control) and si-CDKN1C and subsequent treatment with erdafitinib. 2-way ANOVA (a = 0.05).
CDKN1C binds to E2F1 and regulates the transactivation of its target genes
CDKN1C (p57Kip2) is a tumor suppressor gene and inhibits cyclin-CDK complex activity [46], thereby regulating the retinoblastoma RB-E2F pathway [49]. Since Rb phosphorylation and its subsequent dissociation from transcription factor E2F1 (Fig. 5A) is a significant determinant of cell-cycle progression from the G1-phase [50], we found Rb-phosphorylation was reduced upon FGFR1 loss in MCL cells compared to controls (Fig. 5B). Similar results were obtained by treating MCL cells with erdafitinib (Fig. 5C). As shown in Fig. 5D, transactivation of the representative bona fide E2F1 target genes (CDK1, TK1, UHRF1, PCNA, TUBA1B [51,52,53,54]), which are also part of the PSG in MCL, were down-regulated in shFGFR1 cells. Similar results were obtained by treating the MCL cells with erdafitinib (Fig. 5E), strengthening our observation that loss or inhibition of FGFR1 regulates E2F1-mediated transactivation. Further, ectopic expression of EZH2 in erdafitinib-treated MCL cells restored the expression of phosphorylation of RB, KDM2B, and downregulated CDKN1C protein levels (Fig. 5F). As expected, the loss of CDKN1C in erdafitinib-treated MCL cells restored the expression of RB and its phosphorylation (Fig. 5G) and upregulation in E2F1 target gene expression (Fig. 5H) with no change in levels of E2F1 (Fig. S6A). We further confirmed that CDKN1C immunoprecipitated E2F1 in MCL cells (Fig. 5I), suggesting that CDKN1C may regulate the expression of E2F1-mediated transactivation by direct binding and act as a repressor of E2F1 function.
A Model showing Rb-E2F1 regulation by Cyclin-CDK complex. B Blots show decreased Rb phosphorylation upon FGFR1 knockdown in Z-138 and Granta-519 cell lines. (C) Blots show a decreased Rb phosphorylation upon erdafitinib treatment in Z-138 and Granta-519 cell lines, Actin for Z-138 and Granta-519 in (B) is the same as Fig. 3G, and in (C) is the same as Fig. 3H (different exposures), as the same blots were probed for Rb and EZH2 (D) Heatmap for qPCR analysis depicting the decrease in expression of E2F1 target genes upon FGFR1 knockdown in Z-138 and Granta-519 cell lines. E Heatmap for qPCR analysis depicts decreased expression of E2F1 target genes upon erdafitinib treatment in Z-138 and Granta-519 cell lines. 2-way ANOVA (a = 0.05). F Blots of the rescue of EZH2 expression upon transfection of EZH2 overexpression plasmid in the presence of erdafitinib and the corresponding change in CDKN1C, KDM2B, phospho-Rb expression in Z-138 and Granta-519 cell lines; EZH2-OE (-) lane is a vector control transfected lane, complete blot and exposures used in Suppl Fig. S7A and S7B. G Protein expression of CDKN1C, EZH2, KDM2B, and phospho-Rb upon CDKN1C knockdown in the presence of erdafitinib in Z-138 and Granta-519 cell lines; si-CDKN1C (-) lane is scrambled si-RNA transfected lane. H Heatmap for qPCR analysis in Z-138 cells shows rescue of E2F1 target genes previously downregulated by erdafitinib by now knocking-down CDKN1C in the presence of erdafitinib. I Co-immunoprecipitation of CDKN1C and E2F1 followed by western blot in Granta-519 and Z-138 cells. Model showing proposed hypothesis of CDKN1C mediated regulation of E2F1 target genes.
FGFR1 contributes to a feedback loop to regulate MYC stability, which regulates the EZH2-CDKN1C axis
MYC regulates the expression of EZH2 [55] and KDM2B [56]; hence, we asked if EZH2 and KDM2B expression in erdafitinib-treated MCL is MYC dependent. As shown in Fig. 6A, MYC introduction in erdafitinib-treated MCL cells causes higher levels of expression of KDM2B and EZH2 and subsequent loss of CDKN1C along with reactivation of CDK1 (RB/E2F1 target), suggesting that FGFR1 mediated regulation of KDM2B/EZH2/CDKN1C is MYC dependent. Interestingly, the expression of MYC is significantly lower in either shRNA FGFR1 or erdafitinib-treated MCL cells compared to control (Fig. 6B-C, Fig. S6D), but the expression of MYC was unaltered (Fig. S6B-C). To investigate if the loss of MYC expression was due to a decreased stability of MYC protein with FGFR1 inhibition, we treated MCL cells with cycloheximide and determined that MYC degraded faster in the presence of erdafitinib (Fig. 6D-E). Further, treatment with the proteasomal inhibitor MG132 rescued the expression of MYC in erdafitinib-treated MCL cells (Fig. 6F). The phosphorylation on Serine 62 and Threonine 58 residues largely governs MYC stability, and a higher Ser62/Thr58 ratio indicates greater MYC stability [57,58,59]. We found that the MYC p-Ser(62)/p-Thr(58) ratio is reduced within 6 h of the addition of erdafitinib (Fig. 6G-H). Further, motif scan analysis on MYC found an S/TPx(x)R/K consensus motif on Serine 62 with high-affinity binding scores to two kinases, Erk1 and CDK1 (Fig. 6I). Both p-Erk1/2 and CDK1 levels are reduced upon FGFR1 knockdown and erdafitinib treatment (Fig. 6J, K; Fig. S6E-F). Activated Erk has been established to increase MYC stabilization through Ser62 phosphorylation [57, 60]. When MCL cells were treated with U0126, a MEK1/2 inhibitor that inhibits Erk’s activation, and we found a decrease in MYC expression, consistent with previous observations (Fig. S6G). CDK1 is downregulated in FGFR1 inhibition/loss in MCL cells (Fig. 6J-K) and restored by EZH2 overexpression (Fig. 6L) or CDKN1C downregulation (Fig. 6M) in erdafitinib treated MCL cells. Further, inhibiting the activity of CDK1 using RO-3306, a selective CDK1 inhibitor, we observed subsequent reduction in MYC levels (Fig. 6N) and the MYC p-Ser62/p-Thr58 ratio (Fig. S6H), bolstering the finding that CDK1 regulates MYC stability. Next, the MYC levels were restored by proteasomal inhibition using MG-132 in RO-3306 treated cells (Fig. 6O) or U0126 treated MCL cells (Fig. S6I). The orthogonal validation of our axis was performed in an ex-vivo culture of isolated PDX splenocytes treated with erdafitinib, confirming the loss of MYC, KDM2B, EZH2, CDK1, pERK1/2, pRB, and upregulation of CDKN1C, strengthening our in-vitro results (Fig. 7A; Fig. S6J). Further analysis of the RNA Seq data from healthy donors and MCL primary patients showed a lower expression of CDKN1C and higher expression of FGFR1 and E2F target genes (CDK1, PCNA, UHRF1, and TUBA1B) in MCL primary patients (Fig. 7B).
A Z-138 and Granta-519 cells were transfected with MYC overexpression plasmid in the presence of erdafitinib treatment, and expression of EZH2, KDM2B, CDKN1C, phospho-Rb, CDK1 was evaluated. MYC-OE (-) lane is vector control transfected alone, complete blot and exposures used in Suppl Fig. S7A and S7B. B MYC protein expression upon FGFR1 knockdown and (C) erdafitinib treatment in Z-138 and Granta-519 cells, Actin in (B) is same as Fig. 5B; the same blot was used for MYC and Rb status in this case. D Z-138 and (E) Granta-519 cells were treated with 100ug/ml cycloheximide (CHX) for 20 min, with or without pre-treatment with erdafitinib. MYC protein expression was subsequently evaluated. F Z-138 and Granta-519 cells were treated with erdafitinib and supplied with 10uM proteasomal inhibitor MG-132. MYC expression was then determined. erdafitinib alone treatment lane for Granta-519 was split for representation; See complete blot in Suppl Fig. S7C; G Blots showing phospho-MYC Ser62 and Thr58 expression and ratio upon six hr. treatment with erdafitinib in Z-138 and (H) Granta-519 cells. I ScanSite 4.0 motif scan of Ser62 kinase motif of the human MYC protein revealed Erk1 kinase and CDK1 as top hits with the highest score. J CDK1 protein expression in FGFR1 knockdown and (K) erdafitinib treated cells of Z-138 and Granta-519. Actin in (J) is the same as in Fig. 3G, as the same blots were used for re-probing EZH2 and CDK1 in these cases. L Blots showing CDK1 expression upon EZH2 rescue in the presence of erdafitinib in Z-138 and Granta-519; complete blot and exposures used in Suppl Fig. S7A and S7B. M Blots showing CDK1 expression upon CDKN1C knockdown in the presence of erdafitinib in Z-138 and Granta-519. Note: Actin in (L) and (M) is the same as Fig. 5F and G, respectively, as the same blot was used for probing CDK1 in this case. N MYC expression upon treatment of Z-138 and Granta-519 cells with CDK1 inhibitor RO-3306. O Z-138 and Granta-519 cells were treated with RO-3306 and supplied with 10uM proteasomal inhibitor MG-132. MYC protein expression was then determined. RO-3306 alone treatment lane for Granta-519 was split for representation; See complete blot in Suppl Fig. S7D.
A Western blots showing expression of p-Erk1/2, MYC, p-Rb, EZH2, KDM2B, and CDKN1C upon erdafitinib treatment of 3 MCL Patient-derived xenografts ex-vivo. B Tukey plots showing FGFR1, CDKN1C, TUBA1B, CDK1, UHRF1, and PCNA expression in 20 MCL patients compared to healthy donor B-cells. C Model showing the role of FGFR1 regulating feedback loop in MCL.
Discussion
FGFR1 activation is a known driver in various malignant diseases such as breast, prostate, myeloproliferative neoplasms, AML, ALL, CML, glioblastoma, and lung cancer [61,62,63,64,65]. Therapeutic targeting of FGFRs in cancers is an active area of research. TKI targeting the kinase domain of FGFRs is currently being tested in clinical trials [66, 67]. In particular, the FGFR-selective TKIs, including JNJ42756493 (erdafitinib), BGJ398 (NVP-BGJ398), and AZD4547, are currently under clinical evaluation on different oncotypes [66,67,68]. Our data here support that the selective inhibitor erdafitinib has pre-clinical activity in-vivo in murine models of MCL and presents an opportunity to characterize the potential of TKI targeting in MCL patients. FGF2-mediated activation of FGFR1 promotes resistance to EGFR inhibitors in lung cancer [69], FLT3 inhibitors in AML [70], and KIT inhibitors in gastrointestinal stromal tumors [71]. Noteworthy, FGFR1 can be activated via various ligands (FGF1-6, FGF-8, 19,21, and 23) [72], and previous studies of leukemia and lymphoma cells show higher expression of FGF2 ligands [73]. Recently, a study [74] shows higher levels of FGFR1 in MCL patients who did not respond to Ibrutinib. Our data show that FGFR1 expression correlates with poor prognosis in MCL patients who received CHOP/R-based therapy.
Similarly, the Ki-67 score alone stratified patient outcome, and in Ki67+ cases, FGFR1 continued to show significant adverse effects on overall survival. The data suggest that the prognostic effect of FGFR1 is independent of Ki67, but a future comparative analysis of FGFR1 and Ki-67 index using a more extensive but similarly treated patient cohort is warranted. Additionally, in a larger cohort, the adverse prognostic effects of FGFR1 were observed in cases with high proliferation gene signature (PSG) compared to PSGlow, suggesting that FGFR1 has independent adverse prognostic effects on overall survival in MCL patients. Recent findings of FIGHT-101 [75] using pemigatinib shows that the most frequent alterations co-occurring with FGFR amplifications were TP53 alterations, however genomic analysis of MCL patients (n = 162) did not show any FGFR1 activating mutations. In addition, our data also support that FGFR1 expression can be regulated by stromal cells, thus postulating that both cell intrinsic and extrinsic factors contribute to FGFR1 expression and its activity in MCL.
Coordinated efforts of the polycomb repressor complexes PRC1 and PRC2 are known to maintain epigenetic repression at gene target loci. Canonical PRC1 contains four core subunits: PCGF4, PCGF2, PHC, CBX, and RING1A or RING1B, with E3 ubiquitin ligase activity on histone H2A at lysine-119, thereby inducing the compaction of chromatin [76]. In addition non-canonical PRC1-variants (PRC1.1,PRC1.3,PRC1.5, and PRC1.6) containing a distinct PCGF subunit have been identified [77]. Interestingly, we did not observe any change in protein expression of other PRC1 members other than the PRC1.1 member KDM2B in erdafitinib-treated MCL cells, raising the possibility of KDM2B regulation by FGFR1 in MCL. Ectopic expression of KDM2B in FGFR1 knockdown MCL cells restored EZH2 expression, confirming KDM2B to be downstream of FGFR1, consistent with previous reports [78].
Conversely, the loss of FGFR1 released suppression on CDKN1C, this epigenetic repression of CDKN1C in MCL is consistent with a previous report [79]. Our results indicate an intricate regulation of KDM2B and EZH2 to regulate the repression of CDKN1C in an FGFR1-dependent manner. Interestingly loss of CDKN1C in erdafitinib-treated cells also up-regulated the expression of EZH2, confirming the regulation of Rb-E2F1 transactivation by CDKN1C, as EZH2 expression is regulated by E2F1 [80]. On the contrary, CDKN1C loss in erdafitinib treated cells failed to up-regulate the expression of KDM2B, suggesting that its regulation may be Rb-E2F independent, possibly through EZH2, as ectopic expression of EZH2 restored the expression of KDM2B in erdafitnib treated MCL cells (Fig. 5F). However, a deeper investigation is warranted to delineate whether recruitment of KDM2B and EZH2 to target loci follows a ‘hierarchical’ model or ‘alternative model’ of chromatin assembly in MCL [76, 81,82,83,84].
Our observation is consistent with a previous report that shows G1 arrest upon inhibition of EZH2 through reactivation of p57(CDKN1C) [85]. Interestingly, upon FGFR1 inhibition, levels of total Rb protein were also reduced and subsequently increased upon rescue by EZH2 or loss of CDKN1C, suggesting that RB might be FGFR1 dependent. Our findings indicate that CDKN1C binding to E2F1 in MCL cells may be a repressor of E2F1-mediated transactivation. A recent report discusses transcriptional reprogramming via CDK9 in IR MCL [86], and another report shows that EZH1/2 inhibition overcomes ibrutinib resistance in MCL primarily by up-regulating CDKN1C [79]. The previous report shows CDKN1C binding to CDK7/9 inhibits transcription by preventing RNA Pol-II CTD phosphorylation [49]. Our data here show that FGFR1 signaling contributes to the suppression of CDKN1C via epigenetic regulation in MCL and may be targeted in ibrutinib-resistant patients, but this hypothesis requires further validation.
Overexpression of MYC has been associated with an aggressive blastoid MCL variant [87, 88], and MYC cooperates with Cyclin D1 to drive mouse lymphomagenesis [15, 89, 90]. Consistent with these reports, the introduction of MYC in erdafitinib-treated cells resulted in up-regulated EZH2 and KDM2B. Interestingly ectopic expression of EZH2 and loss of CDKN1C also increased levels of MYC (Fig. S6K), suggesting that epigenetic regulation in MCL may constitute a feedback loop. Further, we show MYC stability depends on phosphorylation at serine 62 [60], and either FGFR1 or CDK1 inhibition in MCL cells decreased S62 phosphorylation. In addition, our data indicate that the downstream mediator of FGFR1 signaling, pERK1, also promotes MYC stability, which eventually results in CDKN1C-mediated regulation of CDK1 and can, in turn, regulate the stability of MYC, resulting in the maintenance of a feedback loop. This possibly explains why CDKN1C downregulation in the presence of the FGFR1 inhibitor also increases MYC levels.
In summary, our data provide a model of cellular proliferation in MCL that is dependent upon FGFR1 levels and shows that FGFR1 loss up-regulates the expression of CDKN1C via MYC/KDM2B/EZH2 axis and regulates E2F target gene expression such as CDK1, which forms a feedback loop to maintain MYC levels and provide a proliferative advantage to MCL cells (Fig. 7C). Further, we show that FGFR1 is therapeutically targetable with erdafitinib to achieve lower tumor burdens in multiple in-vivo MCL models to improve overall survival. This report provides the first compelling pre-clinical rationale for targeting FGFR1 in MCL.
Availability of data materials
Data sets used or generated in this study can be accessed through the GEO portal using the accession numbers mentioned in respective methods.
Data availability
Data sets used or generated in this study can be accessed through the GEO portal using the accession numbers mentioned in respective methods.
References
A clinical evaluation of the International Lymphoma Study Group classification of non-Hodgkin’s lymphoma. The Non-Hodgkin’s Lymphoma Classification Project. Blood. 1997;89:3909–18.
Vose JM. Mantle cell lymphoma: 2013 Update on diagnosis, risk-stratification, and clinical management. Am J Hematol. 2013;88:1082–8.
Scott DW, Gascoyne RD. The tumour microenvironment in B cell lymphomas. Nat Rev Cancer. 2014;14:517–34.
Tiemann M, Schrader C, Klapper W, Dreyling MH, Campo E, Norton A, et al. Histopathology, cell proliferation indices and clinical outcome in 304 patients with mantle cell lymphoma (MCL): a clinicopathological study from the European MCL Network. Br J Haematol. 2005;131:29–38.
Abrahamsson A, Albertsson-Lindblad A, Brown PN, Baumgartner-Wennerholm S, Pedersen LM, D’Amore F, et al. Real world data on primary treatment for mantle cell lymphoma: a Nordic Lymphoma Group observational study. Blood. 2014;124:1288–95.
Zhou Y, Wang H, Fang W, Romaguer JE, Zhang Y, Delasalle KB, et al. Incidence trends of mantle cell lymphoma in the United States between 1992 and 2004. Cancer. 2008;113:791–8.
Kumar A, Sha F, Toure A, Dogan A, Ni A, Batlevi CL, et al. Patterns of survival in patients with recurrent mantle cell lymphoma in the modern era: progressive shortening in response duration and survival after each relapse. Blood Cancer J. 2019;9:50.
Martin P, Maddocks K, Leonard JP, Ruan J, Goy A, Wagner-Johnston N, et al. Postibrutinib outcomes in patients with mantle cell lymphoma. Blood. 2016;127:1559–63.
Cheah CY, Chihara D, Romaguera JE, Fowler NH, Seymour JF, Hagemeister FB, et al. Patients with mantle cell lymphoma failing ibrutinib are unlikely to respond to salvage chemotherapy and have poor outcomes. Ann Oncol. 2015;26:1175–9.
Cheah CY, Seymour JF, Wang ML. Mantle cell lymphoma. J Clin Oncol. 2016;34:1256–69.
Epperla N, Hamadani M, Cashen AF, Ahn KW, Oak E, Kanate AS, et al. Predictive factors and outcomes for ibrutinib therapy in relapsed/refractory mantle cell lymphoma-a “real world” study. Hematol Oncol. 2017;35:528–35.
Sircar A, Chowdhury SM, Hart A, Bell WC, Singh S, Sehgal L, et al. Impact and intricacies of bone marrow microenvironment in B-cell lymphomas: from biology to therapy. Int J Mol Sci. 2020;21:904.
Chiron D, Martin P, Di Liberto M, Huang X, Ely S, Lannutti BJ, et al. Induction of prolonged early G1 arrest by CDK4/CDK6 inhibition reprograms lymphoma cells for durable PI3Kδ inhibition through PIK3IP1. Cell Cycle. 2013;12:1892–1900.
Determann O, Hoster E, Ott G, Wolfram Bernd H, Loddenkemper C, Leo Hansmann M, et al. Ki-67 predicts outcome in advanced-stage mantle cell lymphoma patients treated with anti-CD20 immunochemotherapy: results from randomized trials of the European MCL Network and the German Low Grade Lymphoma Study Group. Blood. 2008;111:2385–7.
Bodrug SE, Warner BJ, Bath ML, Lindeman GJ, Harris AW, Adams JM. Cyclin D1 transgene impedes lymphocyte maturation and collaborates in lymphomagenesis with the myc gene. EMBO J. 1994;13:2124–30.
Edwards DR, Kuo P-Y, Lagana A, Park S, Nedumaran P, Leshchenko V, et al. Aberrant cell cycle programming confers rapid lethality in the EuSOX11+ CCND1 MCL mouse model. Blood. 2020;136:6–7.
Walter DM, Yates TJ, Ruiz-Torres M, Kim-Kiselak C, Gudiel AA, Deshpande C, et al. RB constrains lineage fidelity and multiple stages of tumour progression and metastasis. Nature. 2019;569:423–7.
Wander SA, Cohen O, Gong X, Johnson GN, Buendia-Buendia JE, Lloyd MR, et al. The genomic landscape of intrinsic and acquired resistance to cyclin-dependent kinase 4/6 inhibitors in patients with hormone receptor-positive metastatic breast cancer. Cancer Discov. 2020;10:1174–93.
Roskoski R Jr. Cyclin-dependent protein serine/threonine kinase inhibitors as anticancer drugs. Pharmacol Res. 2019;139:471–88.
Martin P, Bartlett NL, Blum KA, Park S, Maddocks K, Ruan J, et al. A phase 1 trial of ibrutinib plus palbociclib in previously treated mantle cell lymphoma. Blood. 2019;133:1201–4.
Dvorak P, Dvorakova D, Hampl A. Fibroblast growth factor signaling in embryonic and cancer stem cells. FEBS Lett. 2006;580:2869–74.
Grose R, Dickson C. Fibroblast growth factor signaling in tumorigenesis. Cytokine Growth Factor Rev. 2005;16:179–86.
Korc M, Friesel RE. The role of fibroblast growth factors in tumor growth. Current Cancer Drug Targets. 2009;9:639–51.
Moroni E, Dell’Era P, Rusnati M, Presta M. Fibroblast growth factors and their receptors in hematopoiesis and hematological tumors. J Hematother Stem Cell Res. 2002;11:19–32.
Ornitz DM, Itoh N. The fibroblast growth factor signaling pathway. Wiley Interdisc Rev Dev Biol. 2015;4:215–66.
Pazgal I, Zimra Y, Tzabar C, Okon E, Rabizadeh E, Shaklai M, et al. Expression of basic fibroblast growth factor is associated with poor outcome in non-Hodgkin’s lymphoma. Br J Cancer. 2002;86:1770–5.
Turner N, Grose R. Fibroblast growth factor signalling: from development to cancer. Nat Rev Cancer. 2010;10:116–29.
Xian W, Pappas L, Pandya D, Selfors LM, Derksen PW, de Bruin M, et al. Fibroblast growth factor receptor 1-transformed mammary epithelial cells are dependent on RSK activity for growth and survival. Cancer Res. 2009;69:2244–51.
Zhang X, Ibrahimi OA, Olsen SK, Umemori H, Mohammadi M, Ornitz DM. Receptor specificity of the fibroblast growth factor family. The complete mammalian FGF family. J Biol Chem. 2006;281:15694–15700.
Zhang Y, Lin Y, Bowles C, Wang F. Direct cell cycle regulation by the fibroblast growth factor receptor (FGFR) kinase through phosphorylation-dependent release of Cks1 from FGFR substrate 2. J Biol Chem. 2004;279:55348–54.
Helsten T, Elkin S, Arthur E, Tomson BN, Carter J, Kurzrock R. The FGFR landscape in cancer: analysis of 4,853 tumors by next-generation sequencing. Clinical Cancer Research. 2016;22:259–67.
Agerstam H, Jaras M, Andersson A, Johnels P, Hansen N, Lassen C, et al. Modeling the human 8p11-myeloproliferative syndrome in immunodeficient mice. Blood. 2010;116:2103–11.
Peiris MN, Meyer AN, Nelson KN, Bisom-Rapp EW, Donoghue DJ. Oncogenic fusion protein BCR-FGFR1 requires the breakpoint cluster region-mediated oligomerization and chaperonin Hsp90 for activation. Haematologica. 2020;105:1262–73.
Wakim JJ, Tirado CA, Chen W, Collins R. t(8;22)/BCR-FGFR1 myeloproliferative disorder presenting as B-acute lymphoblastic leukemia: report of a case treated with sorafenib and review of the literature. Leuk Res. 2011;35:e151–153.
Formisano L, Lu Y, Servetto A, Hanker AB, Jansen VM, Bauer JA, et al. Aberrant FGFR signaling mediates resistance to CDK4/6 inhibitors in ER+ breast cancer. Nature Communications. 2019;10:1373.
Zhao X, Lwin T, Silva A, Shah B, Tao J, Fang B, et al. Unification of de novo and acquired ibrutinib resistance in mantle cell lymphoma. Nat Commun. 2017;8:14920.
Rosenwald A, Wright G, Wiestner A, Chan WC, Connors JM, Campo E, et al. The proliferation gene expression signature is a quantitative integrator of oncogenic events that predicts survival in mantle cell lymphoma. Cancer Cell. 2003;3:185–97.
Pararajalingam P, Coyle KM, Arthur SE, Thomas N, Alcaide M, Meissner B, et al. Coding and noncoding drivers of mantle cell lymphoma identified through exome and genome sequencing. Blood. 2020;136:572–84.
Nadeu F, Martin-Garcia D, Clot G, Díaz-Navarro A, Duran-Ferrer M, Navarro A, et al. Genomic and epigenomic insights into the origin, pathogenesis, and clinical behavior of mantle cell lymphoma subtypes. Blood. 2020;136:1419–32.
Perera TPS, Jovcheva E, Mevellec L, Vialard J, De Lange D, Verhulst T, et al. Discovery and pharmacological characterization of JNJ-42756493 (Erdafitinib), a functionally selective small-molecule FGFR family inhibitor. Mol Cancer Ther. 2017;16:1010–20.
Markham A. Erdafitinib: First Global Approval. Drugs. 2019;79:1017–21.
Xie Z, Bailey A, Kuleshov MV, Clarke DJB, Evangelista JE, Jenkins SL, et al. Gene set knowledge discovery with enrichr. Current Protocols. 2021;1:e90.
Queiros AC, Beekman R, Vilarrasa-Blasi R, Duran-Ferrer M, Clot G, Merkel A, et al. Decoding the DNA methylome of mantle cell lymphoma in the light of the entire B cell lineage. Cancer Cell. 2016;30:806–21.
Lachmann A, Torre D, Keenan AB, Jagodnik KM, Lee HJ, Wang L, et al. Massive mining of publicly available RNA-seq data from human and mouse. Nature Communications. 2018;9:1366.
Wang X, Sehgal L, Jain N, Khashab T, Mathur R, Samaniego F. LncRNA MALAT1 promotes development of mantle cell lymphoma by associating with EZH2. J Transl Med. 2016;14:346.
Matsuoka S, Edwards MC, Bai C, Parker S, Zhang P, Baldini A, et al. p57KIP2, a structurally distinct member of the p21CIP1 Cdk inhibitor family, is a candidate tumor suppressor gene. Genes Dev. 1995;9:650–62.
Sherr CJ, Roberts JM. CDK inhibitors: positive and negative regulators of G1-phase progression. Genes Dev. 1999;13:1501–12.
Samuelsson MKR, Pazirandeh A, Davani B, Okret S. p57Kip2, a Glucocorticoid-Induced Inhibitor of Cell Cycle Progression in HeLa Cells. Molecular Endocrinology. 1999;13:1811–22.
Ma Y, Chen L, Wright GM, Pillai SR, Chellappan SP, Cress WD. CDKN1C negatively regulates RNA polymerase II C-terminal domain phosphorylation in an E2F1-dependent manner. J Biol Chem. 2010;285:9813–22.
Qin XQ, Livingston DM, Ewen M, Sellers WR, Arany Z, Kaelin WG Jr. The transcription factor E2F-1 is a downstream target of RB action. Mol Cell Biol. 1995;15:742–55.
Wells J, Graveel CR, Bartley SM, Madore SJ, Farnham PJ. The identification of E2F1-specific target genes. Proc Nat Acad Sci. 2002;99:3890–5.
Magri L, Swiss VA, Jablonska B, Lei L, Pedre X, Walsh M, et al. E2F1 coregulates cell cycle genes and chromatin components during the transition of oligodendrocyte progenitors from proliferation to differentiation. J Neurosci. 2014;34:1481–93.
Polager S, Kalma Y, Berkovich E, Ginsberg D. E2Fs up-regulate expression of genes involved in DNA replication, DNA repair and mitosis. Oncogene. 2002;21:437–46.
Bracken AP, Ciro M, Cocito A, Helin K. E2F target genes: unraveling the biology. Trends Biochem Sci. 2004;29:409–17.
Wu X, Liu D, Tao D, Xiang W, Xiao X, Wang M, et al. BRD4 regulates EZH2 transcription through upregulation of C-MYC and represents a novel therapeutic target in bladder cancer. Mol Cancer Therap. 2016;15:1029–42.
Tzatsos A, Paskaleva P, Lymperi S, Contino G, Stoykova S, Chen Z, et al. Lysine-specific Demethylase 2B (KDM2B)-let-7-Enhancer of Zester Homolog 2 (EZH2) pathway regulates cell cycle progression and senescence in primary cells. J Biolog Chem. 2011;286:33061–9.
Sears R, Nuckolls F, Haura E, Taya Y, Tamai K, Nevins JR. Multiple Ras-dependent phosphorylation pathways regulate Myc protein stability. Genes Dev. 2000;14:2501–14.
Yada M, Hatakeyama S, Kamura T, Nishiyama M, Tsunematsu R, Imaki H, et al. Phosphorylation-dependent degradation of c-Myc is mediated by the F-box protein Fbw7. EMBO J. 2004;23:2116–25.
Welcker M, Orian A, Jin J, Grim JE, Harper JW, Eisenman RN, et al. The Fbw7 tumor suppressor regulates glycogen synthase kinase 3 phosphorylation-dependent c-Myc protein degradation. Proc Natl Acad Sci USA. 2004;101:9085–90.
Lee T, Yao G, Nevins J, You L. Sensing and integration of Erk and PI3K signals by Myc. PLoS Comput Biol. 2008;4:e1000013.
Acevedo VD, Gangula RD, Freeman KW, Li R, Zhang Y, Wang F, et al. Inducible FGFR-1 activation leads to irreversible prostate adenocarcinoma and an epithelial-to-mesenchymal transition. Cancer Cell. 2007;12:559–71.
Nguyen PT, Tsunematsu T, Yanagisawa S, Kudo Y, Miyauchi M, Kamata N, et al. The FGFR1 inhibitor PD173074 induces mesenchymal-epithelial transition through the transcription factor AP-1. Br J Cancer. 2013;109:2248–58.
Pacini L, Jenks AD, Lima NC, Huang PH. Targeting the fibroblast growth factor receptor (FGFR) family in lung cancer. Cells. 2021;10:1154.
Inhorn RC, Aster JC, Roach SA, Slapak CA, Soiffer R, Tantravahi R, et al. A syndrome of lymphoblastic lymphoma, eosinophilia, and myeloid hyperplasia/malignancy associated with t(8;13)(p11;q11): description of a distinctive clinicopathologic entity. Blood. 1995;85:1881–7.
Sohal J, Chase A, Mould S, Corcoran M, Oscier D, Iqbal S, et al. Identification of four new translocations involving FGFR1 in myeloid disorders. Genes Chromosomes Cancer. 2001;32:155–63.
Kommalapati A, Tella SH, Borad M, Javle M, Mahipal A. FGFR inhibitors in oncology: insight on the management of toxicities in clinical practice. Cancers. 2021;13:2968.
Katoh M. FGFR inhibitors: Effects on cancer cells, tumor microenvironment and whole-body homeostasis (Review). Int J Mol Med. 2016;38:3–15.
Xie Y, Su N, Yang J, Tan Q, Huang S, Jin M, et al. FGF/FGFR signaling in health and disease. Signal Transd Targ Ther. 2020;5:181.
Ware KE, Hinz TK, Kleczko E, Singleton KR, Marek LA, Helfrich BA, et al. A mechanism of resistance to gefitinib mediated by cellular reprogramming and the acquisition of an FGF2-FGFR1 autocrine growth loop. Oncogenesis. 2013;2:e39.
Traer E, Martinez J, Javidi-Sharifi N, Agarwal A, Dunlap J, English I, et al. FGF2 from marrow microenvironment promotes resistance to FLT3 inhibitors in acute myeloid leukemia. Cancer Res. 2016;76:6471–82.
Li F, Huynh H, Li X, Ruddy DA, Wang Y, Ong R, et al. FGFR-mediated reactivation of MAPK signaling attenuates antitumor effects of imatinib in gastrointestinal stromal tumors. Cancer Discov. 2015;5:438–51.
Ferguson HR, Smith MP, Francavilla C. Fibroblast growth factor receptors (FGFRs) and noncanonical partners in cancer signaling. Cells. 2021;10:1201.
Krejci P, Faitova J, Laurell H, Hampl A, Dvorak P. FGF-2 expression and its action in human leukemia and lymphoma cell lines. Leukemia. 2003;17:818–20.
Zhang L, Yao Y, Zhang S, Liu Y, Guo H, Ahmed M, et al. Metabolic reprogramming toward oxidative phosphorylation identifies a therapeutic target for mantle cell lymphoma. Sci Transl Med. 2019;11:eaau1167.
Subbiah V, Iannotti NO, Gutierrez M, Smith DC, Féliz L, Lihou CF, et al. FIGHT-101, a first-in-human study of potent and selective FGFR 1-3 inhibitor pemigatinib in pan-cancer patients with FGF/FGFR alterations and advanced malignancies. Ann Oncol. 2022;33:522–33.
Wang H, Wang L, Erdjument-Bromage H, Vidal M, Tempst P, Jones RS, et al. Role of histone H2A ubiquitination in Polycomb silencing. Nature. 2004;431:873–8.
Gao Z, Zhang J, Bonasio R, Strino F, Sawai A, Parisi F, et al. PCGF homologs, CBX proteins, and RYBP define functionally distinct PRC1 family complexes. Mol Cell. 2012;45:344–56.
Kottakis F, Polytarchou C, Foltopoulou P, Sanidas I, Kampranis SC, Tsichlis PN. FGF-2 regulates cell proliferation, migration, and angiogenesis through an NDY1/KDM2B-miR-101-EZH2 pathway. Mol Cell. 2011;43:285–98.
Kagiyama Y, Fujita S, Shima Y, Yamagata K, Katsumoto T, Nakagawa M, et al. CDKN1C-mediated growth inhibition by an EZH1/2 dual inhibitor overcomes resistance of mantle cell lymphoma to ibrutinib. Cancer Sci. 2021;112:2314–24.
Bracken AP, Pasini D, Capra M, Prosperini E, Colli E, Helin K. EZH2 is downstream of the pRB-E2F pathway, essential for proliferation and amplified in cancer. EMBO J. 2003;22:5323–35.
Cao R, Wang L, Wang H, Xia L, Erdjument-Bromage H, Tempst P, et al. Role of histone H3 lysine 27 methylation in Polycomb-group silencing. Science. 2002;298:1039–43.
Min J, Zhang Y, Xu RM. Structural basis for specific binding of Polycomb chromodomain to histone H3 methylated at Lys 27. Genes Dev. 2003;17:1823–8.
Farcas AM, Blackledge NP, Sudbery I, Long HK, McGouran JF, Rose NR, et al. KDM2B links the Polycomb Repressive Complex 1 (PRC1) to recognition of CpG islands. Elife. 2012;1:e00205.
Sashida G, Oshima M, Iwama A. Deregulated Polycomb functions in myeloproliferative neoplasms. Int J Hematol. 2019;110:170–8.
Li W, Bi C, Han Y, Tian T, Wang X, Bao H, et al. Targeting EZH1/2 induces cell cycle arrest and inhibits cell proliferation through reactivation of p57(CDKN1C) and TP53INP1 in mantle cell lymphoma. Cancer Biol Med. 2019;16:530–41.
Zhao X, Wang MY, Jiang H, Lwin T, Park PM, Gao J, et al. Transcriptional programming drives Ibrutinib-resistance evolution in mantle cell lymphoma. Cell Rep. 2021;34:108870.
Hernandez L, Hernandez S, Bea S, Pinyol M, Ferrer A, Bosch F, et al. c-myc mRNA expression and genomic alterations in mantle cell lymphomas and other nodal non-Hodgkin’s lymphomas. Leukemia. 1999;13:2087–93.
Zhao X, Lwin T, Zhang X, Huang A, Wang J, Marquez VE, et al. Disruption of the MYC-miRNA-EZH2 loop to suppress aggressive B-cell lymphoma survival and clonogenicity. Leukemia. 2013;27:2341–50.
Adams JM, Harris AW, Pinkert CA, Corcoran LM, Alexander WS, Cory S, et al. The c-myc oncogene driven by immunoglobulin enhancers induces lymphoid malignancy in transgenic mice. Nature. 1985;318:533–8.
Lovec H, Grzeschiczek A, Kowalski MB, Moroy T. Cyclin D1/bcl-1 cooperates with myc genes in the generation of B-cell lymphoma in transgenic mice. EMBO J. 1994;13:3487–95.
Acknowledgements
We thank the Flow Cytometry, Target validation, Cellular Imaging Facility, Leukemia Tissue Bank, and DNA Analysis shared resource at The Ohio State University Comprehensive Cancer Center, Columbus, OH. We thank Dr. Christopher Smith for the Fc-uMCL1 cells. We thank Janssen Pharmaceuticals for providing erdafitinib.
Funding
Research reported in this publication was supported by the National Cancer Institute of the National Institutes of Health under Award Numbers R01CA282483 (LS), 5KL2TR2734 (LS), R01CA233490, R01CA187415, R01CA138688, and RC1CA146299 (ZYXM and KHY), P01CA214274-01A1 (RAB, LA), IRP (Young investigator award to LS), Pelotonia Graduate fellowship to AS, Pelotonia Postdoctoral fellowship to SS, Ramalingaswami grant (BT/RLF/Re-entry/18/2017) from the DBT, India (NJ), American Society of Hematology Global Research Award (NJ), an operating grant from the Canadian Institutes for Health Research and a New Investigator Award from the Canadian Institutes for Health Research (RD.M). RDM. is a Michael Smith Foundation for Health Research Scholar. KMC is a recipient of the Scholarship for the Next Generation of Scientists from the Cancer Research Society. Research reported in this publication was supported by The Ohio State University Comprehensive Cancer Center and the National Institutes of Health under grant number P30 CA016058, supporting OSU Flow Cytometry, Target validation, Cellular Imaging Facility, Leukemia Tissue Bank, DNA Analysis Core Facility, and IGM Genomic shared resource. The IGM Genomic service core is also supported by Nationwide Foundation Pediatric Innovation Fund.
Author information
Authors and Affiliations
Contributions
AS and SS designed, performed most experiments, and analyzed the data. KMC, LK, and RDM. performed RNA seq and Genomic analysis on MCL Patients. NJ, EC, and PT performed certain experiments and analyzed the data, ZYXM and KHY performed IHC and clinical correlation, and PR, BRP, RM, JT, WH, NE, and RAB, provided critical suggestions for research. LS contributed to the overall research design; LS and AS contributed to the writing of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Sircar, A., Singh, S., Xu-Monette, Z.Y. et al. Exploiting the fibroblast growth factor receptor-1 vulnerability to therapeutically restrict the MYC-EZH2-CDKN1C axis-driven proliferation in Mantle cell lymphoma. Leukemia 37, 2094–2106 (2023). https://doi.org/10.1038/s41375-023-02006-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41375-023-02006-8
This article is cited by
-
Beyond Bruton’s tyrosine kinase inhibitors in mantle cell lymphoma: bispecific antibodies, antibody–drug conjugates, CAR T-cells, and novel agents
Journal of Hematology & Oncology (2023)