Abstract
Tyrosine kinase inhibitors (TKIs) targeting epidermal growth factor receptors (EGFR) significantly prolong the survival of lung adenocarcinoma patients with sensitizing EGFR mutations. Unfortunately, 10–30% patients do not show objective responses to EGFR-TKIs, and undergo rapid disease progression during the EGFR-TKIs therapy. Single nucleotide polymorphisms (SNPs) in mature microRNA (miRNA) sequences may influence target site interactions and modulate downstream pathways, such as the EGFR pathway. For this reason, we hypothesized that miRNA SNPs may impact the prognosis of lung adenocarcinoma patients after EGFR-TKI treatment. By systematically screening of the miRbase and the 1000 genomes project databases, we successfully identified five mature miRNA SNPs. Genotypes were determined in two independent cohorts (Hubei and Shandong cohorts) that include 319 EGFR-TKI treated stage IIIB/IV patients. The impact of miR-608 and miR-4513 on the drug sensitivity of gefitinib was examined in lung adenocarcinoma cells. miR-608 rs4919510 or miR-4513 rs2168518 significantly contributed to the progression-free survival (PFS) in the Hubei cohort (hazard ratio [HR] = 0.63, confidence interval [CI] = 0.49–0.81, P = 3.0 × 10−4 or HR = 0.46, 95% CI = 0.31–0.67, P = 8.0 × 10−5). These observations were further validated in the Shandong cohort (P = 0.005 or P = 0.001). Similarly, the miR-608 rs4919510 CC genotype or the miR-4513 rs2168518 GA genotype was significantly associated with decreased death risk after gefitinib treatment, compared with the rs4919510 GG genotype (Hubei cohort: P = 5.0 × 10−4; Shandong cohort: P = 0.004) or the rs2168518 GG genotype (P = 4.9 × 10−5; P = 0.002). Consistently, miR-608 significantly increased the anti-proliferation effect of gefitinib in both lung adenocarcinoma PC9 and H1299 cells, whereas miR-4513 increased cells’ resistance to gefitinib. Our findings suggest that miR-608 and miR-4513 SNPs are independent candidate biomarkers to predict lung adenocarcinoma patients’ survival after EGFR-TKIs treatment. These miRNAs and polymorphisms provide clinical potential in patient-tailored treatment decision-making.
Similar content being viewed by others
Introduction
Lung cancer is a leading cancer and has one of the highest number of annually estimated newly diagnosed cases among all cancers. In addition, lung cancer ranks the no. 1 cancer-related mortality among malignancies in the world [1]. Patients diagnosed as advanced non-squamous non-small cell lung cancer (NSCLC) have extremely poor overall survival (OS) ranged from 8 months to 10 months [2, 3]. For this group of patients with prevalent activating somatic mutations in the epidermal growth factor receptor (EGFR) gene, EGFR-tyrosine kinase inhibitors (EGFR-TKIs) are recommended as the first-line therapy, which has dramatically changed the landscape of therapeutic management of non-squamous NSCLC [4,5,6]. Gefitinib as an FDA approved first-generation EGFR-TKI has been widely used in patients harboring the EGFR exon 19 deletions or exon 21 (Leu858Arg) substitution mutation and has significantly improved disease prognosis [7, 8]. Unfortunately, 10–30% of lung adenocarcinoma patients do not show objective responses, and may undergo rapid disease progression during the EGFR-TKI therapy [4,5,6, 9]. Tumor heterogeneity is one of the most important factors associated with the treatment failure of NSCLC [10]. Accumulating evidence suggests that germline genetic polymorphisms are involved in EGFR-TKI resistance, and can significantly influence the OS of patients [11, 12]. Despite efforts that have been made, reliable markers to guide individualized treatment are still lacking, and the prognosis of lung adenocarcinoma patients remains poor. Therefore, biomarkers in addition to the EGFR mutations need to be identified to predict patients’ response to the EGFR-TKIs treatment.
As evolutionarily conserved short non-coding RNAs (19–23 nucleotides [nt] in length), microRNAs (miRNAs) function essentially in negative, post-transcriptional gene regulation via targeting the 3′-untranslated region of mRNAs with the seed region of miRNAs [13, 14]. One miRNA can bind to hundreds of target mRNAs that regulate essential biological processes [15]. Profiles of miRNAs have been extensively correlated with the development and progression of various cancers [16, 17]. Accumulating evidence has shown that miRNAs can be used as predictive or prognostic biomarkers for lung cancer treatments [18,19,20]. Single nucleotide polymorphisms (SNPs) in the mature sequence of miRNA, especially in the seed region, can alter target site interactions by either strengthening or weakening hybridization kinetics. Interestingly, multiple SNPs located within mature miRNA sequences are significantly associated with lung cancer susceptibility and prognosis of lung cancer patients treated with chemotherapy [21,22,23].
However, it remains unclear whether miRNA SNPs impact the prognosis of advanced lung adenocarcinoma patients that harbor activating EGFR mutations and are treated with gefitinib. Herein, we specifically examined the role of five miRNA polymorphisms (miR-499 rs3746444, miR-608 rs4919510, miR-3152 rs13299349, miR-4513 rs2168518, and miR-4520a rs8078913) on predicting the prognosis of lung adenocarcinoma patients after gefitinib therapy in two cohorts. In addition, to validate the biological functions of two miRNAs (miR-608 and miR-4513), we determined their roles on the gefitinib sensitivity in different lung adenocarcinoma cell lines.
Materials and methods
Study subjects
This study consisted of two cohorts including 319 stage IIIB or IV lung adenocarcinoma patients with prevalent EGFR activating somatic mutations. (a) Hubei cohort: 162 lung adenocarcinoma patients were recruited at Tongji Hospital, Tongji Medical College, Huazhong University of Science and Technology (Wuhan, Hubei Province, China) between January 2009 and May 2015. (b) Shandong cohort: 157 lung adenocarcinoma patients were recruited at Shandong Cancer Hospital affiliated to Shandong University (Jinan, Shandong Province, China) between May 2011 and October 2015. All patients were histologically confirmed lung adenocarcinoma with clinical data available. Each patient was treated with gefitinib orally at a daily dose of 250 mg as the first line monotherapy. Progression-free survival (PFS) was defined as the time from the date of diagnosis to the date of disease progression, or death due to any cause. OS was calculated from the date of diagnosis to the date of death from any cause or the latest follow-up. All subjects were ethnic Han Chinese. This study was approved by the Institutional Review Boards of Tongji Hospital and Shandong Cancer Hospital affiliated to Shandong University. At recruitment, written informed consent was obtained from each patient. The methods were carried out in accordance with the approved guidelines.
Genotyping
After systematically screening the miRbase database and the 1000 genomes project Phase 3 CHB (Chinese Han in Beijing) data, we successfully identified five SNPs in human mature miRNA sequences (miR-499 rs3746444, miR-608 rs4919510, miR-3152 rs13299349, miR-4513 rs2168518, and miR-4520a rs8078913) with minor allele frequencies (MAF) > 0.05 in Chinese [24]. Genotypes of these miRNA polymorphisms in both Hubei cohort and Shandong cohort were determined using the iPLEX Sequenom MassARRAY platform (Sequenom) as previously reported [25,26,27]. A 15% random sample size was reciprocally tested and the reproducibility was 100%.
Cell culture and reagents
Lung adenocarcinoma H1299 and PC9 cells were cultured at 37 °C with 5% CO2 using RPMI-1640 media with 10% fetal bovine serum (Hyclone). miRNA mimics of miR-608 and miR-4513 as well as the negative control RNA duplex (NC) for miRNA mimics were synthesized by Genepharma (Shanghai, China). NC RNA was nonhomologous to any human genome sequence. Small RNAs were transfected into H1299 and PC9 cells with INTERFERin® (Polyplus). Gefitinib was purchased from Amquar Co. (EY0917).
Quantitative reverse transcription PCR (qRT-PCR)
Total RNA was isolated from H1299 and PC9 cells with Trizol reagent (Invitrogen). To avoid contamination of genomic DNA, each RNA sample was treated with RNase-Free DNase to remove genomic DNA (Invitrogen). The corresponding cDNAs were generated using the Revert Ace kit (TOYOBO, Osaka, Japan). Human miR-608, miR-4513, and U6 small RNA were examined with their specific stem-loop RT-PCR primers (Ribobio, Guangzhou, China) using SYBR-Green qRT-PCR as described previously [28, 29].
Cell proliferation and gefitinib drug sensitivity assays
H1299 and PC9 cells were seeded at 7500 cells per well in 96-well plates and transfected with 20 nmol/L miR-608 mimics, miR-4513 mimics or NC RNA (Genepharma), respectively. The day after seeding, gefitinib diluted in DMSO was added to the wells to achieve the desired final concentrations (H1299: 1, 5, 10, 25, and 50 μM; PC9: 1, 5, 10, 20, and 30 μM). H1299 or PC9 cells were incubated with gefitinib at different concentrations for 48 h and survival rates were measured using the MTT assay (Sigma) according to manufacturer’s instructions.
Statistics
The differences of patient clinical characteristics were estimated with Student’s t test or χ2 test. The Kaplan–Meier method was used to examine the genotype impact on PFS or OS. Survival curves were compared using the log-rank test. Multivariate Cox regression analysis was used to access prognostic factors for PFS or OS. Hazard ratios (HRs) and 95% confidence intervals (CIs) for the association between the miRNA SNPs and recurrence risk or death risk were adjusted for age of onset, sex, smoking, and stage status, where it was appropriate. A P value of less than 0.05 was used as the criterion of statistical significance. All statistical tests were two-sided. All analyses were performed with GraphPad or SPSS software package.
Results
Patient characteristics and clinical features
The demographic characteristics and clinical information of the lung adenocarcinoma patients from the two cohorts are summarized in Supplementary Table 1. All patients received gefitinib target therapy. In the Hubei discovery cohort, the median age was 56 years (range, 37–80 years), and there were 90 males (55.6%) and 66 smokers (40.7%). Among these patients with positive EGFR mutations, 30 (18.6%) had stage IIIB diseases and 132 (81.4%) had stage IV diseases. At the final analysis, 142 patients (87.7%) died and the median OS time was 17.2 months (range, 0.6–90.0 months). The median PFS time was 10.4 months (range, 0.1–90.0 months). In the Shandong validation cohort, the median age was 57 years (range, 29–82 years), 81 patients (51.6%) were male, 62 (39.5%) were smokers, 28 (17.8%) had stage IIIB diseases, and 129 (82.2%) had stage IV diseases. At the final analysis, 130 patients (82.8%) died and the median OS or PFS time was 16.2 months (range, 1.0–78.0 months) or 9.9 months (range, 0.4–76.3 months).
Effect of miRNA genetic polymorphisms on PFS
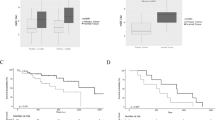
Cox regression analyses were used to access the association of miRNA polymorphisms with PFS of lung adenocarcinoma patients treated with gefitinib in both the discovery cohort and the validation cohort (Table 1). In the discovery cohort, miR-608 rs4919510, miR-4513 rs2168518, and miR-4520a rs8078913 were significantly associated with better or worse PFS. However, only miR-608 rs4919510 and miR-4513 rs2168518 were significantly associated with PFS in both the discovery cohort and validation cohort (Table 1). The miR-608 rs4919510 CC genotype was a significantly favorable PFS factor (Hubei cohort: HR = 0.63, 95% CI = 0.49–0.81, P = 3.0 × 10−4; Shandong cohort: HR = 0.63, 95% CI = 0.45–0.87, P = 0.005). The miR-4513 rs2168518 GA genotype was also associated with better PFS (Hubei cohort: HR = 0.46, 95% CI = 0.31–0.67, P = 8.0 × 10−5; Shandong cohort: HR = 0.40, 95% CI = 0.24–0.68, P = 0.001). Lung adenocarcinoma patients carrying the miR-608 rs4919510 GC or GG genotype had a shorter PFS than patients carrying the CC genotype (both log-rank P < 0.001) (Fig. 1a, b). In addition, patients with the miR-4513 rs2168518 GG genotype had a shorter PFS than patients carrying the GA or AA genotype (Hubei cohort: log-rank P < 0.001; Shandong cohort: log-rank P = 0.003) (Fig. 2a, b). The other three SNPs were not significantly associated with PFS in either cohorts (Supplementary Figure 1-3).
Impact of miRNA genetic polymorphisms on OS
As shown in Table 2, multivariate Cox proportional hazards model showed that only the miR-608 rs4919510 CC or miR-4513 rs2168518 GA genotype was significantly associated with decreased death risk after gefitinib treatment, compared with the miR-608 rs4919510 GG genotype (Hubei cohort: HR = 0.65, 95% CI = 0.51–0.83, P = 5.0 × 10−4; Shandong cohort: HR = 0.59, 95% CI = 0.41–0.85, P = 0.004) or the miR-4513 rs2168518 GG genotype (Hubei cohort: HR = 0.45, 95% CI = 0.31–0.66, P = 4.9 × 10−5; Shandong cohort: HR = 0.38, 95% CI = 0.21–0.69, P = 0.002). Kaplan–Meier survival estimates showed that patients with the miR-608 rs4919510 GC or GG genotype had a worse OS than patients carrying the CC genotype (Hubei cohort: log-rank P = 0.002; Shandong cohort: log-rank P = 0.001) (Fig. 1c, d). Similarly, individuals with the miR-4513 rs2168518 GG genotype had a worse survival than patients carrying the GA or AA genotype (Hubei cohort: log-rank P < 0.001; Shandong cohort: log-rank P = 0.004) (Fig. 2c, d).
miR-608 and miR-4513 oppositely impact drug sensitivity of gefitinib
To further determine the function of miR-608 and miR-4513, we determined if they impact the drug sensitivity of lung adenocarcinoma PC9 and H1299 cells to gefitinib. As shown in Fig. 3, miR-608 significantly increased the anti-proliferation effects of gefitinib in both PC9 and H1299 cells (P < 0.05). On the contrary, although significantly decreased cell viability was observed in lung adenocarcinoma cells treated with gefitinib, miR-4513 rescued the inhibition effect. PC9 and H1299 cells with overexpressed miR-4513 mimics showed obvious resistance to gefitinib (P < 0.05) (Fig. 4).
miR-608 significantly promoted the anti-proliferation effect of gefitinib in PC9 cells (a) and H1299 cells (b). Elevated miR-608 expression was observed in both cell lines (c). All experiments were performed in triplicates in three independent transfection experiments and each value represents mean ± standard deviation (SD). *P < 0.05, **P < 0.01
miR-4513 significantly increased the drug resistance to gefitinib in PC9 (a) and H1299 cells (b). Elevated miR-4513 expression was observed in both cell lines (c). All experiments were performed in triplicates in three independent transfection experiments and each value represents mean ± SD. *P < 0.05, ***P < 0.001
Discussion
EGFR-TKIs have dramatically changed the landscape of the therapeutic strategy of non-squamous NSCLC patients with sensitive EGFR mutation. Nevertheless, more than 10% of sensitive EGFR mutated patients do not respond to the EGFR-TKIs treatment [30]. The IPASS study has shown that during gefitinib treatment, 20.5% of patients with EGFR mutations stayed stable disease and 7.6% of them showed progressive disease [9]. This observation indicates that further studies are required to identify factors impacting the sensitivity of lung adenocarcinoma patients to EGFR-TKIs.
The crucial roles of miRNAs on cancer biology have gradually been revealed. The potential applications of using miRNA as predictive or prognostic biomarkers have generated great interests in cancer risk factor evaluation and prognosis estimation [31,32,33,34]. Genetic variants such as SNPs in miRNAs, especially in their seed region, can significantly transform the nature of a miRNA and change its target library, resulting in altered expression of target mRNA and various functional endings [35]. A series of epidemiologic studies have revealed that the miR-608 rs4919510 SNP has a vital role on the etiology and prognosis of cancer [36,37,38,39]. For example, after genotyping six polymorphisms of miRNAs in 575 patients with lung cancer and 608 healthy controls in a non-smoking female population in northeast China, Yin et al. found that miR-608 rs4919510 was not associated with lung cancer risk in any models. In a meta-analysis including 18 case–control studies (2517 cancer cases and 15,624 controls), Wu et al. observed a significant association between the miR-608 rs4919510 polymorphism and cancer risk in Chinese (OR = 1.11; 95% CI = 1.04–1.19). In further stratified analyses based on cancer types, miR-608 rs4919510 exhibited borderline significant associations with an increased risk of lung cancer (OR = 1.14; 95% CI = 0.99–1.32). miR-4513 rs2168518 polymorphism was reported to be associated with cardiovascular diseases and elderly eye diseases [40, 41]. However, the importance of these two miRNA SNPs in the survival of lung adenocarcinoma patients who received gefitinib therapy was unclear.
In our present study, we found that miR-608 rs4919510 and miR-4513 rs2168518 were significantly associated with prognosis of lung adenocarcinoma patients after gefitinib treatment in two cohorts. Patients carrying the homozygous CC genotype of miR-608 rs4919510 had significantly longer OS and PFS, compared with those carrying the GG genotype. Patients with the heterozygous GA genotype of miR-4513 rs2168518 had better prognosis relative to the GG genotype. To our knowledge, this study provides the first evidence that the rs4919510 polymorphism in miR-608 and the rs2168518 polymorphism in miR-4513 can serve as prognostic biomarkers for lung adenocarcinoma during gefitinib therapy.
These results are biologically plausible considering the tumor suppressor nature of miR-608 in NSCLC [42]. In addition, our drug sensitivity results showed that miR-608 significantly enhances the anti-proliferation effect of gefitinib in both PC9 and H1299 cells. Consistent to our findings, Othman et al. reported that miR-608 could significantly induced cell death of A549 and SK-LU1 cells in comparison to negative control cells [43]. Accumulated evidence has demonstrated that miR-608 plays a critical role in EGFR and its downstream signaling. For instance, AKT2, a crucial regulator of the mitogen activated protein kinase (MAPK) and phosphatidylinositol 3-kinase/protein kinase B (PI3K/AKT) pathways, both of which are downstream of EGFR signaling, has been identified as a novel target of miR-608 in lung adenocarcinoma [42]. Similarly, miRNA-608 inhibits the proliferation of bladder cancer cells via the AKT/FOXO3a signaling pathway [44]. In addition, miR-608 directly targets oncogenes EGFR and Bcl-xL in chordoma [45].
It remains largely unknown how miR-4513 is involved in tumorigenesis and the prognosis of lung cancer. Results from our functional experiments suggest that miR-4513 functions like an oncogene and increases the resistance of lung adenocarcinoma cells to gefitinib. Ghanbari et al. reported that rs2168518 in the seed region of miR-4513 shows pleiotropic effect on lipid and glucose homeostasis, blood pressure and coronary artery disease [41]. There are two mechanisms through which rs2168518 affects miR-4513 function [41]. First, the rs2168518 A allele decreases the expression level of miR-4513 by reducing miRNA processing, and second, the A allele reduces the ability of miR-4513 to interact with its target genes [41]. In line with these findings, we demonstrated that patients with rs2168518 GA or AA genotypes (possible reduced oncogene miR-4513 expression in cells) showed better prognosis compared to carriers of the GG genotype.
In conclusion, our results provide the first evidence that miR-608 rs4919510 and miR-4513 rs2168518 polymorphisms significantly contribute to lung adenocarcinoma patients’ survival to gefitinib in two cohorts of Chinese Han population. The identified miRNA polymorphisms could serve as useful prognostic biomarkers and be applied to improve the treatment strategy for patients with lung adenocarcinoma.
References
Siegel RL, Miller KD, Jemal A. Cancer statistics, 2018. CA Cancer J Clin. 2018;68:7–30.
Liu KJ, Guan ZZ, Liang Y, Yang XQ, Peng J, Huang H, et al. A double-blind, randomized phase II study of dicycloplatin plus paclitaxel versus carboplatin plus paclitaxel as first-line therapy for patients with advanced non-small-cell lung cancers. Arch Med Sci. 2014;10:717–24.
Hotta K, Matsuo K, Ueoka H, Kiura K, Tabata M, Tanimoto M. Addition of platinum compounds to a new agent in patients with advanced non-small-cell lung cancer: a literature based meta-analysis of randomised trials. Ann Oncol. 2004;15:1782–9.
Zhou C, Wu YL, Chen G, Feng J, Liu XQ, Wang C, et al. Erlotinib versus chemotherapy as first-line treatment for patients with advanced EGFR mutation-positive non-small-cell lung cancer (OPTIMAL, CTONG-0802): a multicentre, open-label, randomised, phase 3 study. Lancet Oncol. 2011;12:735–42.
Maemondo M, Inoue A, Kobayashi K, Sugawara S, Oizumi S, Isobe H, et al. Gefitinib or chemotherapy for non-small-cell lung cancer with mutated EGFR. N Engl J Med. 2010;362:2380–8.
Rosell R, Carcereny E, Gervais R, Vergnenegre A, Massuti B, Felip E, et al. Erlotinib versus standard chemotherapy as first-line treatment for European patients with advanced EGFR mutation-positive non-small-cell lung cancer (EURTAC): a multicentre, open-label, randomised phase 3 trial. Lancet Oncol. 2012;13:239–46.
Mitsudomi T, Morita S, Yatabe Y, Negoro S, Okamoto I, Tsurutani J, et al. Gefitinib versus cisplatin plus docetaxel in patients with non-small-cell lung cancer harbouring mutations of the epidermal growth factor receptor (WJTOG3405): an open label, randomised phase 3 trial. Lancet Oncol. 2010;11:121–8.
Douillard JY, Ostoros G, Cobo M, Ciuleanu T, McCormack R, Webster A, et al. First-line gefitinib in Caucasian EGFR mutation-positive NSCLC patients: a phase-IV, open-label, single-arm study. Br J Cancer. 2014;110:55–62.
Mok TS, Wu YL, Thongprasert S, Yang CH, Chu DT, Saijo N, et al. Gefitinib or carboplatin-paclitaxel in pulmonary adenocarcinoma. N Engl J Med. 2009;361:947–57.
De Sousa EMF, Vermeulen L, Fessler E, Medema JP. Cancer heterogeneity—a multifaceted view. EMBO Rep. 2013;14:686–95.
Ng KP, Hillmer AM, Chuah CT, Juan WC, Ko TK, Teo AS, et al. A common BIM deletion polymorphism mediates intrinsic resistance and inferior responses to tyrosine kinase inhibitors in cancer. Nat Med. 2012;18:521–8.
Yuan J, Li B, Zhang N, Zhu H, Zhou L, Zhang L, et al. Clinical implications of the BIM deletion polymorphism in advanced lung adenocarcinoma treated with gefitinib. Clin Lung Cancer. 2018;19:e431–8.
Kong YW, Ferland-McCollough D, Jackson TJ, Bushell M. microRNAs in cancer management. Lancet Oncol. 2012;13:e249–58.
Bartel DP. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell. 2004;116:281–97.
Wu L, Belasco JG. Let me count the ways: mechanisms of gene regulation by miRNAs and siRNAs. Mol Cell. 2008;29:1–7.
Esquela-Kerscher A, Slack FJ. Oncomirs—microRNAs with a role in cancer. Nat Rev Cancer. 2006;6:259–69.
Kumar MS, Lu J, Mercer KL, Golub TR, Jacks T. Impaired microRNA processing enhances cellular transformation and tumorigenesis. Nat Genet. 2007;39:673–7.
Luo P, Yang Q, Cong LL, Wang XF, Li YS, Zhong XM, et al. Identification of miR124a as a novel diagnostic and prognostic biomarker in nonsmall cell lung cancer for chemotherapy. Mol Med Rep. 2017;16:238–46.
Zhang Y, Li H, Han J, Zhang Y. Down-regulation of microRNA-124 is correlated with tumor metastasis and poor prognosis in patients with lung cancer. Int J Clin Exp Pathol. 2015;8:1967–72.
Saito M, Shiraishi K, Matsumoto K, Schetter AJ, Ogata-Kawata H, Tsuchiya N, et al. A three-microRNA signature predicts responses to platinum-based doublet chemotherapy in patients with lung adenocarcinoma. Clin Cancer Res. 2014;20:4784–93.
Pu X, Roth JA, Hildebrandt MA, Ye Y, Wei H, Minna JD, et al. MicroRNA-related genetic variants associated with clinical outcomes in early-stage non-small cell lung cancer patients. Cancer Res. 2013;73:1867–75.
Qiu F, Yang L, Ling X, Yang R, Yang X, Zhang L, et al. Sequence variation in mature microRNA-499 confers unfavorable prognosis of lung cancer patients treated with platinum-based chemotherapy. Clin Cancer Res. 2015;21:1602–13.
Xie K, Chen M, Zhu M, Wang C, Qin N, Liang C, et al. A polymorphism in miR-1262 regulatory region confers the risk of lung cancer in Chinese population. Int J Cancer. 2017;141:958–66.
Zheng J, Deng J, Xiao M, Yang L, Zhang L, You Y, et al. A sequence polymorphism in miR-608 predicts recurrence after radiotherapy for nasopharyngeal carcinoma. Cancer Res. 2013;73:5151–62.
Yang X, Yu D, Ren Y, Wei J, Pan W, Zhou C, et al. Integrative functional genomics implicates EPB41 dysregulation in hepatocellular carcinoma risk. Am J Hum Genet. 2016;99:275–86.
Shi M, Xia J, Xing H, Yang W, Xiong X, Pan W, et al. The Sp1-mediaded allelic regulation of MMP13 expression by an ESCC susceptibility SNP rs2252070. Sci Rep. 2016;6:27013.
Shi M, Ma F, Liu J, Xing H, Zhu H, Yu J, et al. A functional BRCA1 coding sequence genetic variant contributes to prognosis of triple-negative breast cancer, especially after radiotherapy. Breast Cancer Res Treat. 2017;166:109–16.
Ren Y, Shang J, Li J, Liu W, Zhang Z, Yuan J, et al. The long noncoding RNA PCAT-1 links the microRNA miR-215 to oncogene CRKL-mediated signaling in hepatocellular carcinoma. J Biol Chem. 2017;292:17939–49.
Ren Y, Chen Y, Liang X, Lu Y, Pan W, Yang M. miRNA-638 promotes autophagy and malignant phenotypes of cancer cells via directly suppressing DACT3. Cancer Lett. 2017;390:126–36.
Chan SK, Gullick WJ, Hill ME. Mutations of the epidermal growth factor receptor in non-small cell lung cancer—search and destroy. Eur J Cancer. 2006;42:17–23.
Cho WC, Chow AS, Au JS. Restoration of tumour suppressor hsa-miR-145 inhibits cancer cell growth in lung adenocarcinoma patients with epidermal growth factor receptor mutation. Eur J Cancer. 2009;45:2197–206.
Arora S, Ranade AR, Tran NL, Nasser S, Sridhar S, Korn RL, et al. MicroRNA-328 is associated with (non-small) cell lung cancer (NSCLC) brain metastasis and mediates NSCLC migration. Int J Cancer. 2011;129:2621–31.
Mizuno K, Seki N, Mataki H, Matsushita R, Kamikawaji K, Kumamoto T, et al. Tumor-suppressive microRNA-29 family inhibits cancer cell migration and invasion directly targeting LOXL2 in lung squamous cell carcinoma. Int J Oncol. 2016;48:450–60.
Liu C, Luo J, Zhao YT, Wang ZY, Zhou J, Huang S, et al. TWIST1 upregulates miR-214 to promote epithelial-to-mesenchymal transition and metastasis in lung adenocarcinoma. Int J Mol Med. 2018;42:461–70.
Ryan BM, Robles AI, Harris CC. Genetic variation in microRNA networks: the implications for cancer research. Nat Rev Cancer. 2010;10:389–402.
Sclafani F, Chau I, Cunningham D, Lampis A, Hahne JC, Ghidini M, et al. Sequence variation in mature microRNA-608 and benefit from neo-adjuvant treatment in locally advanced rectal cancer patients. Carcinogenesis. 2016;37:852–7.
Yin Z, Cui Z, Ren Y, Xia L, Wang Q, Zhang Y, et al. Association between polymorphisms in pre-miRNA genes and risk of lung cancer in a Chinese non-smoking female population. Lung Cancer. 2016;94:15–21.
Wu S, Yuan W, Shen Y, Lu X, Li Y, Tian T, et al. The miR-608rs4919510 polymorphism may modify cancer susceptibility based on type. Tumour Biol. 2017;39:1010428317703819.
Pardini B, Rosa F, Naccarati A, Vymetalkova V, Ye Y, Wu X, et al. Polymorphisms in microRNA genes as predictors of clinical outcomes in colorectal cancer patients. Carcinogenesis. 2015;36:82–6.
Ghanbari M, Erkeland SJ, Xu L, Colijn JM, Franco OH, Dehghan A, et al. Genetic variants in microRNAs and their binding sites within gene 3′UTRs associate with susceptibility to age-related macular degeneration. Hum Mutat. 2017;38:827–38.
Ghanbari M, de Vries PS, de Looper H, Peters MJ, Schurmann C, Yaghootkar H, et al. A genetic variant in the seed region of miR-4513 shows pleiotropic effects on lipid and glucose homeostasis, blood pressure, and coronary artery disease. Hum Mutat. 2014;35:1524–31.
Othman N, Nagoor NH. miR-608 regulates apoptosis in human lung adenocarcinoma via regulation of AKT2. Int J Oncol. 2017;51:1757–64.
Othman N, In LL, Harikrishna JA, Hasima N. Bcl-xL silencing induces alterations in hsa-miR-608 expression and subsequent cell death in A549 and SK-LU1 human lung adenocarcinoma cells. PLoS One. 2013;8:e81735.
Liang Z, Wang X, Xu X, Xie B, Ji A, Meng S, et al. MicroRNA-608 inhibits proliferation of bladder cancer via AKT/FOXO3a signaling pathway. Mol Cancer. 2017;16:96.
Zhang Y, Schiff D, Park D, Abounader R. MicroRNA-608 and microRNA-34a regulate chordoma malignancy by targeting EGFR, Bcl-xL and MET. PLoS One. 2014;9:e91546.
Acknowledgements
This study was financially supported by the National Natural Science Foundation of China (31671300 and 31871306); Taishan Scholars Program of Shandong Province (tsqn20161060).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Zhang, N., Li, Y., Zheng, Y. et al. miR-608 and miR-4513 significantly contribute to the prognosis of lung adenocarcinoma treated with EGFR-TKIs. Lab Invest 99, 568–576 (2019). https://doi.org/10.1038/s41374-018-0164-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41374-018-0164-y
This article is cited by
-
MicroRNAs as the critical regulators of tyrosine kinase inhibitors resistance in lung tumor cells
Cell Communication and Signaling (2022)
-
LncRNAs LCETRL3 and LCETRL4 at chromosome 4q12 diminish EGFR-TKIs efficiency in NSCLC through stabilizing TDP43 and EIF2S1
Signal Transduction and Targeted Therapy (2022)
-
mTOR-Rictor-EGFR axis in oncogenesis and diagnosis of glioblastoma multiforme
Molecular Biology Reports (2021)
-
LINC02418 promotes malignant behaviors in lung adenocarcinoma cells by sponging miR-4677-3p to upregulate KNL1 expression
BMC Pulmonary Medicine (2020)