Abstract
This article provides a brief review of the literature on the interaction between synthetic particles and biomacromolecules. Understanding the nonspecific interactions between biomacromolecules and synthetic particles is of great importance for using materials in vitro and in vivo. The interaction between functionalized polymer particles and proteins has been studied extensively to identify the main factor that governs the interaction in vitro. Recently, the composition and properties of the protein corona that forms on the surface of nanoparticles (NPs) in biofluids have been studied in the context of the function and distribution of the particles in vivo. In the meantime, NPs that recognize specific biomacromolecules have also been designed by tuning the combination and distribution of functional groups on gold NPs and dendrimers. It has also been shown that nano-gel particles that recognize target molecules can be achieved by (A) optimizing the combination and amount of functional groups in the particles and (B) molecular imprinting polymerization, in combination with (C) affinity purification. Some of these particles are capable of recognizing target molecules and neutralizing their function, even in the bloodstream of living animals, as ‘plastic antibodies.’
Similar content being viewed by others
Introduction
For a protein to strongly and specifically bind to a target molecule, a three-dimensional structure that is complementary to the target molecule should be formed on the protein surface. In addition, functional groups that have positive and negative charges and polar, hydrophobic and aromatic groups are precisely arranged on the binding surface that extends to 100 Å2. These functional groups contact the target through electrostatic interactions, hydrogen bonding and hydrophobic interactions, and they achieve strong and specific target recognition (Figure 1). This elaborate three-dimensional structure is achieved by arranging and folding polypeptides consisting of 20 amino acids.
If various functional groups that contact with target molecules through various interaction modes are arranged and fixed on a three-dimensional space on or in synthetic particles, it would possibly be useful as inexpensive and robust substitute for proteins. To date, interactions between various synthetic particles and biomacromolecules such as proteins, peptides and polysaccharides have been investigated. The studies can be broadly classified as (a) fundamental studies of ‘nonspecific’ adsorption of biomacromolecules and (b) the design of particles that recognize specific molecules (Figure 2). In the former case, interaction is usually not very strong. Thus, amount of proteins adsorbed on the surface of the particles is quantified, compared and discussed in the high protein concentration. In the latter case, affinity to the target molecules has to be quantified and strong; usually, dissociation constants are lower than micro molar level. Some of them show strong affinity comparable to antibodies. Various particles have been used in those studies, including solid polystyrene, silica and gold particles that are denser than protein and swollen, low-density dendrimers and gel particles.
Overview of this review. (a) Left: Fundamental studies of protein adsorption on the surface of functionalized polystyrene latexes and pNIPAm microgels in vitro. Right: Fundamental study of protein adsorption on the surface of NPs in vivo. (b) Design of gold NPs, dendritic polymers and nanogel particles that recognize target biomacromolecules.
Several fundamental studies have revealed the factors that control the nonspecific protein adsorption on synthetic particles in vitro. Recently, the composition and properties of the protein corona that forms on the surface of nanoparticles (NPs) in biofluids have been extensively studied in the context of the function and distribution of the particles in vivo. NPs that strongly interact with specific molecules are designed by tuning the combination and distribution of functional groups on synthetic particles such as gold NPs and particles formed from dendritic polymers. Recently, it has been shown that nano-gel particles that recognize the target molecule can be achieved by (A) optimizing the combination and amount of functional groups in the particles and (B) molecular imprinting polymerization, in combination with (C) affinity purification. NPs that can recognize, neutralize and eliminate a target molecule in a bloodstream of living animals have also been reported.
Fundamental study of protein adsorption on the surface of synthetic particles
Solid particles
The binding behavior of proteins on the surface of polystyrene latex beads has been studied since the late 1970s.1, 2, 3, 4, 5, 6, 7 The effects of the surface functional groups of polystyrene microparticles on the adsorption of proteins were extensively studied in the late 1980s. Kawaguchi et al.8 quantitatively functionalized the surface of polystyrene beads with hydroxyl, carboxyl and amino groups, systematically analyzing their interaction with human serum albumin, γ-globulin, cytochrome C, myoglobin and horseradish peroxidase. They reported that the amount of protein binding to the surface generally increases as the hydrophobicity of the particle surface increases. In addition, they also concluded that particles with a higher charge density interacted with a larger amount of protein if the particles are hydrophilic.8
Lyklema and coworkers9, 10 analyzed the effect of solution pH on the interaction of bovine serum albumin and IgG with polystyrene latex, the surface of which had been modified with charges of various densities. They reported that the maximum adsorption occurred at the isoelectric point of the protein-covered polystyrene particles, rather than at the isoelectric point of the protein in solution.
It has been reported that the combination of hydrophobic and electrostatic interactions is important. In 1999, it was reported that protein adsorption mainly occurs by hydrophobic interactions when a hydrophobic surface was only slightly modified with ionic functional groups. In contrast, adsorption primarily occurs by hydrogen bonding or electrostatic interactions when the surface is highly modified with ionic groups (Figure 3).11, 12, 13
Gel particles
From the late 1980s to the early 2000s, procedures to prepare poly-N-isopropylacrylamide (pNIPAm) hydrogel particles having various functional groups were established. First, Pelton and Chibante14 succeeded in synthesizing temperature-responsive pNIPAm microgels with a size of several hundred nanometers to several microns by polymerizing pNIPAm in water at temperatures above the lower critical solution temperature of pNIPAm. They subsequently showed that it is possible to synthesize gel particles having a diameter of several tens of nanometers by polymerizing the particles in the presence of a surfactant.15, 16 Kokufuta and coworkers17, 18 and Debord and Lyon19 established methods to synthesize multifunctional and temperature-responsive nanogel particles by the copolymerization of NIPAm with functional monomers having positively charged, negatively charged and hydrophobic functional groups.
It has been reported that poly(NIPAm)-based hydrogels undergo a volume phase transition from a solvent-swollen ‘hydrophilic’ state to a collapsed ‘hydrophobic’ state at the lower critical solution temperature, which is typically 31 °C (Figure 4). Kawaguchi et al.20 discovered that human γ-globulin is adsorbed by pNIPAm gel particles above the transition temperature (possibly through hydrophobic interactions) but little is adsorbed by the swollen gel below the transition temperature. A change in the amount of protein that binds to gel particles in response to temperature changes has also been observed with the binding of HIV-1 capsid protein21 and IgG22 to cationic pNIPAm.
Smith and Lyon23 reported that when the amount of carboxylic acid introduced into a microgel is increased, it is possible to encapsulate large amounts of cytochrome C into the particles by electrostatic interactions. This interaction can be attenuated by increasing the salt concentration from 20 to 200 mM (Figure 5). Hansson and coworkers24 also reported that it is possible to encapsulate 2.4 g of lysozyme per gram of negatively charged gel particles through electrostatic interactions.
Illustration of salt-responsive encapsulation of positively charged proteins by negatively charged pNIPAm gel particles.23 A full color version of this figure is available at Polymer Journal online.
Shea and coworkers25 analyzed the interaction between an anionic polysaccharide (heparin) and positively charged acrylamide nano-gel particles that had various cationic side chains using isothermal titration calorimetry, and discussed the contribution of entropy and enthalpy to the interaction. They concluded from the isothermal titration calorimetry results that hydrogen bonding, ionic interactions and dehydration all have important roles in the NP−heparin interaction. Interestingly, they find that NPs with low crosslinking may make accessible more of the NP interior to heparin, resulting in a higher capacity.
The adsorption of proteins on gel particles can be reduced by modification of the gel surface with polyethyleneglycol26 and the use of long polyethyleneglycol chains as crosslinking agents.27
Fundamental study of protein adsorption on the surface of synthetic particles in biological fluids
Attempts to use small particles (NPs) having a diameter of ∼100 nm in the bloodstream of animals have been reported since the mid 2000s.28 Unless the particles are specifically designed to avoid passive protein interactions, for example, by polyethyleneglycol coating, NP in biological fluids are rapidly covered by proteins, forming a corona that interacts with biological systems.29 The corona around the particles is expected to have a dominant role in determining the function of the particles. Therefore, detailed analyses have addressed the types and amounts of proteins in the corona that forms on the surfaces of gel particles,29, 30, 31 gold NPs,32 polystyrene NPs,33, 34, 35 silica NPs,33, 36, 37 titania NPs38 and zinc oxide NPs38 in addition to effects on the exchange rate.30
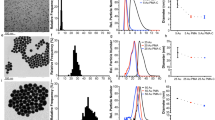
For example, Dawson and coworkers29 developed a methodology to measure the exchange rate and affinity of the protein on the corona in addition to the amount and types of the proteins. They found that gel particles introduced into the blood are first covered by weakly binding serum albumins (soft corona) and subsequently covered by a hard (slow exchange) corona of strongly binding apolipoprotein (Figure 6).30 The relationships between the type of protein corona and the surface functional group composition and NP diameter were also analyzed. The type of proteins that bind to hydrophobic NPs was found to not depend on the diameter (50 or 100 nm) of the particles (∼80% homology), but the particle diameter affected the type of proteins that bound to the surface of either positively or negatively charged NPs.34 In addition, among NPs of the same size, 30% of the proteins bound to the NPs regardless of surface hydrophobicity or charge. Thus, the binding of 30% of the proteins in the corona was completely independent of the surface properties.34
Design of synthetic particles that recognize biomacromolecules by optimizing combination of functional groups
Gold NPs
If combinations of functional groups rather than the 20 amino acids are randomly presented on NPs, the NPs are, to a certain degree, capable of recognizing biomacromolecules. For example, when the surfaces of gold NPs are modified with surfactants having various functional groups, it is possible for the NPs to discriminate proteins. Rotello and coworkers39, 40, 41, 42 found that when the surface of gold NPs is modified with both a carboxylic acid and a hydrophobic group, chymotrypsin can be recognized by electrostatic and hydrophobic interactions, and its enzymatic activity can further be inhibited by the interaction. The affinity for chymotrypsin can be controlled from 106–107 M by tuning the type of hydrophobic group (Figure 7).39, 40, 41, 42 In addition, by merely introducing a slightly different type of functional group on the gold surface, it is possible to distinguish proteins41 or to achieve recognition of a specific domain of a protein.42
Dendritic polymers
Some dendritic polymers recognize target proteins if their terminals are functionalized and their size is optimized. Haag and coworkers43 developed a dendritic polyglycerol (dPG) modified with sulfate groups and analyzed its interaction with selectin (Figure 8). They discovered that it was possible to control the affinity for L-selectin having cationic pockets from approximately millimolar to the nanomolar level by adjusting the dPG size or the sulfate group density. In contrast, as E-selectin does not have cationic pockets, almost no interaction was observed for E-selectin. Thus, it was concluded that the modified dPG interacted with L-selectin through multipoint electrostatic interactions.43 In addition, as the size of the dPG significantly affected the suppression of selectin activity, it was suggested that the strong inhibitory activity of dPG is not only on a strong electrostatic interaction but also on steric shielding by large flexible dPG particles.44 Surprisingly, dPG was also shown to suppress selectin-mediated inflammation in vivo.43 Furthermore, when the anion on the dPG terminal was changed to bisphosphate, sulfonate, phosphonate, phosphate or carboxylate, the bonds to L-selectin gradually weakened (Figure 8).45 These results suggest that for dPG to strongly bind to selectin, simply displaying a large number of anions is not sufficient. Rather, it is necessary to design appropriate molecular structure.
Hydrogel NPs
Free radical polymerization of inexpensive functional acrylic monomers is a simple method to produce NPs having a biomolecular recognition capability.46 For example, a library of pNIPAm nano-gel particles with a size of several tens of nanometers was synthesized from functional acrylic monomers (for example, acrylic acid, 3-N-aminopropylmethacrylamide and N-tert-butylacrylamide) having negatively charged, positively charged and hydrophobic functional groups (Figure 9). Melittin, a 26-amino acid hemolytic peptide consisting of hydrophobic and positively charged amino acids was used as a model target peptide. pNIPAm particles consisting of only negatively charged or hydrophobic functional monomers did not interact with melittin, but NPs with both negatively charged and hydrophobic functional groups interacted with melittin with a 10−6 M dissociation constant (Figure 10), indicating that melittin was recognized by the NPs through multipoint electrostatic and hydrophobic interactions.46, 47 As these nano-gel particles do not strongly interact with plasma proteins such as albumin and fibrinogen, melittin is recognized and its toxicity can be completely neutralized by the particles even under physiological conditions.46, 47 Because of their strong and specific interaction, the C-terminal 12 amino acids of the peptide can further be used as a tag for protein purification.48
The specificity of target recognition can be tuned through selection of functional monomers; NPs with carboxylic acid and sulfonic acid49 or primary amine, tertiary amine and guanidium groups50 showed completely different binding affinities to target proteins (Figure 11). The library of particles can be easily diversified because dozens of functional monomers are commercially available, which makes it theoretically possible to recognize a wide range of biomacromolecules. In fact, NPs that strongly bind to histone,49 fibrinogen51 and lysozyme51 have already been developed. In addition, NPs that bind to a specific domain of the target protein (for example, IgG Fc fragment) can be synthesized.52
Interaction between positively charged groups on NPs and carboxylic acids on proteins.49
Poly(NIPAm) NPs undergo a volume phase transition from a swollen state to a collapsed state in response to temperature and pH changes (Figure 4). Together with large changes in the density of the functional groups in the NPs,51, 53 the mobility of polymer chains54, 55 and the ionization states of the acidic and basic functional groups in the nanogel56, 57, 58, 59, 60, 61 change reversibly. As the functional group density in the nanogel increases 10–100 times with phase transition, it is possible to almost completely switch on and off the interaction with the target protein by applying external stimuli. It should be noted that the binding constant for multipoint binding recognition increases exponentially with the number of contact points.47, 53 Therefore, based on the stimulus response, these particles can be used to isolate target proteins from a protein mixture in solution by precipitating target proteins as complex with nanogels at a high temperature and eluting the targets by cooling.48
The binding and release kinetics of the multipoint target recognition process can be tuned by manipulating the mobility of the polymer chains.54, 55 PNIPAm nanogel particles with a ‘flexible’ random-coil conformation were found to have a faster binding rate than nanogel particles with a ‘rigid’ globule conformation; however, the dissociation rate constant remained unchanged, resulting in stronger affinity. The dissociation rate of the ‘flexible’ nanogel particles was found to be decreased by the ‘induced-fit’ conformation change of polymers around the coil-globule phase-transition temperature, resulting in the formation of highly stable particle–protein complexes.54 In addition, both the binding and dissociation rate constants of the swollen phase nanogel particles increased as the crosslinking degree decreased, indicating that the polymer density of the particles governs the binding and dissociation rates.55 These results also suggest that the mechanical flexibility of gel particles, defined as the Young’s modulus, does not always have a crucial role in the multipoint molecular recognition process.55 Tuning of the binding/dissociation rate will be important for the future use of the particles in non-equilibrium systems.
The nanogel particles have a cost advantage over other materials because they can be synthesized on a large scale using inexpensive monomers. However, in the free radical polymerization process described above, the arrangement of the functional monomers and the three-dimensional structure of the polymer chains are not controlled during synthesis. Thus, the structure of the binding site is completely random, resulting in limited specificity and binding strength.
Preparation of synthetic particles that recognize biomacromolecules by molecular imprinting polymerization
To achieve stronger and more specific target recognition, it is necessary to prepare materials presenting a three-dimensional structure that is complementary to the target molecules. Therefore, a molecular imprinting polymerization method has been proposed. In the molecular imprinting procedure, the three-dimensional shape of the target molecule is transferred into the polymer network by polymerizing the materials in the presence of target molecules, leaving complementary binding sites in the network after the removal of the target (Figure 12). This method was proposed by Mosbach, Wulff, Shea and others in the 1980s.62, 63, 64, 65, 66, 67, 68 Recently, molecular imprint materials that target biomacromolecules have been actively studied by Shea’s group.69
Schematic representation of the peptide imprinting process.70 A full color version of this figure is available at Polymer Journal online.
It is possible to prepare nanogel particles that recognize targets with a dissociation constant comparable to that of antibodies (<nM) by polymerizing the nanogel particles in the presence of the target peptide, using a combination of functional monomers that are capable of contacting the target through electrostatic and hydrophobic interactions.70 As this NP does not strongly interact with plasma proteins such as albumin or fibrinogen, it is stable even in the plasma.
Nanogel particles are also being tested as antitoxins in animal experiments. It has been reported that when mice were intravenously administered with a lethal dose of melittin, a 100% mortality rate was observed. However, on intravenous infusion of melittin-imprinted nanogel particles, a significant decrease in mortality was observed. In contrast, particles synthesized with the same monomers in the absence of melittin did not significantly neutralize melittin in vivo, indicating that a strong and specific interaction of the imprinted NPs enabled the neutralization of target in vivo.71 Furthermore, biodistribution studies of nanogel particles and melittin using fluorescent or radioisotope-labeled materials revealed that the particles bind to target peptides in the bloodstream.71
Although peptide-imprinted particles can strongly and specifically recognize targets as ‘plastic antibodies,’ the method of molecular imprinting requires a large amount of expensive template biomacromolecules for NP synthesis. Thus, the use of imprinted particles negates the cost advantage of polymer materials. Therefore, affinity purification has recently been developed for isolation of particles that strongly recognize biomacromolecules.
Isolation of synthetic NPs that recognize biomacromolecules by affinity purification
Although the binding capacity of randomly polymerized particles with an optimized combination of functional monomers is high, the average binding affinity of the particles to melittin is much lower than that of natural antibodies because the statistically created affinity sites have a broad distribution of affinities for the target. However, the distribution of affinities suggests that high-affinity nanogel particles are present in the mixture. To isolate NPs with a high affinity and a narrow affinity distribution, the NPs were sorted on the basis of their peptide affinity similar to that in proteins by performing affinity chromatography (Figure 13). When melittin, the target molecule, was fixed onto agarose beads and melittin-binding NPs prepared using random copolymerization were affinity purified, it was possible to isolate NPs with a nanomolar level dissociation constant from the NP solution with an overall submicromolar dissociation constant.72 The purified particles showed a much narrower affinity distribution than the average value obtained for the particles before purification. The yield of high-affinity NPs isolated using this process was only several percent, but even when taking into account the cost of the commercial monomers the preparation of the purified particles is much less expensive than that of the protein antibodies. Affinity purification is thus a useful process for isolating nano-size materials, including inorganic particles, with high and/or tailor-made affinity and a narrow affinity distribution for target biomacromolecules.
Protocol for affinity purification. Randomly co-polymerized NPs are incubated with target-immobilized beads and then washed to remove the weakly binding NPs. Strongly binding NPs are eluted by changing the buffer conditions and/or temperature.70 A full color version of this figure is available at Polymer Journal online.
Prospective
In this article, the literature on the interaction of synthetic particles and biomacromolecules was reviewed. On the basis of the fundamental study of ‘nonspecific’ protein–particle interactions, strategies for the design of NPs that strongly interact with specific molecules have been developed. Further fusion of the approaches of supramolecular chemistry,73 stereocontrolled polymerization,74 topology control75 and living polymerization76 is expected to enable future development of uniform polymer ligands77, 78 as ‘monoclonal plastic antibodies,79 that more strongly and specifically recognize target biomacromolecules.
References
Norde, W. & Lyklema, J. The adsorption of human plasma albumin and bovine pancreas ribonuclease at negatively charged polystyrene surfaces: I. Adsorption isotherms. Effects of charge, ionic strength, and temperature. J. Colloid Interface Sci. 66, 257–265 (1978).
Suzawa, T., Shirahama, H. & Fujimoto, T. Adsorption of bovine serum albumin onto homo- and copolymer lattices. J. Colloid Interface Sci. 86, 144–150 (1982).
Fair, B. D. & Jamieson, A. M. Studies of protein adsorption on polystyrene latex surfaces. J. Colloid Interface Sci. 77, 525–534 (1980).
Bagchi, P. & Birnbaum, S. M. Effect of pH on the adsorption of immunoglobulin G on anionic poly(vinyltoluene) model latex particles. J. Colloid Interface Sci. 83, 460–478 (1981).
de Bruin, H. G., van Oss, C. J. & Absolom, D. R. Desorption of protein from polystyrene latex particles. J. Colloid Interface Sci. 76, 254–255 (1980).
Van Dulm, P., Norde, W. & Lyklema, J. Ion participation in protein adsorption at solid surfaces. J. Colloid Interface Sci. 82, 77–82 (1981).
Koutsoukos, P. G., Mumme-Young, C. A., Norde, W. & Lyklema, J. Effect of the nature of the substrate on the adsorption of human plasma albumin. J. Colloids Surf. 5, 93–104 (1982).
Kawaguchi, H., Amagasa, H., Hagiya, T., Kimura, N. & Ohtsuka, Y. Interaction between proteins and latex particles having different surface structures. Colloids Surf. 13, 295–311 (1985).
Elgersma, A. V., Zsom, R. L. J., Norde, W. & Lyklema, J. The adsorption of bovine serum albumin on positively and negatively charged polystyrene lattices. J. Colloid. Interf. Sci. 138, 145–156 (1990).
Elgersma, A. V., Zsom, R. L. J., Norde, W. & Lyklema. The asorption of different types of monoclonal immunoglobulin on positively and negatively charged polystyrene lattices. J. Colloid Surf. 54, 89–101 (1991).
Kondo, A. & Higashitani, K. Adsorption of model proteins with wide variation in molecular properties on colloidal particles. J. Colloid Interface Sci. 150, 344–351 (1992).
Ortega-Vinuesa, J. L. & Hidalgo-Álvarez, R. Sequential adsorption of F(ab′)2 and BSA on negatively and positively charged polystyrene latexes. Biotechnol. Bioeng. 47, 633–639 (1995).
Yoon, J.-Y., Kim, W. S. & Kim, J.-H. The relationship of interaction forces in the protein adsorption onto polymeric microspheres. Colloids Surf. A 153, 413–419 (1999).
Pelton, R. H. & Chibante, P. Preparation of aqueous latices with N-isopropylacrylamide. Colloids Surf. 20, 247–256 (1986).
McPhee, W., Tam, K. C. & Pelton, R. Poly(N-isopropylacrylamide) latices prepared with sodium dodecyl sulfate. J. Colloid Interface Sci. 156, 24–30 (1993).
Pelton, R. Temperature-sensitive aqueous microgels. Adv. Colloid Interface Sci. 85, 1–33 (2000).
Ito, S., Ogawa, K., Suzuki, H., Wang, B. L., Yoshida, R. & Kokufuta, E. Preparation of thermosensitive submicrometer gel particles with anionic and cationic charges. Langmuir 15, 4289–4294 (1999).
Ogawa, K., Nakayama, A. & Kokufuta, E. Preparation and characterization of thermosensitive polyampholyte nanogels. Langmuir 19, 3178–3184 (2003).
Debord, J. D. & Lyon, L. A. Synthesis and characterization of pH-responsive copolymer microgels with tunable volume phase transition temperatures. Langmuir 19, 7662–7664 (2003).
Kawaguchi, H., Fujimoto, K. & Mizuhara, Y. Hydrogel microspheres III. Temperature-dependent adsorption of proteins on poly-N-isopropylacrylamide hydrogel microspheres. Colloid Polym. Sci. 270, 53–57 (1992).
Duracher, D., Elaissari, A., Mallet, F. & Pichot, C. Adsorption of modified HIV-1 capsid p24 protein onto thermosensitive and cationic core−shell poly(styrene)−poly(N-isopropylacrylamide) particles. Langmuir 16, 9002–9008 (2000).
Silva, C. S. O., Baptista, R. P., Santos, A. M., Martinho, J. M. G., Cabral, J. M. S. & Taipa, M. A. Adsorption of human IgG on to poly(N-isopropylacrylamide)-based polymer particles. Biotechnol. Lett. 28, 2019–2025 (2006).
Smith, D. & Lyon, L. A. Tunable encapsulation of proteins within charged microgels. Macromolecules 44, 8154–8160 (2011).
Johansson, C., Gernandt, J., Bradley, M., Vincent, B. & Hansson, P. Interaction between lysozyme and colloidal poly(NIPAM-co-acrylic acid) microgels. J. Colloid Interface Sci. 347, 241–251 (2010).
Zeng, Z., Patel, J., Lee, S.-H., McCallum, M., Tyagi, A., Yan, M. & Shea, K. J. Synthetic polymer nanoparticle–polysaccharide interactions: a systematic study. J. Am. Chem. Soc. 134, 2681–2690 (2012).
Gan, D. & Lyon, L. A. Synthesis and protein adsorption resistance of PEG-modified poly(N-isopropylacrylamide) core/shell microgels. Macromolecules 35, 9634–9639 (2002).
Nolan, C. M., Reyes, C. D., Debord, J. D., Garcia, A. J. & Lyon, L. A. phase transition behavior, protein adsorption, and cell adhesion resistance of poly(ethylene glycol) cross-linked microgel particles. Biomacromolecules 6, 2032–2039 (2005).
Haag, R. & Kratz, F. Polymer therapeutics: concepts and applications. Angew. Chem. Int. Ed. 45, 1198–1215 (2006).
Cedervall, T., Lynch, I., Lindman, S., Berggard, T., Thulin, E., Nilsson, H., Dawson, K. A. & Linse, S. Understanding the nanoparticle–protein corona using methods to quantify exchange rates and affinities of proteins for nanoparticles. Proc. Natl Acad. Sci. USA 104, 2050–2055 (2007).
Cedervall, T., Lynch, I., Foy, M., Berggård, T., Donnelly, S. C., Cagney, G., Linse, S. & Dawson, K. A. Detailed identification of plasma proteins adsorbed on copolymer nanoparticles. Angew. Chem. Int. Ed. 46, 5754–5756 (2007).
Lindman, S., Lynch, I., Thulin, E., Nilsson, H., Dawson, K. A. & Linse, S. Systematic investigation of the thermodynamics of HSA adsorption to N-iso-propylacrylamide/N-tert-butylacrylamide copolymer nanoparticles. Effects of particle size and hydrophobicity. Nano Lett. 7, 914–920 (2007).
Casals, E., Pfaller, T., Duschl, A., Oostingh, G. J. & Puntes, V. Time evolution of the nanoparticle protein corona. ACS Nano 4, 3623–3632 (2010).
Walczyk, D., Baldelli Bombelli, F., Monopoli, M. P., Lynch, I. & Dawson, K. A. What the cell “sees” in bionanoscience. J. Am. Chem. Soc. 132, 5761–5768 (2010).
Lundqvist, M., Stigler, J., Cedervall, T., Elia, G., Lynch, I. & Dawson., K. Nanoparticle size and surface properties determine the protein corona with possible implications for biological impacts. Proc. Natl Acad. Sci. USA 105, 14265–14270 (2008).
Monopoli, M., Aberg, C., Salvati, A. & Dawson, K. A. Biomolecular coronas provide the biological identity of nanosized materials. Nat. Nanotechnol. 7, 779–786 (2012).
Tenzer, S., Docter, D., Rosfa, S., Wlodarski, A., Kuharev, J., Rekik, A., Knauer, S. K., Bantz, C., Nawroth, T., Bier, C., Sirirattanapan, J., Mann, W., Treuel, L., Zellner, R., Maskos, M., Schild, H. & Stauber, R. H. nanoparticle size is a critical physicochemical determinant of the human blood plasma corona: a comprehensive quantitative proteomic analysis. ACS Nano 5, 7155–7167 (2011).
Monopoli, M. P., Walczyk, D., Campbell, A., Elia, G., Lynch, I., Bombelli, F. B. & Dawson, K. A. Physical−chemical aspects of protein corona: relevance to in vitro and in vivo biological impacts of nanoparticles. J. Am. Chem. Soc. 133, 2525–2534 (2011).
Deng, Z. J., Mortimer, G., Schiller, T., Musumeci, A., Martin, D. & Minchin, R. F. Differential plasma protein binding to metal oxide nanoparticles. Nanotechnology 20, 455101 (2009).
Fischer, N. O., Verma, A., Goodman, C. M., Simard, J. M. & Rotello, V. M. Reversible “irreversible” inhibition of chymotrypsin using nanoparticle receptors. J. Am. Chem. Soc. 125, 13387–13391 (2003).
You, C.-C., De, M., Han, G. & Rotello, V. M. Tunable inhibition and denaturation of α-chymotrypsin with amino acid-functionalized gold nanoparticles. J. Am. Chem. Soc. 127, 12873–12881 (2005).
De, M., You, C.-C., Srivastava, S. & Rotello, V. M. Biomimetic interactions of proteins with functionalized nanoparticles: a thermodynamic study. J. Am. Chem. Soc. 129, 10747–10753 (2007).
Bayraktar, H., You, C.-C., Rotello, V. M. & Knapp, M. J. Facial control of nanoparticle binding to cytochrome c. J. Am. Chem. Soc. 129, 2732–2733 (2007).
Dernedde, J., Rausch, A., Weinhart, M., Enders, S., Tauber, R., Licha, K., Schirner, M., Zügel, U., von Bonin, A. & Haag, R. Dendritic polyglycerol sulfates as multivalent inhibitors of inflammation. Proc. Natl Acad. Sci. USA 107, 19679–19684 (2010).
Weinhart, M., Groger, D., Enders, S., Riese, S. B., Dernedde, J., Kainthan, R. K., Brooks, D. E. & Haag, R. The role of dimension in multivalent binding events: structure–activity relationship of dendritic polyglycerol sulfate binding to L-selectin in correlation with size and surface charge density. Macromol. Biosci. 11, 1088–1098 (2011).
Weinhart, M., Groger, D., Enders, S., Dernedde, J. & Haag, R. Synthesis of dendritic polyglycerol anions and their efficiency toward l-selectin inhibition. Biomacronolecules 12, 2502–2511 (2011).
Hoshino, Y., Urakami, T., Kodama, T., Koide, H., Oku, N., Okahata, Y. & Shea, K. J. Design of synthetic polymer nanoparticles that capture and neutralize a toxic peptide. Small 5, 1562–1568 (2009).
Hoshino, Y., Koide, H., Furuya, K., Haberaecker, W. W. III, Lee, S.-H., Kodama, T., Kanazawa, H., Oku, N. & Shea, K. J. The rational design of a synthetic polymer nanoparticle that neutralizes a toxic peptide in vivo. Proc. Natl Acad. Sci. USA 109, 33–38 (2012).
Yoshimatsu, K., Yamazaki, T., Hoshino, Y., Rose, P. E., Epstein, L. F., Miranda, L. P., Tagari, P., Beierle, J. M., Yonamine, Y. & Shea, K. J. Epitope discovery for a synthetic polymer nanoparticle: a new strategy for developing a peptide tag. J. Am. Chem. Soc. 136, 1194–1197 (2014).
Yonamine, Y, Hoshino, Y. & Shea, K. J. An ELISA-mimic screen for synthetic polymer nanoparticles with high affinity to target proteins. Biomacromolecules 13, 2952–2957 (2012).
Yonamine, Y., Yoshimatsu, K., Lee, S.-H., Hoshino, Y., Okahata, Y. & Shea, K. J. Polymer nanoparticle–protein interface. evaluation of the contribution of positively charged functional groups to protein affinity. ACS Appl. Mater. Interfaces 5, 374–379 (2013).
Yoshimatsu, K., Lesel, B. K., Yonamine, Y., Beierle, J. M., Hoshino, Y. & Shea, K. J. Temperature-responsive “catch and release” of proteins by using multifunctional polymer-based nanoparticles. Angew. Chem. Int. Ed. 51, 2405–2408 (2012).
Lee, S.-H., Hoshino, Y., Randall, A., Zeng, Z., Baldi, P., Doong, R. & Shea, K. J. engineered synthetic polymer nanoparticles as IgG affinity ligands. J. Am. Chem. Soc. 134, 15765–15772 (2012).
Oya, T., Enoki, T., Grosberg, A. Y., Masamune, S., Sakiyama, T., Takeoka, Y., Tanaka, K., Wang, G., Yilmaz, Y., Feld, M. S., Dasari, R. & Tanaka, T. reversible molecular adsorption based on multiple-point interaction by shrinkable gels. Science 286, 1543–1545 (1999).
Hoshino, Y., Nakamoto, M. & Miura, Y. Control of protein-binding kinetics on synthetic polymer nanoparticles by tuning flexibility and inducing conformation changes of polymer chains. J. Am. Chem. Soc. 134, 15209–15212 (2012).
Nakamoto, M., Hoshino, Y. & Miura, Y. Effect of physical properties of nanogel particles on the kinetic constants of multipoint protein recognition process. Biomacromolecules 15, 541–547 (2014).
Hoshino, Y., Imamura, K., Yue, M., Inoue, G. & Miura, Y. Reversible absorption of CO2 triggered by phase transition of amine-containing micro- and nanogel particles. J. Am. Chem. Soc. 134, 18177–18180 (2012).
Yue, M., Hoshino, Y., Ohshiro, Y., Imamura, K. & Miura, Y. Temperature-responsive microgel films as reversible carbon dioxide absorbents in wet environment. Angew. Chem. Int. Ed. 53, 2654–2657 (2014).
Hoshino, Y., Ohashi, R. C. & Miura, Y. Rational design of synthetic nanoparticles with a large reversible shift of acid dissociation constants: proton imprinting in stimuli responsive nanogel particles. Adv. Mater. 26 doi:10.1002/adma.201305957 (2014).
Nagase, K., Kobayashi, J., Kikuchi, A., Akiyama, Y., Kanazawa, H. & Okano, T. Preparation of thermoresponsive cationic copolymer brush surfaces and application of the surface to separation of biomolecules. Biomacromolecules 9, 1340–1347 (2008).
Feil, H., Bae, Y. H., Feijen, J. & Kim, S. W. Mutual influence of pH and temperature on the swelling of ionizable and thermosensitive hydrogels. Macromolecules 25, 5528–5530 (1992).
Yang, Y., Mijalis, A. J., Fu, H., Agosto, C., Tan, K., Batteas, J. D. & Bergbreiter, D. E. Reversible changes of solution pH resulting from changes in thermoresponsive polymer solubility. J. Am. Chem. Soc. 134, 7378–7383 (2012).
Wulff, G. Molecular imprinting in cross-linked materials with the aid of molecular templates— a way towards artificial antibodies. Angew. Chem. Int. Ed. Engl. 34, 1812–1832 (1995).
Vlatakis, G., Andersson, L. I., Müller, R. & Mosbach, K. Drug assay using antibody mimics made by molecular imprinting. Nature 361, 645–647 (1993).
Yan, M. & Ramstrom, O. (Eds) Molecularly Imprinted Materials: Science and Technology, (CRC Press, Boca Raton, FL, USA, 2004).
Sellergren, B. Molecularly Imprinted Polymers, (Elsevier, Amsterdam, 2001).
Shea, K. Molecular imprinting of synthetic network polymers: the de novo synthesis of macromolecular binding and catalytic sites. Trends Polym. Sci. 2, 166–173 (1994).
Haupt, K. Molecularly imprinted polymers: the next generation. Anal. Chem. 75, 376A–383A (2003).
Cakir, P., Cutivet, A., Resmini, M., Bui, B. T. & Haupt, K. Protein-size molecularly imprinted polymer nanogels as synthetic antibodies, by localized polymerization with multi-initiators. Adv. Mater. 25, 1048–1051 (2013).
Zeng, Z., Hoshino, Y., Rodriguez, A., Yoo, H. & Shea, K. J. Synthetic polymer nanoparticles with antibody-like affinity for a hydrophilic peptide. ACS Nano 4, 199–204 (2010).
Hoshino, Y., Kodama, T., Okahata, Y. & Shea, K. J. Peptide imprinted polymer nanoparticles: a plastic antibody. J. Am. Chem. Soc. 130, 15242–15243 (2008).
Hoshino, Y., Koide, H., Urakami, T., Kanazawa, H., Kodama, T., Oku, N. & Shea, K. J. Recognition, neutralization and clearance of target peptides in the blood stream of living mice by molecular imprinted polymer nanoparticles: a plastic antibody. J. Am. Chem. Soc. 132, 6644–6645 (2010).
Hoshino, Y., Haberaecker, W. W. III, Kodama, T., Zeng, Z., Okahata, Y. & Shea, K. J. affinity purification of multi functional polymer nanoparticles. J. Am. Chem. Soc. 132, 13648–13650 (2010).
Hirao, A. & Yoo, H.-S. Dendrimer-like star-branched polymers: novel structurally well-defined hyperbranched polymers. Polym. J. 43, 2–17 (2011).
Takeuchi, D. Stereo-controlled synthesis of polyolefins with cycloalkane groups by using late transition metals. Polym. J. 44, 919–928 (2012).
Tezuka, Y. Topological polymer chemistry for designing multicyclic macromolecular architectures. Polym. J. 44, 1159–1169 (2012).
Kamigaito, M. recent developments in metal-catalyzed living radical polymerization. Polym. J. 43, 105–120 (2011).
Hatada, K., Kitayama, T., Ute, K. & Nishiura, T. Synthetic uniform polymers and their use for understanding fundamental problems in polymer chemistry. Macromol. Rapid Commun. 25, 1447–1477 (2004).
Hatada, K. Stereoregular uniform polymers. J. Polym. Sci. Part A Polym. Chem. 37, 245–260 (1999).
Hoshino, Y. & Shea, K. J. The evolution of plastic antibodies. J. Mater. Chem. 21, 3517–3521 (2011).
Acknowledgements
The financial support from MEXT (23111716 and 25107726), JSPS (23750193), Ogasawara Foundation and the Kao Foundation for Arts and Sciences is greatly appreciated.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Hoshino, Y., Lee, H. & Miura, Y. Interaction between synthetic particles and biomacromolecules: fundamental study of nonspecific interaction and design of nanoparticles that recognize target molecules. Polym J 46, 537–545 (2014). https://doi.org/10.1038/pj.2014.33
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/pj.2014.33
Keywords
This article is cited by
-
Quantifying intracellular trafficking of silica-coated magnetic nanoparticles in live single cells by site-specific direct stochastic optical reconstruction microscopy
Journal of Nanobiotechnology (2021)
-
Hollow multishell structures exercise temporal–spatial ordering and dynamic smart behaviour
Nature Reviews Chemistry (2020)
-
Functionalized Nano-adsorbent for Affinity Separation of Proteins
Nanoscale Research Letters (2018)
-
Preparation of abiotic polymer nanoparticles for sequestration and neutralization of a target peptide toxin
Nature Protocols (2015)
-
Preparation of nanogel-immobilized porous gel beads for affinity separation of proteins: fusion of nano and micro gel materials
Polymer Journal (2015)