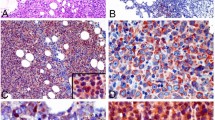
Abstract
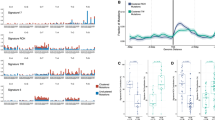
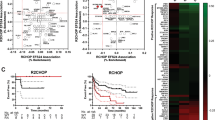
Diffuse large B-cell lymphoma (DLBCL) has been categorized into two molecular subtypes that have prognostic significance, namely germinal center B-cell like (GCB) and activated B-cell like (ABC). Although ABC-DLBCL has been associated with NF-κB activation, the relationships between activation of specific NF-κB signals and DLBCL phenotype remain unclear. Application of novel gene expression classifiers identified two new DLBCL categories characterized by selective p100 (NF-κB2) and p105 (NF-κB1) signaling. Interestingly, our molecular studies showed that p105 signaling is predominantly associated with GCB subtype and histone mutations. Conversely, most tumors with p100 signaling displayed ABC phenotype and harbored ABC-associated mutations in genes such as MYD88 and PIM1. In vitro, MYD88 L265P mutation promoted p100 signaling through TAK1/IKKα and GSK3/Fbxw7a pathways, suggesting a novel role for this protein as an upstream regulator of p100. p100 signaling was engaged during activation of normal B cells, suggesting p100’s role in ABC phenotype development. Additionally, silencing p100 in ABC-DLBCL cells resulted in a GCB-like phenotype, with suppression of Blimp, IRF4 and XBP1 and upregulation of BCL6, whereas introduction of p52 or p100 into GC cells resulted in differentiation toward an ABC-like phenotype. Together, these findings identify specific roles for p100 and p105 signaling in defining DLBCL molecular subtypes and posit MYD88/p100 signaling as a regulator for B-cell activation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Teras LR, DeSantis CE, Cerhan JR, Morton LM, Jemal A, Flowers CR . 2016 US lymphoid malignancy statistics by World Health Organization subtypes. CA Cancer J Clin 2016; 66: 443–459.
Alizadeh AA, Eisen MB, Davis RE, Ma C, Lossos IS, Rosenwald A et al. Distinct types of diffuse large B-cell lymphoma identified by gene expression profiling. Nature 2000; 403: 503–511.
Rosenwald A, Wright G, Chan WC, Connors JM, Campo E, Fisher RI et al. The use of molecular profiling to predict survival after chemotherapy for diffuse large-B-cell lymphoma. N Engl J Med 2002; 346: 1937–1947.
Shipp MA, Ross KN, Tamayo P, Weng AP, Kutok JL, Aguiar RCT et al. Diffuse large B-cell lymphoma outcome prediction by gene-expression profiling and supervised machine learning. Nat Med 2002; 8: 68–74.
Compagno M, Lim WK, Grunn A, Nandula SV, Brahmachary M, Shen Q et al. Mutations of multiple genes cause deregulation of NF-[kgr]B in diffuse large B-cell lymphoma. Nature 2009; 459: 717–721.
Zhang B, Calado Dinis P, Wang Z, Fröhler S, Köchert K, Qian Y et al. An oncogenic role for alternative NF-κB signaling in DLBCL revealed upon deregulated BCL6 expression. Cell Rep 2015; 11: 715–726.
Gasparini C, Celeghini C, Monasta L, Zauli G . NF-κB pathways in hematological malignancies. Cell Mol Life Sci 2014; 71: 2083–2102.
Luo J-L, Kamata H, Karin M . IKK/NF-κB signaling: balancing life and death – a new approach to cancer therapy. J Clin Invest 2005; 115: 2625–2632.
Oeckinghaus A, Hayden MS, Ghosh S . Crosstalk in NF-κB signaling pathways. Nat Immunol 2011; 12: 695–708.
Ramachandiran S, Adon A, Guo X, Wang Y, Wang H, Chen Z et al. Chromosome instability in diffuse large B cell lymphomas is suppressed by activation of the noncanonical NF-κB pathway. Int J Cancer 2014; 136: 2341–2351.
Sha WC, Liou H-C, Tuomanen EI, Baltimore D . Targeted disruption of the p50 subunit of NF-κB leads to multifocal defects in immune responses. Cell 1995; 80: 321–330.
Bernal-Mizrachi L, Lovly CM, Ratner L . The role of NF-κB-1 and NF-κB-2-mediated resistance to apoptosis in lymphomas. PNAS 2006; 103: 9220–9225.
Lam LT, Wright G, Davis RE, Lenz G, Farinha P, Dang L et al. Cooperative signaling through the signal transducer and activator of transcription 3 and nuclear factor-κB pathways in subtypes of diffuse large B-cell lymphoma. Blood 2008; 111: 3701–3713.
Thompson MP, Aggarwal BB, Shishodia S, Estrov Z, Kurzrock R . Autocrine lymphotoxin production in Epstein-Barr virus-immortalized B cells: induction via NF-κB activation mediated by EBV-derived latent membrane protein 1. Leukemia 2003; 17: 2196–2201.
Hummel M, Bentink S, Berger H, Klapper W, Wessendorf S, Barth TFE et al. A biologic definition of Burkitt's lymphoma from transcriptional and genomic profiling. N Engl J Med 2006; 354: 2419–2430.
McCool KW, Miyamoto S . DNA damage-dependent NF-κB activation: NEMO turns nuclear signaling inside out. Immunol Rev 2012; 246: 311–326.
Jazirehi AR, Huerta-Yepez S, Cheng G, Bonavida B . Rituximab (chimeric anti-CD20 monoclonal antibody) inhibits the constitutive nuclear factor-κB signaling pathway in non-hodgkin's lymphoma B-cell lines: role in sensitization to chemotherapeutic drug-induced apoptosis. Cancer Res 2005; 65: 264–276.
Davis RE, Brown KD, Siebenlist U, Staudt LM . Constitutive nuclear factor κB activity is required for survival of activated B cell-like diffuse large B cell lymphoma cells. J Exp Med 2001; 194: 1861–1874.
Lenz G, Wright G, Dave SS, Xiao W, Powell J, Zhao H et al. Stromal gene signatures in large-B-cell lymphomas. N Engl J Med 2008; 359: 2313–2323.
Zhang J, Grubor V, Love CL, Banerjee A, Richards KL, Mieczkowski PA et al. Genetic heterogeneity of diffuse large B-cell lymphoma. Proc Natl Acad Sci 2013; 110: 1398–1403.
Ngo VN, Young RM, Schmitz R, Jhavar S, Xiao W, Lim K-H et al. Oncogenically active MYD88 mutations in human lymphoma. Nature 2011; 470: 115–119.
Pasqualucci L, Neumeister P, Goossens T, Nanjangud G, Chaganti RSK, Kuppers R et al. Hypermutation of multiple proto-oncogenes in B-cell diffuse large-cell lymphomas. Nature 2001; 412: 341–346.
Lohr JG, Stojanov P, Lawrence MS, Auclair D, Chapuy B, Sougnez C et al. Discovery and prioritization of somatic mutations in diffuse large B-cell lymphoma (DLBCL) by whole-exome sequencing. Proc Natl Acad Sci 2012; 109: 3879–3884.
Morin RD, Johnson NA, Severson TM, Mungall AJ, An J, Goya R et al. Somatic mutations altering EZH2 (Tyr641) in follicular and diffuse large B-cell lymphomas of germinal-center origin. Nat Genet 2010; 42: 181–185.
Lenz G, Staudt LM . Aggressive lymphomas. N Engl J Med 2010; 362: 1417–1429.
Shaffer AL, Young RM, Staudt LM . Pathogenesis of human B cell lymphomas. Annu Rev Immunol 2012; 30: 565–610.
Shaffer 3rd AL, Young RM, Staudt LM . Pathogenesis of human B cell lymphomas. Annu Rev Immunol 2012; 30: 565–610.
Klein U, Dalla-Favera R . Germinal centres: role in B-cell physiology and malignancy. Nat Rev Immunol 2008; 8: 22–33.
Bowles JA, Wang S-Y, Link BK, Allan B, Beuerlein G, Campbell M-A et al. Anti-CD20 monoclonal antibody with enhanced affinity for CD16 activates NK cells at lower concentrations and more effectively than rituximab. Blood 2006; 108: 2648–2654.
Lepin EJM, Bastin JM, Allan DSJ, Roncador G, Braud VM, Mason DY et al. Functional characterization of HLA-F and binding of HLA-F tetramers to ILT2 and ILT4 receptors. Eur J Immunol 2000; 30: 3552–3561.
Navarro F, Llano M, Bellón T, Colonna M, Geraghty DE, López-Botet M . The ILT2(LIR1) and CD94/NKG2A NK cell receptors respectively recognize HLA-G1 and HLA-E molecules co-expressed on target cells. Eur J Immunol 1999; 29: 277–283.
Chang H, Schimmer AD . Livin/melanoma inhibitor of apoptosis protein as a potential therapeutic target for the treatment of malignancy. Mol Cancer Ther 2007; 6: 24–30.
Eckfeld K, Hesson L, Vos MD, Bieche I, Latif F, Clark GJ . RASSF4/AD037 is a potential Ras effector/tumor suppressor of the RASSF family. Cancer Res 2004; 64: 8688–8693.
Wu RP, Hayashi T, Cottam HB, Jin G, Yao S, Wu CCN et al. Nrf2 responses and the therapeutic selectivity of electrophilic compounds in chronic lymphocytic leukemia. Proc Natl Acad Sci 2010; 107: 7479–7484.
Asanuma K, Yanagida-Asanuma E, Faul C, Tomino Y, Kim K, Mundel P . Synaptopodin orchestrates actin organization and cell motility via regulation of RhoA signalling. Nat Cell Biol 2006; 8: 485–491.
Hale JS, Frock RL, Mamman SA, Fink PJ, Kennedy BK . Cell-extrinsic defective lymphocyte development in Lmna-/- mice. PLoS ONE 2010; 5: e10127.
Liao Y-C, Si L, de Vere White RW, Lo SH . The phosphotyrosine-independent interaction of DLC-1 and the SH2 domain of cten regulates focal adhesion localization and growth suppression activity of DLC-1. J Cell Biol 2007; 176: 43–49.
Roodink I, Kats G, van Kempen L, Grunberg M, Maass C, Verrijp K et al. Semaphorin 3E expression correlates inversely with plexin D1 during tumor progression. Am J Pathol 2008; 173: 1873–1881.
Chan YW, Fava LL, Uldschmid A, Schmitz MHA, Gerlich DW, Nigg EA et al. Mitotic control of kinetochore-associated dynein and spindle orientation by human spindly. J Cell Biol 2009; 185: 859–874.
De Antoni A, Pearson CG, Cimini D, Canman JC, Sala V, Nezi L et al. The Mad1/Mad2 complex as a template for Mad2 activation in the spindle assembly checkpoint. Curr Biol 2005; 15: 214–225.
Loïodice I, Alves A, Rabut G, van Overbeek M, Ellenberg J, Sibarita J-B et al. The entire Nup107-160 complex, including three new members, is targeted as one entity to kinetochores in mitosis. Mol Biol Cell 2004; 15: 3333–3344.
Roman Y, Oshige M, Lee Y-J, Goodwin K, Georgiadis MM, Hromas RA et al. Biochemical characterization of a SET and transposase fusion protein, metnase: its DNA binding and DNA cleavage activity. Biochemistry 2007; 46: 11369–11376.
Wolf A, Keil R, Gotzl O, Mun A, Schwarze K, Lederer M et al. The armadillo protein p0071 regulates Rho signalling during cytokinesis. Nat Cell Biol 2006; 8: 1432–1440.
Zhu H, Coppinger JA, Jang C-Y, Yates JR, Fang G . FAM29A promotes microtubule amplification via recruitment of the NEDD1–γ-tubulin complex to the mitotic spindle. J Cell Biol 2008; 183: 835–848.
Chattopadhyay S, Bielinsky A-K . Human Mcm10 regulates the catalytic subunit of DNA polymerase-α and prevents DNA damage during replication. Mol Biol Cell 2007; 18: 4085–4095.
Klon AE, Héroux A, Ross LJ, Pathak V, Johnson CA, Piper JR et al. Atomic structures of human dihydrofolate reductase complexed with NADPH and two lipophilic antifolates at 1.09 a and 1.05 a resolution. J Mol Biol 2002; 320: 677–693.
Nandi AK, Ford T, Fleksher D, Neuman B, Rapoport AP . Attenuation of DNA damage checkpoint by PBK, a novel mitotic kinase, involves protein–protein interaction with tumor suppressor p53. Biochem Biophys Res Commun 2007; 358: 181–188.
Nijman SMB, Huang TT, Dirac AMG, Brummelkamp TR, Kerkhoven RM, D'Andrea AD et al. The deubiquitinating enzyme USP1 regulates the fanconi anemia pathway. Mol Cell 2005; 17: 331–339.
Piao L, Nakagawa H, Ueda K, Chung S, Kashiwaya K, Eguchi H et al. C12orf48, termed PARP-1 binding protein, enhances poly(ADP-ribose) polymerase-1 (PARP-1) activity and protects pancreatic cancer cells from DNA damage. Genes Chromosomes Cancer 2011; 50: 13–24.
Dephoure N, Zhou C, Villén J, Beausoleil SA, Bakalarski CE, Elledge SJ et al. A quantitative atlas of mitotic phosphorylation. Proc Natl Acad Sci 2008; 105: 10762–10767.
Joshi B, Cameron A, Jagus R . Characterization of mammalian eIF4E-family members. Eur J Biochem 2004; 271: 2189–2203.
Kaser A, Bogengruber E, Hallegger M, Doppler E, Lepperdinger G, Jantsch M et al. Brix from Xenopus laevis and Brx1p from yeast define a new family of proteins involved in the biogenesis of large ribosomal subunits. Biol Chem 2001; 382: 1637–1647.
Gerondakis S, Grumont R, Gugasyan R, Wong L, Isomura I, Ho W et al. Unravelling the complexities of the NF-κB signalling pathway using mouse knockout and transgenic models. Oncogene 2006; 25: 6781–6799.
Basak S, VF-S Shih, Hoffmann A . Generation and activation of multiple dimeric transcription factors within the NF-κB signaling system. Mol Cell Biol 2008; 28: 3139–3150.
Dejardin E, Droin NM, Delhase M, Haas E, Cao Y, Makris C et al. The lymphotoxin-β receptor induces different patterns of gene expression via two NF-κB pathways. Immunity 2002; 17: 525–535.
Shih VF-S, Tsui R, Caldwell A, Hoffmann A . A single NFκB system for both canonical and non-canonical signaling. Cell Res 2011; 21: 86–102.
Zhao B, Barrera Luis A, Ersing I, Willox B, Schmidt Stefanie CS, Greenfeld H et al. The NF-κB genomic landscape in lymphoblastoid B cells. Cell Rep 2014; 8: 1595–1606.
Braggio E, Dogan A, Keats JJ, Chng WJ, Huang G, Matthews JM et al. Genomic analysis of marginal zone and lymphoplasmacytic lymphomas identified common and disease-specific abnormalities. Mod Pathol 2012; 25: 651–660.
Harvey D, Pointon JJ, Karaderi T, Appleton LH, Farrar C, Wordsworth BP . A common functional variant of endoplasmic reticulum aminopeptidase 2 (ERAP2) that reduces major histocompatibility complex class I expression is not associated with ankylosing spondylitis. Rheumatology 2011; 50: 1720–1721.
Kamphausen E, Kellert C, Abbas T, Akkad N, Tenzer S, Pawelec G et al. Distinct molecular mechanisms leading to deficient expression of ER-resident aminopeptidases in melanoma. Cancer Immunol Immunother 2010; 59: 1273–1284.
Zhu D, Ikpatt OF, Dubovy SR, Lossos C, Natkunam Y, Chapman-Fredricks JR et al. Molecular and genomic aberrations in chlamydophila psittaci negative ocular adnexal marginal zone lymphomas. Am J Hematol 2013; 88: 730–735.
Morin RD, Mendez-Lago M, Mungall AJ, Goya R, Mungall KL, Corbett RD et al. Frequent mutation of histone-modifying genes in non-Hodgkin lymphoma. Nature 2011; 476: 298–303.
Angus SP, Nevins JR . A role for mediator complex subunit MED13L in Rb/E2F-induced growth arrest. Oncogene 2012; 31: 4709–4717.
Marill J, Cresteil T, Lanotte M, Chabot GG . Identification of human cytochrome P450s involved in the formation of all-trans-retinoic acid principal metabolites. Mol Pharmacol 2000; 58: 1341–1348.
Shimizu N, Yoshikawa N, Ito N, Maruyama T, Suzuki Y, Takeda S-i et al. Crosstalk between glucocorticoid receptor and nutritional sensor mTOR in skeletal muscle. Cell Metab 2011; 13: 170–182.
Ansell SM, Hodge LS, Secreto FJ, Manske M, Braggio E, Price-Troska T et al. Activation of TAK1 by MYD88 L265P drives malignant B-cell growth in non-Hodgkin lymphoma. Blood Cancer J 2014; 4: e183.
Gerondakis S, Siebenlist U . Roles of the NF-κB pathway in lymphocyte development and function. Cold Spring Harb Perspect Biol 2010; 2: a000182.
Franzoso G, Carlson L, Poljak L, Shores EW, Epstein S, Leonardi A et al. Mice deficient in nuclear factor (NF)-κ B/p52 present with defects in humoral responses, germinal center reactions, and splenic microarchitecture. J Exp Med 1998; 187: 147–159.
Lo JC, Basak S, James ES, Quiambo RS, Kinsella MC, Alegre M-L et al. Coordination between NF-κB family members p50 and p52 is essential for mediating LTbetaR signals in the development and organization of secondary lymphoid tissues. Blood 2006; 107: 1048–1055.
Claudio E, Brown K, Park S, Wang H, Siebenlist U . BAFF-induced NEMO-independent processing of NF-κB2 in maturing B cells. Nat Immunol 2002; 3: 958–965.
Staudt LM . Oncogenic activation of NF-κB. Cold Spring Harb Perspect Biol 2010; 2: a000109.
Zhang B, Wang Z, Li T, Tsitsikov EN, Ding H-F . NF-κB2 mutation targets TRAF1 to induce lymphomagenesis. Blood 2007; 110: 743–751.
Acknowledgements
We thank Dr Sagar Lonial and Dr Jing Chen for their critical review and helpful comments. We thank Dr Anne J Novak for providing the L265P MYD88 lentivirus. CRF was supported by NIH (R21 CA158686 and K24 CA208132) and GCCDSW. ISL by the NIH (CA109335 and CA122105) and the Dwoskin and Rizzo Family and Fidelity. JK was supported by GCCDSW. LHB received funding support from the NIH (CA127910 and CA129968) and a GCCDSW. LB-M received funding from the Byron Davis Research Fund.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on the Oncogene website
Supplementary information
Rights and permissions
About this article
Cite this article
Guo, X., Koff, J., Moffitt, A. et al. Molecular impact of selective NFKB1 and NFKB2 signaling on DLBCL phenotype. Oncogene 36, 4224–4232 (2017). https://doi.org/10.1038/onc.2017.90
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2017.90
This article is cited by
-
Genome-wide CRISPR screens reveal synthetic lethal interaction between CREBBP and EP300 in diffuse large B-cell lymphoma
Cell Death & Disease (2021)
-
Identification of Hub Genes and Key Pathways Associated with Peripheral T-cell Lymphoma
Current Medical Science (2020)
-
NFKB2 gene expression in patients with peptic ulcer diseases and gastric cancer
Molecular Biology Reports (2020)
-
Novel phosphorylated TAK1 species with functional impact on NF-κB and β-catenin signaling in human Cutaneous T-cell lymphoma
Leukemia (2018)
-
Oncogenic MYD88 mutations in lymphoma: novel insights and therapeutic possibilities
Cancer Immunology, Immunotherapy (2018)