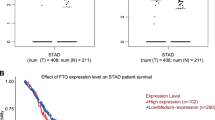
Abstract
Six GATA transcription factors play important roles in eukaryotic development. Among these, GATA2, an essential factor for the hematopoietic cell lineage, exhibits low expression in human gastric tissues, whereas GATA6, which is crucial for gastrointestinal development and differentiation, is frequently amplified and/or overexpressed in human gastric cancer. Interestingly, we found that GATA6 was overexpressed in human gastric cancer cells only when GATA2 expression was completely absent, thereby showing an inverse correlation between GATA2 and GATA6. In gastric cancer cells that express high GATA6 levels, a GATA2 CpG island is hypermethylated, repressing expression in these cells. In contrast, GATA6 expression is undetectable in GATA2-overexpressing gastric cancer cells, which lack GATA2 DNA methylation. Furthermore, PRC2 complex-mediated transcriptional silencing of GATA6 was observed in the GATA2-overexpressing cells. We also show that the GATA2 and PRC2 complexes are enriched within the GATA6 locus, and that the recruitment of the PRC2 complex is impaired by disrupting GATA2 expression, resulting in GATA6 upregulation. In addition, ectopic GATA2 expression significantly downregulates GATA6 expression, suggesting GATA2 directly represses GATA6. Furthermore, GATA6 downregulation showed antitumor activity by inducing growth arrest. Finally, we show that aberrant GATA2 methylation occurs early during the multistep process of gastric carcinogenesis regardless of Helicobacter pylori infection. Taken together, GATA2 dysregulation by epigenetic modification is associated with unfavorable phenotypes in human gastric cancer cells by allowing GATA6 expression.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout









Similar content being viewed by others
References
Molkentin JD . The zinc finger-containing transcription factors GATA-4, -5, and -6. Ubiquitously expressed regulators of tissue-specific gene expression. J Biol Chem 2000; 275: 38949–38952.
Bresnick EH, Katsumura KR, Lee HY, Johnson KD, Perkins AS . Master regulatory GATA transcription factors: mechanistic principles and emerging links to hematologic malignancies. Nucleic Acids Res 2012; 40: 5819–5831.
Johnson KD, Hsu AP, Ryu MJ, Wang J, Gao X, Boyer ME et al. Cis-element mutated in GATA2-dependent immunodeficiency governs hematopoiesis and vascular integrity. J Clin Invest 2012; 122: 3692–3704.
Kang Y, Kim YW, Kang J, Yun WJ, Kim A . Erythroid specific activator GATA-1-dependent interactions between CTCF sites around the beta-globin locus. Biochim Biophy Acta 2017; 1860: 416–426.
Kang Y, Kim YW, Yun J, Shin J, Kim A . KLF1 stabilizes GATA-1 and TAL1 occupancy in the human beta-globin locus. Biochim Biophys Acta 2015; 1849: 282–289.
Moriguchi T, Yamamoto M . A regulatory network governing Gata1 and Gata2 gene transcription orchestrates erythroid lineage differentiation. Int J Hematol 2014; 100: 417–424.
Bresnick EH, Lee HY, Fujiwara T, Johnson KD, Keles S . GATA switches as developmental drivers. J Biol Chem 2010; 285: 31087–31093.
Grass JA, Boyer ME, Pal S, Wu J, Weiss MJ, Bresnick EH . GATA-1-dependent transcriptional repression of GATA-2 via disruption of positive autoregulation and domain-wide chromatin remodeling. Proc Natl Acad Sci USA 2003; 100: 8811–8816.
Katsumura KR, Ong IM, DeVilbiss AW, Sanalkumar R, Bresnick EH . GATA factor-dependent positive-feedback circuit in acute myeloid leukemia cells. Cell Rep 2016; 16: 2428–2441.
Beuling E, Baffour-Awuah NY, Stapleton KA, Aronson BE, Noah TK, Shroyer NF et al. GATA factors regulate proliferation, differentiation, and gene expression in small intestine of mature mice. Gastroenterology 2011; 140: 1219–1229 e1211-1212.
Morrisey EE, Ip HS, Tang Z, Lu MM, Parmacek MS . GATA-5: a transcriptional activator expressed in a novel temporally and spatially-restricted pattern during embryonic development. Dev Biol 1997; 183: 21–36.
Morrisey EE, Tang Z, Sigrist K, Lu MM, Jiang F, Ip HS et al. GATA6 regulates HNF4 and is required for differentiation of visceral endoderm in the mouse embryo. Genes Dev 1998; 12: 3579–3590.
Cheung WK, Zhao M, Liu Z, Stevens LE, Cao PD, Fang JE et al. Control of alveolar differentiation by the lineage transcription factors GATA6 and HOPX inhibits lung adenocarcinoma metastasis. Cancer Cell 2013; 23: 725–738.
Capo-chichi CD, Cai KQ, Testa JR, Godwin AK, Xu XX . Loss of GATA6 leads to nuclear deformation and aneuploidy in ovarian cancer. Mol Cell Biol 2009; 29: 4766–4777.
Lin L, Bass AJ, Lockwood WW, Wang Z, Silvers AL, Thomas DG et al. Activation of GATA binding protein 6 (GATA6) sustains oncogenic lineage-survival in esophageal adenocarcinoma. Proc Natl Acad Sci USA 2012; 109: 4251–4256.
Haveri H, Westerholm-Ormio M, Lindfors K, Maki M, Savilahti E, Andersson LC et al. Transcription factors GATA-4 and GATA-6 in normal and neoplastic human gastrointestinal mucosa. BMC Gastroenterol 2008; 8: 9.
Chia NY, Deng N, Das K, Huang D, Hu L, Zhu Y et al. Regulatory crosstalk between lineage-survival oncogenes KLF5, GATA4 and GATA6 cooperatively promotes gastric cancer development. Gut 2015; 64: 707–719.
Deng N, Goh LK, Wang H, Das K, Tao J, Tan IB et al. A comprehensive survey of genomic alterations in gastric cancer reveals systematic patterns of molecular exclusivity and co-occurrence among distinct therapeutic targets. Gut 2012; 61: 673–684.
Sulahian R, Casey F, Shen J, Qian ZR, Shin H, Ogino S et al. An integrative analysis reveals functional targets of GATA6 transcriptional regulation in gastric cancer. Oncogene 2014; 33: 5637–5648.
Baylin SB, Jones PA . A decade of exploring the cancer epigenome - biological and translational implications. Nat Rev Cancer 2011; 11: 726–734.
Hattori N, Ushijima T . Compendium of aberrant DNA methylation and histone modifications in cancer. Biochem Biophysical Res Commun 2014; 455: 3–9.
Deng D, Liu Z, Du Y . Epigenetic alterations as cancer diagnostic, prognostic, and predictive biomarkers. Adv Genet 2010; 71: 125–176.
Liu-Chittenden Y, Jain M, Gaskins K, Wang S, Merino MJ, Kotian S et al. RARRES2 functions as a tumor suppressor by promoting beta-catenin phosphorylation/degradation and inhibiting p38 phosphorylation in adrenocortical carcinoma. Oncogene 2017; 36: 3541–3552.
Song SH, Han SW, Bang YJ . Epigenetic-based therapies in cancer: progress to date. Drugs 2011; 71: 2391–2403.
Liu Z, Zhang J, Gao Y, Pei L, Zhou J, Gu L et al. Large-scale characterization of DNA methylation changes in human gastric carcinomas with and without metastasis. Clin Cancer Res 2014; 20: 4598–4612.
Nakajima T, Maekita T, Oda I, Gotoda T, Yamamoto S, Umemura S et al. Higher methylation levels in gastric mucosae significantly correlate with higher risk of gastric cancers. Cancer Epidemiol Biomarkers Prev 2006; 15: 2317–2321.
Karimi P, Islami F, Anandasabapathy S, Freedman ND, Kamangar F . Gastric cancer: descriptive epidemiology, risk factors, screening, and prevention. Cancer Epidemiol Biomarkers Prev 2014; 23: 700–713.
Dyson MT, Roqueiro D, Monsivais D, Ercan CM, Pavone ME, Brooks DC et al. Genome-wide DNA methylation analysis predicts an epigenetic switch for GATA factor expression in endometriosis. PLoS Genet 2014; 10: e1004158.
Shih AH, Jiang Y, Meydan C, Shank K, Pandey S, Barreyro L et al. Mutational cooperativity linked to combinatorial epigenetic gain of function in acute myeloid leukemia. Cancer Cell 2015; 27: 502–515.
Kim HP, Cho GA, Han SW, Shin JY, Jeong EG, Song SH et al. Novel fusion transcripts in human gastric cancer revealed by transcriptome analysis. Oncogene 2014; 33: 5434–5441.
Ku JL, Park JG . Biology of SNU cell lines. Cancer Res Treat 2005; 37: 1–19.
Rodriguez-Paredes M, Esteller M . Cancer epigenetics reaches mainstream oncology. Nat Med 2011; 17: 330–339.
Pan X, Minegishi N, Harigae H, Yamagiwa H, Minegishi M, Akine Y et al. Identification of human GATA-2 gene distal IS exon and its expression in hematopoietic stem cell fractions. J Biochem 2000; 127: 105–112.
Yun J, Song SH, Kang JY, Park J, Kim HP, Han SW et al. Reduced cohesin destabilizes high-level gene amplification by disrupting pre-replication complex bindings in human cancers with chromosomal instability. Nucleic Acids Res 2016; 44: 558–572.
Ping N, Sun A, Song Y, Wang Q, Yin J, Cheng W et al. Exome sequencing identifies highly recurrent somatic GATA2 and CEBPA mutations in acute erythroid leukemia. Leukemia 2017; 31: 195–202.
Kang JY, Song SH, Yun J, Jeon MS, Kim HP, Han SW et al. Disruption of CTCF/cohesin-mediated high-order chromatin structures by DNA methylation downregulates PTGS2 expression. Oncogene 2015; 34: 5677–5684.
Endoh M, Endo TA, Endoh T, Fujimura Y, Ohara O, Toyoda T et al. Polycomb group proteins Ring1A/B are functionally linked to the core transcriptional regulatory circuitry to maintain ES cell identity. Development 2008; 135: 1513–1524.
Kang X, Qi Y, Zuo Y, Wang Q, Zou Y, Schwartz RJ et al. SUMO-specific protease 2 is essential for suppression of polycomb group protein-mediated gene silencing during embryonic development. Mol Cell 2010; 38: 191–201.
Lu R, Yang A, Jin Y . Dual functions of T-box 3 (Tbx3) in the control of self-renewal and extraembryonic endoderm differentiation in mouse embryonic stem cells. J Biol Chem 2011; 286: 8425–8436.
Schuettengruber B, Chourrout D, Vervoort M, Leblanc B, Cavalli G . Genome regulation by polycomb and trithorax proteins. Cell 2007; 128: 735–745.
Cao R, Wang L, Wang H, Xia L, Erdjument-Bromage H, Tempst P et al. Role of histone H3 lysine 27 methylation in Polycomb-group silencing. Science 2002; 298: 1039–1043.
Shalem O, Sanjana NE, Hartenian E, Shi X, Scott DA, Mikkelsen TS et al. Genome-scale CRISPR-Cas9 knockout screening in human cells. Science 2014; 343: 84–87.
Yu M, Riva L, Xie H, Schindler Y, Moran TB, Cheng Y et al. Insights into GATA-1-mediated gene activation versus repression via genome-wide chromatin occupancy analysis. Mol Cell 2009; 36: 682–695.
Cancer Genome Atlas Research N, Kandoth C, Schultz N, Cherniack AD, Akbani R, Liu Y et al. Integrated genomic characterization of endometrial carcinoma. Nature 2013; 497: 67–73.
Kang JY, Song SH, Yun J, Jeon MS, Cha Y, Lee SH et al. Identification of long-range epigenetic silencing on chromosome 15q25 and its clinical implication in gastric cancer. Am J Pathol 2015; 185: 666–678.
Tahara T, Arisawa T . DNA methylation as a molecular biomarker in gastric cancer. Epigenomics 2015; 7: 475–486.
Park SY, Yoo EJ, Cho NY, Kim N, Kang GH . Comparison of CpG island hypermethylation and repetitive DNA hypomethylation in premalignant stages of gastric cancer, stratified for Helicobacter pylori infection. J Pathol 2009; 219: 410–416.
Matsusaka K, Funata S, Fukayama M, Kaneda A . DNA methylation in gastric cancer, related to Helicobacter pylori and Epstein-Barr virus. World J Gastroenterol 2014; 20: 3916–3926.
Maekita T, Nakazawa K, Mihara M, Nakajima T, Yanaoka K, Iguchi M et al. High levels of aberrant DNA methylation in Helicobacter pylori-infected gastric mucosae and its possible association with gastric cancer risk. Clin Cancer Res 2006; 12: 989–995.
Ushijima T, Hattori N . Molecular pathways: involvement of Helicobacter pylori-triggered inflammation in the formation of an epigenetic field defect, and its usefulness as cancer risk and exposure markers. Clin Cancer Res 2012; 18: 923–929.
Song SH, Hou C, Dean A . A positive role for NLI/Ldb1 in long-range beta-globin locus control region function. Mol Cell 2007; 28: 810–822.
Acknowledgements
This research was supported by a grant of the Korea Health Technology R&D Project through the Korea Health Industry Development Institute, funded by the Ministry of Health & Welfare, Republic of Korea (HI14C1277), by the Bio & Medical Technology Development Program of the National Research Foundation (NRF) funded by the Ministry of Science, ICT & Future Planning, Republic of Korea (NRF-2016M3A9B6026918, NRF-2016M3A9B6026921 and 2011-0030049) and by Basic Science Research Program through the NRF funded by the Ministry of Education, Republic of Korea (NRF-2016R1D1A1B03930736).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on the Oncogene website
Rights and permissions
About this article
Cite this article
Song, S., Jeon, M., Nam, J. et al. Aberrant GATA2 epigenetic dysregulation induces a GATA2/GATA6 switch in human gastric cancer. Oncogene 37, 993–1004 (2018). https://doi.org/10.1038/onc.2017.397
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2017.397
This article is cited by
-
Role of the pioneer transcription factor GATA2 in health and disease
Journal of Molecular Medicine (2023)
-
Prognostic and predictive biomarkers for response to neoadjuvant chemoradiation in esophageal adenocarcinoma
Biomarker Research (2022)
-
The dynamic alteration of transcriptional regulation by crucial TFs during tumorigenesis of gastric cancer
Molecular Medicine (2022)
-
5-AZA-dC induces epigenetic changes associated with modified glycosylation of secreted glycoproteins and increased EMT and migration in chemo-sensitive cancer cells
Clinical Epigenetics (2021)
-
CircPIP5K1A facilitates gastric cancer progression via miR-376c-3p/ZNF146 axis
Cancer Cell International (2020)