Abstract
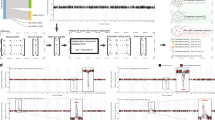
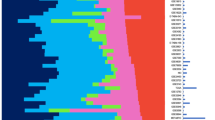
Transcriptional profile-based subtypes of cancer are often viewed as identifying different diseases from the same tissue origin. Understanding the mechanisms driving the subtypes may be key in development of novel therapeutics but is challenged by lineage-specific expression signals. Using a t-test statistics approach, we compared gene expression subtypes across 12 tumor types, which identified eight transcriptional superclusters characterized by commonly activated disease pathways and similarities in gene expression. One of the largest superclusters was determined by the upregulation of a proliferation signature, significant enrichment in TP53 mutations, genomic loss of CDKN2A (p16ARF), evidence of increased numbers of DNA double strand breaks and high expression of cyclin B1 protein. These correlations suggested that abrogation of the P53-mediated apoptosis response to DNA damage results in activation of cell cycle pathways and represents a common theme in cancer. A second consistent pattern, observed in 9 of 11 solid tumor types, was a subtype related to an activated tumor-associated stroma. The similarity in transcriptional footprints across cancers suggested that tumor subtypes are commonly unified by a limited number of molecular themes.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
TCGA. Comprehensive genomic characterization defines human glioblastoma genes and core pathways. Nature 2008; 455: 1061–1068.
Brennan CW, Verhaak RG, McKenna A, Campos B, Noushmehr H, Salama SR et al. The somatic genomic landscape of glioblastoma. Cell 2013; 155: 462–477.
TCGA. Integrated genomic analyses of ovarian carcinoma. Nature 2011; 474: 609–615.
TCGA. Comprehensive genomic characterization of squamous cell lung cancers. Nature 2012; 489: 519–525.
TCGA. Comprehensive molecular characterization of human colon and rectal cancer. Nature 2012; 487: 330–337.
TCGA. Comprehensive molecular portraits of human breast tumours. Nature 2012; 490: 61–70.
TCGA. Comprehensive molecular characterization of clear cell renal cell carcinoma. Nature 2013; 499: 43–49.
TCGA. Genomic and epigenomic landscapes of adult de novo acute myeloid leukemia. N Engl J Med 2013; 368: 2059–2074.
TCGA. Comprehensive molecular characterization of urothelial bladder carcinoma. Nature 2014; 507: 315–322.
Perou CM, Sørlie T, Eisen MB, van de Rijn M, Jeffrey SS, Rees CA et al. Molecular portraits of human breast tumours. Nature 2000; 406: 747–752.
Verhaak RG, Wouters BJ, Erpelinck CA, Abbas S, Beverloo HB, Lugthart S et al. Prediction of molecular subtypes in acute myeloid leukemia based on gene expression profiling. Haematologica 2009; 94: 131–134.
Verhaak RG, Hoadley KA, Purdom E, Wang V, Qi Y, Wilkerson MD et al. Integrated genomic analysis identifies clinically relevant subtypes of glioblastoma characterized by abnormalities in PDGFRA, IDH1, EGFR, and NF1. Cancer Cell 2010; 17: 98–110.
van de Vijver MJ, He YD, van't Veer LJ, Dai H, Hart AA, Voskuil DW et al. A gene-expression signature as a predictor of survival in breast cancer. N Engl J Med 2002; 347: 1999–2009.
Verhaak RG, Tamayo P, Yang JY, Hubbard D, Zhang H, Creighton CJ et al. Prognostically relevant gene signatures of high-grade serous ovarian carcinoma. J Clin Invest 2013; 123: 517–525.
Valk PJ, Verhaak RG, Beijen MA, Erpelinck CA, Barjesteh van Waalwijk van Doorn-Khosrovani S, Boer JM et al. Prognostically useful gene-expression profiles in acute myeloid leukemia. N Engl J Med 2004; 350: 1617–1628.
Brunet JP, Tamayo P, Golub TR, Mesirov JP . Metagenes and molecular pattern discovery using matrix factorization. Proc Natl Acad Sci USA 2004; 101: 4164–4169.
Barbie DA, Tamayo P, Boehm JS, Kim SY, Moody SE, Dunn IF et al. Systematic RNA interference reveals that oncogenic KRAS-driven cancers require TBK1. Nature 2009; 462: 108–112.
Vaske CJ, Benz SC, Sanborn JZ, Earl D, Szeto C, Zhu J et al. Inference of patient-specific pathway activities from multi-dimensional cancer genomics data using PARADIGM. Bioinformatics 2010; 26: i237–i245.
Subramanian A, Tamayo P, Mootha VK, Mukherjee S, Ebert BL, Gillette MA et al. Gene set enrichment analysis: A knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci USA 2005; 102: 15545–15550.
Heiser LM, Sadanandam A, Kuo WL, Benz SC, Goldstein TC, Ng S et al. Subtype and pathway specific responses to anticancer compounds in breast cancer. Proc Natl Acad Sci USA 2012; 109: 2724–2729.
Yoshihara K, Shahmoradgoli M, Martínez E, Vegesna R, Kim H, Torres-Garcia W et al. Inferring tumour purity and stromal and immune cell admixture from expression data. Nat Commun 2013; 4: 2612.
Carter SL, Cibulskis K, Helman E, McKenna A, Shen H, Zack T et al. Absolute quantification of somatic DNA alterations in human cancer. Nat Biotechnol 2012; 30: 413–421.
Martinsson-Ahlzén HS, Liberal V, Grünenfelder B, Chaves SR, Spruck CH, Reed SI . Cyclin-dependent kinase-associated proteins Cks1 and Cks2 are essential during early embryogenesis and for cell cycle progression in somatic cells. Mol Cell Biol 2008; 28: 5698–5709.
Li J, Bennett K, Stukalov A, Fang B, Zhang G, Yoshida T et al. Perturbation of the mutated EGFR interactome identifies vulnerabilities and resistance mechanisms. Mol Syst Biol 2013; 9: 705.
Zheng Y, Zhang C, Croucher DR, Soliman MA, St-Denis N, Pasculescu A et al. Temporal regulation of EGF signalling networks by the scaffold protein Shc1. Nature 2013; 499: 166–171.
Bhattacharjee A, Richards WG, Staunton J, Li C, Monti S, Vasa P et al. Classification of human lung carcinomas by mRNA expression profiling reveals distinct adenocarcinoma subclasses. Proc Natl Acad Sci USA 2001; 98: 13790–13795.
Wolchok JD, Kluger H, Callahan MK, Postow MA, Rizvi NA, Lesokhin AM et al. Nivolumab plus ipilimumab in advanced melanoma. N Engl J Med 2013; 369: 122–133.
Hamid O, Robert C, Daud A, Hodi FS, Hwu WJ, Kefford R et al. Safety and tumor responses with lambrolizumab (anti-PD-1) in melanoma. N Engl J Med 2013; 369: 134–144.
Topalian SL, Hodi FS, Brahmer JR, Gettinger SN, Smith DC, McDermott DF et al. Safety, activity, and immune correlates of anti-PD-1 antibody in cancer. N Engl J Med 2012; 366: 2443–2454.
Hodi FS, O'Day SJ, McDermott DF, Weber RW, Sosman JA, Haanen JB et al. Improved survival with ipilimumab in patients with metastatic melanoma. N Engl J Med 2010; 363: 711–723.
Parker JS, Mullins M, Cheang MC, Leung S, Voduc D, Vickery T et al. Supervised risk predictor of breast cancer based on intrinsic subtypes. J Clin Oncol 2009; 27: 1160–1167.
Zack TI, Schumacher SE, Carter SL, Cherniack AD, Saksena G, Tabak B et al. Pan-cancer patterns of somatic copy number alteration. Nat Genet 2013; 45: 1134–1140.
Tusher VG, Tibshirani R, Chu G . Significance analysis of microarrays applied to the ionizing radiation response. Proc Natl Acad Sci USA 2001; 98: 5116–5121.
Barretina J, Caponigro G, Stransky N, Venkatesan K, Margolin AA, Kim S et al. The Cancer Cell Line Encyclopedia enables predictive modelling of anticancer drug sensitivity. Nature 2012; 483: 603–607.
Everitt BS, Landau S, Leese M, Stahl D, Shewhart WA, Wilks SS . Cluster Analysis. John Wiley & Sons Ltd: West Sussex, UK, 2011.
Zheng S, Fu J, Vegesna R, Mao Y, Heathcock LE, Torres-Garcia W et al. A survey of intragenic breakpoints in glioblastoma identifies a distinct subset associated with poor survival. Genes Dev 2013; 27: 1462–1472.
Tibshirani R, Hastie T, Narasimhan B, Chu G . Diagnosis of multiple cancer types by shrunken centroids of gene expression. Proc Natl Acad Sci USA 2002; 99: 6567–6572.
Noble WS . What is a support vector machine? Nat Biotechnol 2006; 24: 1565–1567.
Croft D, Mundo AF, Haw R, Milacic M, Weiser J, Wu G et al. The Reactome pathway knowledgebase. Nucleic Acids Res 2014; 42 (Database issue): D472–D477.
Kanehisa M, Goto S, Sato Y, Kawashima M, Furumichi M, Tanabe M . Data, information, knowledge and principle: back to metabolism in KEGG. Nucleic Acids Res 2014; 42 (Database issue): D199–D205.
Schaefer CF, Anthony K, Krupa S, Buchoff J, Day M, Hannay T et al. PID: the Pathway Interaction Database. Nucleic Acids Res 2009; 37 (Database issue): D674–D679.
Rosner B . Percentage Points for a Generalized ESD Many Outlier Procedure. Technometrics 1983; 25: 165–172.
Team RC . RA: Language and Environment for Statistical Computing, Vol. 13. R Foundation for Statistical Computing: Vienna, Austria, 2013, pp 497–512.
Acknowledgements
The results published here are in whole or part based upon data generated by The Cancer Genome Atlas pilot project established by the NCI and NHGRI. Information about TCGA and the investigators and institutions who constitute the TCGA research network can be found at ‘http://cancergenome.nih.gov’. This work was supported in part by U.S. National Cancer Institute (NCI; MD Anderson TCGA Genome Data Analysis Center) grant numbers CA143883 and CA083639, and MD Anderson Cancer Center Support Grant P30 CA016672. HK is supported in part by the Odyssey Program and the Theodore N, Law Endowment for Scientific Achievement at The University of Texas MD Anderson Cancer Center.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare that they have no conflicts of interest.
Additional information
Supplementary Information accompanies this paper on the Oncogene website
Supplementary information
Rights and permissions
About this article
Cite this article
Martínez, E., Yoshihara, K., Kim, H. et al. Comparison of gene expression patterns across 12 tumor types identifies a cancer supercluster characterized by TP53 mutations and cell cycle defects. Oncogene 34, 2732–2740 (2015). https://doi.org/10.1038/onc.2014.216
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2014.216