Abstract
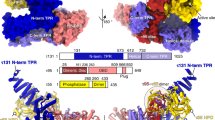
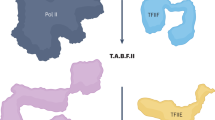
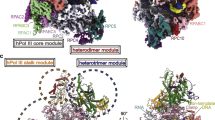
Core Factor (CF) is a conserved RNA polymerase (Pol) I general transcription factor comprising Rrn6, Rrn11 and the TFIIB-related subunit Rrn7. CF binds TATA-binding protein (TBP), Pol I and the regulatory factors Rrn3 and upstream activation factor. We used chemical cross-linking–MS to determine the molecular architecture of CF and its interactions with TBP. The CF subunits assemble through an interconnected network of interactions between five structural domains that are conserved in orthologous subunits of the human Pol I factor SL1. We validated the cross-linking–derived model through a series of genetic and biochemical assays. Our combined results show the architecture of CF and the functions of the CF subunits in assembly of the complex. We extend these findings to model how CF assembles into the Pol I preinitiation complex, providing new insight into the roles of CF, TBP and Rrn3.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Change history
24 August 2014
In the version of this article initially published online, there was a mistake in a grant number. The error has been corrected for the PDF and HTML versions of this article.
References
Knutson, B.A. & Hahn, S. TFIIB-related factors in RNA polymerase I transcription. Biochim. Biophys. Acta 1829, 265–273 (2013).
Schneider, D.A. RNA polymerase I activity is regulated at multiple steps in the transcription cycle: recent insights into factors that influence transcription elongation. Gene 493, 176–184 (2012).
Bedwell, G.J., Appling, F.D., Anderson, S.J. & Schneider, D.A. Efficient transcription by RNA polymerase I using recombinant core factor. Gene 492, 94–99 (2012).
Lin, C.W. et al. A novel 66-kilodalton protein complexes with Rrn6, Rrn7, and TATA-binding protein to promote polymerase I transcription initiation in Saccharomyces cerevisiae. Mol. Cell. Biol. 16, 6436–6443 (1996).
Lalo, D., Steffan, J.S., Dodd, J.A. & Nomura, M. RRN11 encodes the third subunit of the complex containing Rrn6p and Rrn7p that is essential for the initiation of rDNA transcription by yeast RNA polymerase I. J. Biol. Chem. 271, 21062–21067 (1996).
Russell, J. & Zomerdijk, J.C. The RNA polymerase I transcription machinery. Biochem. Soc. Symp. 73, 203–216 (2006).
Gorski, J.J. et al. A novel TBP-associated factor of SL1 functions in RNA polymerase I transcription. EMBO J. 26, 1560–1568 (2007).
Denissov, S. et al. Identification of novel functional TBP-binding sites and general factor repertoires. EMBO J. 26, 944–954 (2007).
Milkereit, P., Schultz, P. & Tschochner, H. Resolution of RNA polymerase I into dimers and monomers and their function in transcription. Biol. Chem. 378, 1433–1443 (1997).
Milkereit, P. & Tschochner, H. A specialized form of RNA polymerase I, essential for initiation and growth-dependent regulation of rRNA synthesis, is disrupted during transcription. EMBO J. 17, 3692–3703 (1998).
Peyroche, G. et al. The recruitment of RNA polymerase I on rDNA is mediated by the interaction of the A43 subunit with Rrn3. EMBO J. 19, 5473–5482 (2000).
Blattner, C. et al. Molecular basis of Rrn3-regulated RNA polymerase I initiation and cell growth. Genes Dev. 25, 2093–2105 (2011).
Stepanchick, A. et al. DNA binding by the ribosomal DNA transcription factor rrn3 is essential for ribosomal DNA transcription. J. Biol. Chem. 288, 9135–9144 (2013).
Aprikian, P., Moorefield, B. & Reeder, R.H. New model for the yeast RNA polymerase I transcription cycle. Mol. Cell. Biol. 21, 4847–4855 (2001).
Engel, C., Sainsbury, S., Cheung, A.C., Kostrewa, D. & Cramer, P. RNA polymerase I structure and transcription regulation. Nature 502, 650–655 (2013).
Fernández-Tornero, C. et al. Crystal structure of the 14-subunit RNA polymerase I. Nature 502, 644–649 (2013).
Vannini, A. A structural perspective on RNA polymerase I and RNA polymerase III transcription machineries. Biochim. Biophys. Acta 1829, 258–264 (2013).
Vannini, A. & Cramer, P. Conservation between the RNA polymerase I, II, and III transcription initiation machineries. Mol. Cell 45, 439–446 (2012).
Geiger, S.R. et al. RNA polymerase I contains a TFIIF-related DNA-binding subcomplex. Mol. Cell 39, 583–594 (2010).
Zomerdijk, J. Structural biology: pivotal findings for a transcription machine. Nature 502, 629–630 (2013).
Bywater, M.J., Pearson, R.B., McArthur, G.A. & Hannan, R.D. Dysregulation of the basal RNA polymerase transcription apparatus in cancer. Nat. Rev. Cancer 13, 299–314 (2013).
Bywater, M.J. et al. Inhibition of RNA polymerase I as a therapeutic strategy to promote cancer-specific activation of p53. Cancer Cell 22, 51–65 (2012).
Drygin, D. et al. Targeting RNA polymerase I with an oral small molecule CX-5461 inhibits ribosomal RNA synthesis and solid tumor growth. Cancer Res. 71, 1418–1430 (2011).
Drygin, D., Rice, W.G. & Grummt, I. The RNA polymerase I transcription machinery: an emerging target for the treatment of cancer. Annu. Rev. Pharmacol. Toxicol. 50, 131–156 (2010).
Hannan, K.M., Sanij, E., Rothblum, L.I., Hannan, R.D. & Pearson, R.B. Dysregulation of RNA polymerase I transcription during disease. Biochim. Biophys. Acta 1829, 342–360 (2013).
Knutson, B.A. & Hahn, S. Yeast Rrn7 and human TAF1B are TFIIB-related RNA polymerase I general transcription factors. Science 333, 1637–1640 (2011).
Naidu, S., Friedrich, J.K., Russell, J. & Zomerdijk, J.C. TAF1B is a TFIIB-like component of the basal transcription machinery for RNA polymerase I. Science 333, 1640–1642 (2011).
Allan, R.K. & Ratajczak, T. Versatile TPR domains accommodate different modes of target protein recognition and function. Cell Stress Chaperones 16, 353–367 (2011).
D'Andrea, L.D. & Regan, L. TPR proteins: the versatile helix. Trends Biochem. Sci. 28, 655–662 (2003).
Stirnimann, C.U., Petsalaki, E., Russell, R.B. & Muller, C.W. WD40 proteins propel cellular networks. Trends Biochem. Sci. 35, 565–574 (2010).
Xu, C. & Min, J. Structure and function of WD40 domain proteins. Protein Cell 2, 202–214 (2011).
Yang, B. et al. Identification of cross-linked peptides from complex samples. Nat. Methods 9, 904–906 (2012).
Chen, Z.A. et al. Architecture of the RNA polymerase II-TFIIF complex revealed by cross-linking and mass spectrometry. EMBO J. 29, 717–726 (2010).
Kalkhof, S. & Sinz, A. Chances and pitfalls of chemical cross-linking with amine-reactive N-hydroxysuccinimide esters. Anal. Bioanal. Chem. 392, 305–312 (2008).
Mädler, S., Bich, C., Touboul, D. & Zenobi, R. Chemical cross-linking with NHS esters: a systematic study on amino acid reactivities. J. Mass Spectrom. 44, 694–706 (2009).
Müller, M.Q. & Sinz, A. Chemical cross-linking and high-resolution mass spectrometry to study protein-drug interactions. Methods Mol. Biol. 803, 205–218 (2012).
Kim, D.E., Chivian, D. & Baker, D. Protein structure prediction and analysis using the Robetta server. Nucleic Acids Res. 32, W526–W531 (2004).
Chao, W.C., Kulkarni, K., Zhang, Z., Kong, E.H. & Barford, D. Structure of the mitotic checkpoint complex. Nature 484, 208–213 (2012).
ter Haar, E., Musacchio, A., Harrison, S.C. & Kirchhausen, T. Atomic structure of clathrin: a β propeller terminal domain joins an α zigzag linker. Cell 95, 563–573 (1998).
Colbert, T. & Hahn, S. A yeast TFIIB-related factor involved in RNA polymerase III transcription. Genes Dev. 6, 1940–1949 (1992).
Hahn, S. & Roberts, S. The zinc ribbon domains of the general transcription factors TFIIB and Brf: conserved functional surfaces but different roles in transcription initiation. Genes Dev. 14, 719–730 (2000).
Kassavetis, G.A. & Geiduschek, E.P. Transcription factor TFIIIB and transcription by RNA polymerase III. Biochem. Soc. Trans. 34, 1082–1087 (2006).
Khoo, S.K., Wu, C.C., Lin, Y.C., Lee, J.C. & Chen, H.T. Mapping the protein interaction network for the TFIIB-related factor Brf1 in the RNA polymerase III pre-initiation complex. Mol. Cell. Biol. 34, 551–559 (2014).
Schramm, L. & Hernandez, N. Recruitment of RNA polymerase III to its target promoters. Genes Dev. 16, 2593–2620 (2002).
Steffan, J.S., Keys, D.A., Dodd, J.A. & Nomura, M. The role of TBP in rDNA transcription by RNA polymerase I in Saccharomyces cerevisiae: TBP is required for upstream activation factor-dependent recruitment of core factor. Genes Dev. 10, 2551–2563 (1996).
Steffan, J.S., Keys, D.A., Vu, L. & Nomura, M. Interaction of TATA-binding protein with upstream activation factor is required for activated transcription of ribosomal DNA by RNA polymerase I in Saccharomyces cerevisiae in vivo. Mol. Cell. Biol. 18, 3752–3761 (1998).
Rudloff, U., Eberhard, D. & Grummt, I. The conserved core domain of the human TATA binding protein is sufficient to assemble the multisubunit RNA polymerase I-specific transcription factor SL1. Proc. Natl. Acad. Sci. USA 91, 8229–8233 (1994).
Grünberg, S. & Hahn, S. Structural insights into transcription initiation by RNA polymerase II. Trends Biochem. Sci. 38, 603–611 (2013).
Grünberg, S., Warfield, L. & Hahn, S. Architecture of the RNA polymerase II preinitiation complex and mechanism of ATP-dependent promoter opening. Nat. Struct. Mol. Biol. 19, 788–796 (2012).
He, Y., Fang, J., Taatjes, D.J. & Nogales, E. Structural visualization of key steps in human transcription initiation. Nature 495, 481–486 (2013).
Murakami, K. et al. Architecture of an RNA polymerase II transcription pre-initiation complex. Science 342, 1238724 (2013).
Wu, C.C. et al. RNA polymerase III subunit architecture and implications for open promoter complex formation. Proc. Natl. Acad. Sci. USA 109, 19232–19237 (2012).
Wu, C.C., Lin, Y.C. & Chen, H.T. The TFIIF-like Rpc37/53 dimer lies at the center of a protein network to connect TFIIIC, Bdp1, and the RNA polymerase III active center. Mol. Cell. Biol. 31, 2715–2728 (2011).
Cavanaugh, A.H., Evans, A. & Rothblum, L.I. Mammalian Rrn3 is required for the formation of a transcription competent preinitiation complex containing RNA polymerase I. Gene Expr. 14, 131–147 (2008).
Kim, Y., Geiger, J.H., Hahn, S. & Sigler, P.B. Crystal structure of a yeast TBP/TATA-box complex. Nature 365, 512–520 (1993).
Bedwell, G.J., Appling, F.D., Anderson, S.J. & Schneider, D.A. Efficient transcription by RNA polymerase I using recombinant core factor. Gene 492, 94–99 (2012).
Studier, F.W. Protein production by auto-induction in high density shaking cultures. Protein Expr. Purif. 41, 207–234 (2005).
Schultz, M.C., Choe, S.Y. & Reeder, R.H. Specific initiation by RNA polymerase I in a whole-cell extract from yeast. Proc. Natl. Acad. Sci. USA 88, 1004–1008 (1991).
Fishburn, J. & Hahn, S. Architecture of the yeast RNA polymerase II open complex and regulation of activity by TFIIF. Mol. Cell. Biol. 32, 12–25 (2012).
McGuffin, L.J., Bryson, K. & Jones, D.T. The PSIPRED protein structure prediction server. Bioinformatics 16, 404–405 (2000).
Ward, J.J., McGuffin, L.J., Bryson, K., Buxton, B.F. & Jones, D.T. The DISOPRED server for the prediction of protein disorder. Bioinformatics 20, 2138–2139 (2004).
Söding, J., Biegert, A. & Lupas, A.N. The HHpred interactive server for protein homology detection and structure prediction. Nucleic Acids Res. 33, W244–W248 (2005).
Yang, Z. et al. UCSF Chimera, MODELLER, and IMP: an integrated modeling system. J. Struct. Biol. 179, 269–278 (2012).
Sainsbury, S., Niesser, J. & Cramer, P. Structure and function of the initially transcribing RNA polymerase II–TFIIB complex. Nature 493, 437–440 (2013).
Napetschnig, J., Blobel, G. & Hoelz, A. Crystal structure of the N-terminal domain of the human protooncogene Nup214/CAN. Proc. Natl. Acad. Sci. USA 104, 1783–1788 (2007).
Wang, J., Dye, B.T., Rajashankar, K.R., Kurinov, I. & Schulman, B.A. Insights into anaphase promoting complex TPR subdomain assembly from a CDC26–APC6 structure. Nat. Struct. Mol. Biol. 16, 987–989 (2009).
Kim, D.E., Chivian, D. & Baker, D. Protein structure prediction and analysis using the Robetta server. Nucleic Acids Res. 32, W526–W531 (2004).
Wallner, B. & Elofsson, A. Can correct protein models be identified? Protein Sci. 12, 1073–1086 (2003).
Benkert, P., Kunzli, M. & Schwede, T. QMEAN server for protein model quality estimation. Nucleic Acids Res. 37, W510–W514 (2009).
Randall, A. & Baldi, P. SELECTpro: effective protein model selection using a structure-based energy function resistant to BLUNDERs. BMC Struct. Biol. 8, 52 (2008).
Bowie, J.U., Luthy, R. & Eisenberg, D. A method to identify protein sequences that fold into a known three-dimensional structure. Science 253, 164–170 (1991).
Schneidman-Duhovny, D., Inbar, Y., Nussinov, R. & Wolfson, H.J. PatchDock and SymmDock: servers for rigid and symmetric docking. Nucleic Acids Res. 33, W363–W367 (2005).
Yang, B. et al. Identification of cross-linked peptides from complex samples. Nat. Methods 9, 904–906 (2012).
Eng, J.K., Fischer, B., Grossmann, J. & Maccoss, M.J. A fast SEQUEST cross correlation algorithm. J. Proteome Res. 7, 4598–4602 (2008).
Nogi, Y., Vu, L. & Nomura, M. An approach for isolation of mutants defective in 35S ribosomal RNA synthesis in Saccharomyces cerevisiae. Proc. Natl. Acad. Sci. USA 88, 7026–7030 (1991).
Nogi, Y., Yano, R. & Nomura, M. Synthesis of large rRNAs by RNA polymerase II in mutants of Saccharomyces cerevisiae defective in RNA polymerase I. Proc. Natl. Acad. Sci. USA 88, 3962–3966 (1991).
Nelson, J., Denisenko, O. & Bomsztyk, K. The fast chromatin immunoprecipitation method. Methods Mol. Biol. 567, 45–57 (2009).
Nelson, J.D., Denisenko, O. & Bomsztyk, K. Protocol for the fast chromatin immunoprecipitation (ChIP) method. Nat. Protoc. 1, 179–185 (2006).
Zhang, Y. et al. The SWI/SNF chromatin remodeling complex influences transcription by RNA polymerase I in Saccharomyces cerevisiae. PLoS ONE 8, e56793 (2013).
Acknowledgements
We thank members of the S.H. laboratory and of the E. Young (University of Washington) and J. Ranish (Institute for Systems Biology) laboratories for comments throughout the course of the work. We also thank P. Gafkin and the Fred Hutchinson Cancer Research Center proteomics shared resources for assistance with MS and the P. Cramer laboratory (University of Munich) for sharing an early release of their yeast Pol I crystal structure. This work was supported by grants from the US National Institutes of Health (NIH) NIGMS (2RO1GM053451 to S.H. and 2P50 GM076547 to J.R.) and NCI (R21CA175849 to J.R.).
Author information
Authors and Affiliations
Contributions
B.A.K., J.L., J.R. and S.H. designed the experiments. B.A.K. performed all structural modeling, genetic assays and biochemical experiments. B.A.K. and J.L. performed the cross-linking experiments, and J.L. performed the database searches for the CXMS analysis. B.A.K. and S.H. prepared the manuscript. All authors discussed and interpreted the data and approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Sequence analysis and structural modeling of Core Factor subunits and domains.
(a-c) Immediately above the protein diagrams for Rrn7 (a), Rrn11 (b), and Rrn6 (c) are the structural and functional domain assignments and the 3D structural models. Position of BS3-reactive lysines and amine groups are shown as vertical purple bars within the protein diagrams. Immediately below are the PSIpred secondary structure predictions (Helix, red; Beta-strand, blue; Coil, grey) followed by the DISOPRED disorder predictions shown in green. The scale bar on the y-axis from 0 to 1 indicates the confidence of the predictions from low to high, respectively.
Supplementary Figure 2 Recombinant GST-Rrn7 complements growth in vivo but cannot rescue transcription activity of an rrn7Δ extract in vitro.
(a) RRN7 growth complementation assay. An Δrrn7 yeast strain containing a plasmid that expresses the rDNA locus under control of the GAL7 promoter was transformed with the indicated RRN7 expression constructs and spotted onto glucose complete plates and scored for growth (Wild-type (WT), +++). (b) In vitro transcription assays with a Pol I promoter plasmid and either WT or Δrrn7 yeast extracts in the presence or absence of GST-Rrn7. Pol I transcripts were detected by primer extension.
Supplementary Figure 3 Growth and biochemical phenotypes of Rrn7-deletion mutants.
(a) Growth of yeast strains with indicated mutations in Rrn7. Below the Rrn7 domain map are schematics of the various deletion variants used for the growth assays. Black bars denote regions included in the deletion construct. Each Rrn7 deletion derivative depicted was transformed in strains containing their respective gene deletions and the plasmid pNOY103 that expresses ribosomal RNA under control of the GAL7 promoter. Strains were grown in galactose containing media, streaked onto glucose-containing plates and scored for growth. WT (+++); Lethal (-). Spheres along the top of the domain maps indicate the approximate sites of BS3 crosslinking and are colored accordingly: orange, Rrn11; blue, Rrn6. (b) Assay for CF integrity defects in the Rrn7 deletion derivatives. WT and the indicated deletion variants of Rrn7-Flag were coexpressed in a WT strain that contained epitope tags on the other CF subunits. Each deletion variant was purified from whole-cell extracts and analyzed by Western blotting using the indicated epitope tag antibodies.
Supplementary Figure 4 Growth and biochemical phenotypes of Rrn11-deletion mutants.
(a) Growth of yeast strains with indicated mutations in Rrn11. Growth assays were as described in Supplementary Figure 3a. Spheres along the top of the domain maps indicate the approximate sites of BS3 crosslinking and are colored accordingly: green, Rrn7; blue, Rrn6. (b) Assay for CF integrity defects in the Rrn11 deletion derivatives. WT and indicated deletion variants of Rrn11-Myc were coexpressed in a WT strain that contained unique epitope tags on the other CF subunits. IP and Western blot analysis were performed as in Supplementary Figure 3b.
Supplementary Figure 5 Growth and biochemical phenotypes of Rrn6-deletion mutants.
(a,c) Growth of yeast strains with indicated mutations in Rrn6. Growth assay were done as described in Supplementary Figure 3a. Spheres along the top of the domain maps indicate the approximate sites of BS3 crosslinking and are colored accordingly: green, Rrn7; orange, Rrn11. N.D., not determined. (b,d) Assay for CF integrity defects in the Rrn6 deletion derivatives. WT and indicated deletion variants of Rrn6-Myc were coexpressed in a WT strain that contained unique epitope tags on the other CF subunits. IP and Western blot analysis were performed as in Supplementary Figure 3b.
Supplementary Figure 6 Recruitment of CF-deletion variants to the rDNA promoter.
(a) Location of PCR primer sets (red bars) within the 35S rDNA locus and Pol I promoter. Promoter, Pro; External Transcribed Spacer, ETS. (b-d) Strains containing either WT Rrn7 (b), Rrn11 (c), Rrn6 (d), and the indicated CF subunit derivatives were crosslinked with formaldehyde and analyzed by chromatin IP and qPCR using the indicated primers sets. Averages of biological duplicate experiments are expressed relative to WT, which was set at 1.0. Error bars indicate standard deviations. An asterisk denotes lethal CF mutants that retain complex integrity.
Supplementary Figure 7 Molecular topology of Core Factor and position of the Rrn7 CTD.
(a) Topological linkage map showing all the observed intermolecular and interdomain crosslinks between the CF subunit domains. Non-crosslinked lysine Cα atoms are depicted as spheres in the same color as the domain. Crosslinked lysine pairs are shown as red spheres connected by black lines. (b) Lysine residue Cα positions within the CF model that crosslink to the Rrn7-CTD are shown as green spheres connected by black lines to numbers indicating the crosslinked Rrn7-CTD lysine residue. Rrn7-CTD lysines K329 and K360 reside in the helical segment H1 and the linker between H2 and H3, respectively. Rrn7-CTD lysine K457 lies within the linker between H5 and H6, while the K473 and K495, 499, and 506 reside in H6 and H7, respectively. The green dashed outline indicates the approximate location of the Rrn7-CTD, suggested by the mapped Rrn7-CTD crosslink positions.
Supplementary Figure 8 Molecular topology and architecture of CF–TBP complex.
(a) Linkage map of crosslinked CF-TBP lysine residues. Schematics of TBP and each CF subunit showing their known and predicted domain organization. Purple bars denote lysine positions and the N-terminal amine, while red spheres connected by dashed black lines indicate intra- and inter-molecular crosslinked lysine pairs within TBP or between TBP and CF. (b) Intramolecular crosslinks within the TBP structure (1YTB). Non-crosslinked lysine Cα atoms are depicted as spheres. Cα atoms of crosslinked lysine pairs are depicted as red spheres connected by black lines (d-e) Model of the CF-TBP-DNA complex shown without (d) or with TBP and DNA (e). (c-f) Calculated Cα-Cα distances between crosslinked lysine pairs within TBP (c) or between CF subunits and TBP (f). Dashed grey line denotes the theoretical maximum crosslinking distance for BS3 of 30 Å.
Supplementary Figure 9 Full images of cropped panels.
The figure number and corresponding panel are indicated at the top of each image.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–9 and Supplementary Tables 1–5 (PDF 6211 kb)
Rights and permissions
About this article
Cite this article
Knutson, B., Luo, J., Ranish, J. et al. Architecture of the Saccharomyces cerevisiae RNA polymerase I Core Factor complex. Nat Struct Mol Biol 21, 810–816 (2014). https://doi.org/10.1038/nsmb.2873
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.2873
This article is cited by
-
Molecular determinants underlying functional innovations of TBP and their impact on transcription initiation
Nature Communications (2020)
-
Structure of the initiation-competent RNA polymerase I and its implication for transcription
Nature Communications (2016)
-
RNA polymerase I–Rrn3 complex at 4.8 Å resolution
Nature Communications (2016)
-
AF4 uses the SL1 components of RNAP1 machinery to initiate MLL fusion- and AEP-dependent transcription
Nature Communications (2015)
-
A strategy for dissecting the architectures of native macromolecular assemblies
Nature Methods (2015)