Abstract
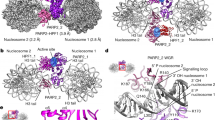
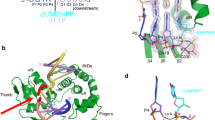
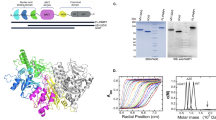
Poly(ADP-ribose) polymerase 1 (PARP1) is a primary DNA damage sensor whose (ADP-ribose) polymerase activity is acutely regulated by interaction with DNA breaks. Upon activation at sites of DNA damage, PARP1 modifies itself and other proteins by covalent addition of long, branched polymers of ADP-ribose, which in turn recruit downstream DNA repair and chromatin remodeling factors. PARP1 recognizes DNA damage through its N-terminal DNA-binding domain (DBD), which consists of a tandem repeat of an unusual zinc-finger (ZnF) domain. We have determined the crystal structure of the human PARP1-DBD bound to a DNA break. Along with functional analysis of PARP1 recruitment to sites of DNA damage in vivo, the structure reveals a dimeric assembly whereby ZnF1 and ZnF2 domains from separate PARP1 molecules form a strand-break recognition module that helps activate PARP1 by facilitating its dimerization and consequent trans-automodification.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Accession codes
Change history
10 July 2015
In the version of this article initially published, the image in the bottom row of Figure 4a (full-length PARP1-EGFP mutant R138E) was mistakenly replaced with a duplicate of the bottom-row image in Figure 4c (full-length PARP1-EGFP mutant M43D F44D) during preparation of the accepted version of the manuscript, after peer review and editorial evaluation had taken place. The error has been corrected in the HTML and PDF versions of the article.
References
Pion, E. et al. DNA-induced dimerization of poly(ADP-ribose) polymerase-1 triggers its activation. Biochemistry 44, 14670–14681 (2005).
Kirsten, E., Kun, E., Mendeleyev, J. & Ordahl, C.P. Activity assays for poly-ADP ribose polymerase. Methods Mol. Biol. 287, 137–149 (2004).
Huang, K. et al. Analysis of nucleotide sequence-dependent DNA binding of poly(ADP-ribose) polymerase in a purified system. Biochemistry 43, 217–223 (2004).
Shall, S. & de Murcia, G. Poly(ADP-ribose) polymerase-1: what have we learned from the deficient mouse model? Mutat. Res. DNA Repair 460, 1–15 (2000).
Amé, J.C., Spenlehauer, C. & de Murcia, G. The PARP superfamily. Bioessays 26, 882–893 (2004).
D'Amours, D., Desnoyers, S., D'Silva, I. & Poirier, G.G. Poly(ADP-ribosyl)ation reactions in the regulation of nuclear functions. Biochem. J. 342, 249–268 (1999).
Altmeyer, M., Messner, S., Hassa, P.O., Fey, M. & Hottiger, M.O. Molecular mechanism of poly(ADP-ribosyl)ation by PARP1 and identification of lysine residues as ADP-ribose acceptor sites. Nucleic Acids Res. 37, 3723–3738 (2009).
Fontan-Lozano, A. et al. Histone H1 poly[ADP]-ribosylation regulates the chromatin alterations required for learning consolidation. J. Neurosci. 30, 13305–13313 (2010).
Messner, S. et al. PARP1 ADP-ribosylates lysine residues of the core histone tails. Nucleic Acids Res. 38, 6350–6362 (2010).
Messner, S. & Hottiger, M.O. Histone ADP-ribosylation in DNA repair, replication and transcription. Trends Cell Biol. 21, 534–542 (2011).
Mathis, G. & Althaus, F.R. Release of core DNA from nucleosomal core particles following (ADP-ribose)N-modification in vitro. Biochem. Biophys. Res. Commun. 143, 1049–1054 (1987).
Althaus, F.R. Poly (ADP-ribose) and chromatin organization in DNA excision repair. Br. J. Cancer 56, 176–176 (1987).
Karras, G.I. et al. The macro domain is an ADP-ribose binding module. EMBO J. 24, 1911–1920 (2005).
Timinszky, G. et al. A macrodomain-containing histone rearranges chromatin upon sensing PARP1 activation. Nat. Struct. Mol. Biol. 16, 923–929 (2009).
Ahel, D. et al. Poly(ADP-ribose)-dependent regulation of DNA repair by the chromatin remodeling enzyme ALC1. Science 325, 1240–1243 (2009).
Gottschalk, A.J. et al. Poly(ADP-ribosyl)ation directs recruitment and activation of an ATP-dependent chromatin remodeler. Proc. Natl. Acad. Sci. USA 106, 13770–13774 (2009).
Ahel, I. et al. Poly(ADP-ribose)-binding zinc finger motifs in DNA repair/checkpoint proteins. Nature 451, 81–85 (2008).
Bryant, H.E. et al. Specific killing of BRCA2-deficient tumours with inhibitors of poly(ADP-ribose) polymerase. Nature 434, 913–917 (2005).
Farmer, H. et al. Targeting the DNA repair defect in BRCA mutant cells as a therapeutic strategy. Nature 434, 917–921 (2005).
Lilyestrom, W., van der Woerd, M.J., Clark, N. & Luger, K. Structural and biophysical studies of human PARP-1 in complex with damaged DNA. J. Mol. Biol. 395, 983–994 (2010).
Eustermann, S. et al. The DNA-binding domain of human PARP-1 interacts with DNA single-strand breaks as a monomer through its second zinc finger. J. Mol. Biol. 407, 149–170 (2011).
Langelier, M.F., Planck, J.L., Roy, S. & Pascal, J.M. Crystal structures of poly(ADP-ribose) polymerase-1 (PARP-1) zinc fingers bound to DNA: structural and functional insights into DNA-dependent PARP-1 activity. J. Biol. Chem. 286, 10690–10701 (2011).
Gradwohl, G. et al. The second zinc-finger domain of poly(ADP-ribose) polymerase determines specificity for single-stranded breaks in DNA. Proc. Natl. Acad. Sci. USA 87, 2990–2994 (1990).
Panzeter, P.L. & Althaus, F.R. DNA strand break-mediated partitioning of poly(ADP-ribose) polymerase function. Biochemistry 33, 9600–9605 (1994).
Kim, J.W., Kim, K., Kang, K. & Joe, C.O. Inhibition of homodimerization of poly(ADP-ribose) polymerase by its C-terminal cleavage products produced during apoptosis. J. Biol. Chem. 275, 8121–8125 (2000).
Mendoza-Alvarez, H. & Alvarez-Gonzalez, R. Poly(ADP-ribose) polymerase is a catalytic dimer and the automodification reaction is intermolecular. J. Biol. Chem. 268, 22575–22580 (1993).
Bauer, P.I., Buki, K.G., Hakam, A. & Kun, E. Macromolecular association of ADP-ribosyltransferase and its correlation with enzymic activity. Biochem. J. 270, 17–26 (1990).
Pion, E. et al. Poly(ADP-ribose) polymerase-1 dimerizes at a 5′ recessed DNA end in vitro: a fluorescence study. Biochemistry 42, 12409–12417 (2003).
Langelier, M.F., Ruhl, D.D., Planck, J.L., Kraus, W.L. & Pascal, J.M. The Zn3 domain of human poly(ADP-ribose) polymerase-1 (PARP-1) functions in both DNA-dependent poly(ADP-ribose) synthesis activity and chromatin compaction. J. Biol. Chem. 285, 18877–18887 (2010).
de Murcia, G. & Menissier de Murcia, J. Poly(ADP-ribose) polymerase: a molecular nick-sensor. Trends Biochem. Sci. 19, 172–176 (1994).
D'Silva, I. et al. Relative affinities of poly(ADP-ribose) polymerase and DNA- dependent protein kinase for DNA strand interruptions. Biochim. Biophys. Acta 1430, 119–126 (1999).
Potaman, V.N., Shlyakhtenko, L.S., Oussatcheva, E.A., Lyubchenko, Y.L. & Soldatenkov, V.A. Specific binding of poly(ADP-ribose) polymerase-1 to cruciform hairpins. J. Mol. Biol. 348, 609–615 (2005).
Lonskaya, I. et al. Regulation of poly(ADP-ribose) polymerase-1 by DNA structure-specific binding. J. Biol. Chem. 280, 17076–17083 (2005).
Bryant, H.E. et al. PARP is activated at stalled forks to mediate Mre11-dependent replication restart and recombination. EMBO J. 28, 2601–2615 (2009).
Ikejima, M. et al. The zinc fingers of human poly(ADP-ribose) polymerase are differentially required for the recognition of DNA breaks and nicks and the consequent enzyme activation. Other structures recognize intact DNA. J. Biol. Chem. 265, 21907–21913 (1990).
Kim, M.Y., Mauro, S., Gevry, N., Lis, J.T. & Kraus, W.L. NAD+-dependent modulation of chromatin structure and transcription by nucleosome binding properties of PARP-1. Cell 119, 803–814 (2004).
Tulin, A. & Spradling, A. Chromatin loosening by poly(ADP)-ribose polymerase (PARP) at Drosophila puff loci. Science 299, 560–562 (2003).
Ju, B.G. et al. Activating the PARP-1 sensor component of the groucho/ TLE1 corepressor complex mediates a CaMKinase IIδ-dependent neurogenic gene activation pathway. Cell 119, 815–829 (2004).
David, K.K., Andrabi, S.A., Dawson, T.M. & Dawson, V.L. Parthanatos, a messenger of death. Front. Biosci. 14, 1116–1128 (2009).
Kotova, E., Jarnik, M. & Tulin, A.V. Uncoupling of the transactivation and transrepression functions of PARP1 protein. Proc. Natl. Acad. Sci. USA 107, 6406–6411 (2010).
Audebert, M., Salles, B. & Calsou, P. Involvement of poly(ADP-ribose) polymerase-1 and XRCC1/DNA ligase III in an alternative route for DNA double-strand breaks rejoining. J. Biol. Chem. 279, 55117–55126 (2004).
Audebert, M., Salles, B., Weinfeld, M. & Calsou, P. Involvement of polynucleotide kinase in a poly(ADP-ribose) polymerase-1-dependent DNA double-strand breaks rejoining pathway. J. Mol. Biol. 356, 257–265 (2006).
Audebert, M., Salles, B. & Calsou, P. Effect of double-strand break DNA sequence on the PARP-1 NHEJ pathway. Biochem. Biophys. Res. Commun. 369, 982–988 (2008).
Cock, J.M., Vanoosthuyse, V. & Gaude, T. Receptor kinase signalling in plants and animals: distinct molecular systems with mechanistic similarities. Curr. Opin. Cell Biol. 14, 230–236 (2002).
Pellegrini, L. Role of heparan sulfate in fibroblast growth factor signalling: a structural view. Curr. Opin. Struct. Biol. 11, 629–634 (2001).
Ali, M.M. et al. Structure of the Ire1 autophosphorylation complex and implications for the unfolded protein response. EMBO J. 30, 894–905 (2011).
Leslie, A.G.W. MOSFLM Users Guide (MRC Laboratory of Molecular Biology, Cambridge, UK, 1995).
CCP4. Programs for protein crystallography. Acta Crystallogr. D Biol. Crystallogr. 50, 760–763 (1994).
McCoy, A.J. Solving structures of protein complexes by molecular replacement with Phaser. Acta Crystallogr. D Biol. Crystallogr. 63, 32–41 (2007).
Adams, P.D. et al. Recent developments in the PHENIX software for automated crystallographic structure determination. J. Synchrotron Radiat. 11, 53–55 (2004).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Chen, V.B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D Biol. Crystallogr. 66, 12–21 (2010).
Zufferey, R., Nagy, D., Mandel, R.J., Naldini, L. & Trono, D. Multiply attenuated lentiviral vector achieves efficient gene delivery in vivo. Nat. Biotechnol. 15, 871–875 (1997).
Hacham, H. & Ben-Ishai, R. Determination of poly(ADP-ribose) chain length distribution on polyacrylamide gels by silver staining. Anal. Biochem. 184, 83–85 (1990).
Malanga, M., Bachmann, S., Panzeter, P.L., Zweifel, B. & Althaus, F.R. Poly(ADP-ribose) quantification at the femtomole level in mammalian cells. Anal. Biochem. 228, 245–251 (1995).
Acknowledgements
We thank M. Roe for assistance with X-ray data collection and K. Caldecott for useful discussion. We are grateful to the Diamond Light Source Ltd., Didcot, UK, for access to synchrotron radiation. We thank the Advanced Light Microscopy Facility (ALMF) at the European Molecular Biology Laboratory (EMBL) and Olympus Europe for supporting EMBL's ALMF. This work was supported by EMBL, the Human Frontiers Science Program and the EU FP6 Marie Curie Research and Training Network “Chromatin Plasticity” to A.G.L. and a Cancer Research UK Programme Grant (C302/A8265) to L.H.P. L.H.P. is supported by a Wellcome Trust Senior Investigator Award.
Author information
Authors and Affiliations
Contributions
A.A.E.A. purified the protein, crystallized the complex and collected the X-ray diffraction data; G.T. designed and constructed the PARP1-EGFP constructs and performed the laser DNA-damage experiments; M.K. performed the FRAP experiments; P.O.H. engineered the knockdown PARP1 cell line and the wild-type imaging reporter constructs and performed the in vitro complementation assays; M.H. and R.A.-B. engineered and purified the mutant PARP1 constructs; A.G.L. designed the study and analyzed the data; L.H.P. designed the study, analyzed the data and wrote the paper; A.W.O. designed the study, made the baculovirus constructs, performed the EMSA experiments, designed the purification protocol and solved and refined the crystal structure. All authors discussed the results and commented on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–2 (PDF 960 kb)
Rights and permissions
About this article
Cite this article
Ali, A., Timinszky, G., Arribas-Bosacoma, R. et al. The zinc-finger domains of PARP1 cooperate to recognize DNA strand breaks. Nat Struct Mol Biol 19, 685–692 (2012). https://doi.org/10.1038/nsmb.2335
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.2335
This article is cited by
-
Comprehensive machine learning boosts structure-based virtual screening for PARP1 inhibitors
Journal of Cheminformatics (2024)
-
Biochemical Markers of Zinc Nutrition
Biological Trace Element Research (2024)
-
HPF1-dependent histone ADP-ribosylation triggers chromatin relaxation to promote the recruitment of repair factors at sites of DNA damage
Nature Structural & Molecular Biology (2023)
-
Structural dynamics of DNA strand break sensing by PARP-1 at a single-molecule level
Nature Communications (2022)
-
G4-quadruplex-binding proteins: review and insights into selectivity
Biophysical Reviews (2022)