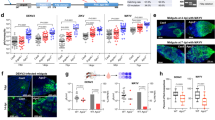
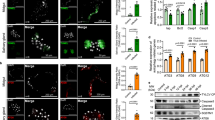
Abstract
Insect viruses have evolved strategies to control the host RNAi antiviral defense mechanism. In nature, Drosophila melanogaster C virus (DCV) infection causes low mortality and persistent infection, whereas the closely related cricket paralysis virus (CrPV) causes a lethal infection. We show that these viruses use different strategies to modulate the host RNAi defense machinery. The DCV RNAi suppressor (DCV-1A) binds to long double-stranded RNA and prevents processing by Dicer2. In contrast, the CrPV suppressor (CrPV-1A) interacts with the endonuclease Argonaute 2 (Ago2) and inhibits its activity without affecting the microRNA (miRNA)-Ago1–mediated silencing. We examined the link between viral RNAi suppressors and the outcome of infection using recombinant Sindbis viruses encoding either CrPV-1A or DCV-1A. Flies infected with Sindbis virus expressing CrPV-1A showed a marked increase in virus production, spread and mortality. In contrast, Sindbis pathogenesis was only modestly increased by expression of DCV- 1A. We conclude that RNAi suppressors function as virulence factors in insects and can target the Drosophila RNAi pathway at different points.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Siomi, H. & Siomi, M.C. On the road to reading the RNA-interference code. Nature 457, 396–404 (2009).
van Rij, R.P. & Andino, R. The silent treatment: RNAi as a defense against virus infection in mammals. Trends Biotechnol. 24, 186–193 (2006).
Forstemann, K., Horwich, M.D., Wee, L., Tomari, Y. & Zamore, P.D. Drosophila microRNAs are sorted into functionally distinct argonaute complexes after production by dicer-1. Cell 130, 287–297 (2007).
Tomari, Y., Du, T. & Zamore, P.D. Sorting of Drosophila small silencing RNAs. Cell 130, 299–308 (2007).
Ding, S.W. & Voinnet, O. Antiviral immunity directed by small RNAs. Cell 130, 413–426 (2007).
Baumberger, N., Tsai, C.H., Lie, M., Havecker, E. & Baulcombe, D.C. The Polerovirus silencing suppressor P0 targets ARGONAUTE proteins for degradation. Curr. Biol. 17, 1609–1614 (2007).
Bortolamiol, D., Pazhouhandeh, M., Marrocco, K., Genschik, P. & Ziegler-Graff, V. The Polerovirus F box protein P0 targets ARGONAUTE1 to suppress RNA silencing. Curr. Biol. 17, 1615–1621 (2007).
Pazhouhandeh, M. et al. F-box-like domain in the polerovirus protein P0 is required for silencing suppressor function. Proc. Natl. Acad. Sci. USA 103, 1994–1999 (2006).
Zhang, X. et al. Cucumber mosaic virus-encoded 2b suppressor inhibits Arabidopsis Argonaute1 cleavage activity to counter plant defense. Genes Dev. 20, 3255–3268 (2006).
Kasschau, K.D. et al. P1/HC-Pro, a viral suppressor of RNA silencing, interferes with Arabidopsis development and miRNA unction. Dev. Cell 4, 205–217 (2003).
Chao, J.A. et al. Dual modes of RNA-silencing suppression by flock house virus protein B2. Nat. Struct. Mol. Biol. 12, 952–957 (2005).
Gomariz-Zilber, E. & Thomas-Orillard, M. Drosophila C virus and Drosophila hosts: a good association in various environments. J. Evol. Biol. 6, 677–689 (1993).
Brun, P. & Plus, N. The viruses of Drosophila. in The Genetics and Biology of Drosophila Vol. 2, 625–702 (Academic Press, New York, 1980).
Jousset, F.-X. & Plus, N. Etude de la transmission horizontale et de la transmis- sion verticale de picornavirus de Drosophila melanogaster et de Drosophila immigrans. Ann. Microbiol. 126, 231–249 (1975).
Thomas, J.M., Klimstra, W.B., Ryman, K.D. & Heidner, H.W. Sindbis virus vectors designed to express a foreign protein as a cleavable component of the viral structural polyprotein. J. Virol. 77, 5598–5606 (2003).
Aravin, A.A. et al. The small RNA profile during Drosophila melanogaster development. Dev. Cell 5, 337–350 (2003).
Gomariz-Zilber, E., Poras, M. & Thomas-Orillard, M. Drosophila C virus: experimental study of infectious yields and underlying pathology in Drosophila melanogaster laboratory populations. J. Invertebr. Pathol. 65, 243–247 (1995).
Reinganum, C. The isolation of cricket paralysis virus from the emperor gum moth, Antheraea eucalypti Scott, and its infectivity towards a range of insect species. Intervirology 5, 97–102 (1975).
Manousis, T. & Moore, N.F. Cricket paralysis virus, a potential control agent for the olive fruit fly, Dacus oleae Gmel. Appl. Environ. Microbiol. 53, 142–148 (1987).
Wilson, J.E., Powell, M.J., Hoover, S.E. & Sarnow, P. Naturally occurring dicistronic cricket paralysis virus RNA is regulated by two internal ribosome entry sites. Mol. Cell. Biol. 20, 4990–4999 (2000).
Galiana-Arnoux, D., Dostert, C., Schneemann, A., Hoffmann, J.A. & Imler, J.L. Essential function in vivo for Dicer-2 in host defense against RNA viruses in Drosophila. Nat. Immunol. 7, 590–597 (2006).
van Rij, R.P. et al. The RNA silencing endonuclease Argonaute 2 mediates specific antiviral immunity in Drosophila melanogaster. Genes Dev. 20, 2985–2995 (2006).
Wang, X.H. et al. RNA interference directs innate immunity against viruses in adult Drosophila. Science 312, 452–454 (2006).
Hahn, H. & Palmenberg, A.C. Mutational analysis of the encephalomyocarditis virus primary cleavage. J. Virol. 70, 6870–6875 (1996).
Chapman, E.J., Prokhnevsky, A.I., Gopinath, K., Dolja, V.V. & Carrington, J.C. Viral RNA silencing suppressors inhibit the microRNA pathway at an intermediate step. Genes Dev. 18, 1179–1186 (2004).
Brennecke, J., Hipfner, D.R., Stark, A., Russell, R.B. & Cohen, S.M. bantam encodes a developmentally regulated microRNA that controls cell proliferation and regulates the proapoptotic gene hid in Drosophila. Cell 113, 25–36 (2003).
Roignant, J.Y. et al. Absence of transitive and systemic pathways allows cell-specific and isoform-specific RNAi in Drosophila. RNA 9, 299–308 (2003).
Czech, B. et al. An endogenous small interfering RNA pathway in Drosophila. Nature 453, 798–802 (2008).
Ghildiyal, M. et al. Endogenous siRNAs derived from transposons and mRNAs in Drosophila somatic cells. Science 320, 1077–1081 (2008).
Kawamura, Y. et al. Drosophila endogenous small RNAs bind to Argonaute 2 in somatic cells. Nature 453, 793–797 (2008).
Miyoshi, K., Tsukumo, H., Nagami, T., Siomi, H. & Siomi, M.C. Slicer function of Drosophila Argonautes and its involvement in RISC formation. Genes Dev. 19, 2837–2848 (2005).
Pham, J.W. & Sontheimer, E.J. Molecular requirements for RNA-induced silencing complex assembly in the Drosophila RNA interference pathway. J. Biol. Chem. 280, 39278–39283 (2005).
Sontheimer, E.J. Assembly and function of RNA silencing complexes. Nat. Rev. Mol. Cell Biol. 6, 127–138 (2005).
Haley, B., Tang, G. & Zamore, P.D. In vitro analysis of RNA interference in Drosophila melanogaster. Methods 30, 330–336 (2003).
Hammond, S.M., Bernstein, E., Beach, D. & Hannon, G.J. An RNA-directed nuclease mediates post-transcriptional gene silencing in Drosophila cells. Nature 404, 293–296 (2000).
Thomas-Orillard, M. & Legendre, S. C virus of Drosophila and dynamics of host population. C.R. Acad. Sci. III 319, 615–621 (1996).
Plus, N., Croizier, G., Jousset, F.X. & David, J. Picornaviruses of laboratory and wild Drosophila melanogaster: geographical distribution and serotypic composition. Ann. Microbiol. 126, 107–117 (1975).
Cullen, B.R. Is RNA interference involved in intrinsic antiviral immunity in mammals? Nat. Immunol. 7, 563–567 (2006).
Mudiganti, U., Hernandez, R., Ferreira, D. & Brown, D.T. Sindbis virus infection of two model insect cell systems—a comparative study. Virus Res. 122, 28–34 (2006).
Hannon, G.J. RNA interference. Nature 418, 244–251 (2002).
Voinnet, O. RNA silencing as a plant immune system against viruses. Trends Genet. 17, 449–459 (2001).
Zamore, P.D. RNA interference: listening to the sound of silence. Nat. Struct. Biol. 8, 746–750 (2001).
Merai, Z. et al. Double-stranded RNA binding may be a general plant RNA viral strategy to suppress RNA silencing. J. Virol. 80, 5747–5756 (2006).
Haley, B. & Zamore, P.D. Kinetic analysis of the RNAi enzyme complex. Nat. Struct. Mol. Biol. 11, 599–606 (2004).
Filipe, D. & Thomas-Orillard, M. Experimental study of a Drosophila melanogaster laboratory population infected by food contamination. Endocytobiosis and Cell Res. 12, 163–176 (1998).
Horwich, M.D. & Zamore, P.D. Design and delivery of antisense oligonucleotides to block microRNA function in cultured Drosophila and human cells. Nat. Protoc. 3, 1537–1549 (2008).
Blethrow, J.D., Tang, C., Deng, C. & Krutchinsky, A.N. Modular mass spectrometric tool for analysis of composition and phosphorylation of protein complexes. PLoS One 2, e358 (2007).
Harper, S.M., Neil, L.C. & Gardner, K.H. Structural basis of a phototropin light switch. Science 301, 1541–1544 (2003).
Ephrussi, B. & Herold, J.L. Studies of eye pigments of Drosophila. I. Methods of extraction and quantitative estimation of the pigment components. Genetics 29, 148–175 (1944).
Berry, B., Deddouche, S., Kirschner, D., Imler, J.L. & Antoniewski, C. Viral suppressors of RNA silencing hinder exogenous and endogenous small RNA pathways in Drosophila. PLoS One 4, e5866 (2009).
Acknowledgements
We thank J. Frydman and Andino laboratory members L. Gitlin, D. Barnes, M. Flenniken and A. Lauring for useful discussion for preparing the manuscript, R. van Rij and C. Saleh for their help, G. Belsham for critical reading of the manuscript and useful comments, P. Zamore (Univ. Massachusetts Medical School), Mikiko Siomi and Keita Miyoshi (Keio Univ. School of Medicine, Japan) for providing antibody directed against Drosophila Argonaute 2, H. Heidner (Univ. Texas, San Antonio) for providing Sindbis virus plasmid and M. Moritz (Univ. California, San Francisco) for Drosophila embryo extract preparation. This work was financially supported by Pasteur Institute, by the Centre National de la Recherche Scientifique, by grants from the Agence Nationale de la Recherche (AKROSS) and by fellowships from the Lebanese Centre National de la Recherche Scientifique to B.B. as well as by US National Institutes of Health grants AI40085 and AI064738 to R.A.
Author information
Authors and Affiliations
Contributions
A.N. and R.A. designed and interpreted most of the experiments; A.N. performed most of the experiments; B.B. and C.A. designed and performed the IR[white], GFP/bantam sensor and retrotransposon experiments in flies; M.T. and M.K. performed the fly injections; A.A. made some plasmid constructs; C.D. and A.K. carried out the MS analysis; J.G. advised on the project; A.N., R.A., A.A. and C.A. prepared the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–5 (PDF 501 kb)
Supplementary Video 1
Serial optical section of Drosophila embryos expressing GFP under Bantam miRNA control in the presence or absence of CrPV-1A suppressor. (MOV 6775 kb)
Supplementary Video 2
Serial Optical Sections 2, Bantam expressionCrPV148 (MOV 6835 kb)
Rights and permissions
About this article
Cite this article
Nayak, A., Berry, B., Tassetto, M. et al. Cricket paralysis virus antagonizes Argonaute 2 to modulate antiviral defense in Drosophila. Nat Struct Mol Biol 17, 547–554 (2010). https://doi.org/10.1038/nsmb.1810
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.1810
This article is cited by
-
Role of the receptor for activated C kinase 1 during viral infection
Archives of Virology (2022)
-
Establishing RNAi for basic research and pest control and identification of the most efficient target genes for pest control: a brief guide
Frontiers in Zoology (2021)
-
Innate and adaptive resistance to RNAi: a major challenge and hurdle to the development of double stranded RNA-based pesticides
3 Biotech (2021)
-
Role of microRNAs in antiviral responses to dengue infection
Journal of Biomedical Science (2020)
-
Isolation and characterization of a novel cripavirus, the first Dicistroviridae family member infecting the cotton mealybug Phenacoccus solenopsis
Archives of Virology (2020)