Key Points
-
Translation factors and translational control mechanisms are downstream targets of several signalling pathways and are crucial in synaptic plasticity. Some forms of translational control alter general protein synthesis, whereas others regulate translation of specific messenger RNAs (mRNAs).
-
Translation initiation refers to the assembly of a translation-competent ribosome at the AUG start codon on an mRNA. The first step involves the binding of the translation-initiation factor eIF2, which is a G protein, to methionyl-transfer RNA in a GTP-dependent manner.
-
eIF2 has three subunits (α, β and γ), and the conversion of inactive eIF2·GDP to active eIF2·GTP by eIF2B is blocked by phosphorylation of eIF2α. Four kinases that are present in the brain — PKR, HRI, PERK and GCN2 — phosphorylate eIF2α on Ser51.
-
The eIF2B enzyme complex consists of five polypeptides (α–ε), with eIF2Bε catalysing guanine nucleotide exchange on eIF2. The importance of eIF2B function in the brain is highlighted by the fact that mutations in each eIF2B subunit can cause leukoencephalopathy with vanishing white matter.
-
The integrity of the eIF4F cap-binding complex and, therefore, translation efficiency, is regulated by 4E-BPs. Phosphorylation of 4E-BPs by the extracellular signal-regulated kinase (ERK), phosphoinositide 3-kinase (PI3K) and mammalian target of rapamycin (mTOR) signalling pathways is crucial for protein synthesis-dependent synaptic plasticity and memory.
-
The cap-binding protein eIF4E is phosphorylated by the protein kinases Mnk1 and Mnk2, which are phosphorylated and activated by ERK and p38. eIF4E phosphorylation is associated with synaptic plasticity and memory.
-
In addition to regulating 4E-BP phosphorylation and function, mTOR directly phosphorylates and activates the S6 kinase (S6K), which phosphorylates the ribosomal protein S6, an essential component of the small, 40S ribosomal subunit. S6K and S6 phosphorylation have been implicated in synaptic plasticity and memory.
-
The cytoplasmic polyadenylation element (CPE) in the 3′-untranslated region is important in extension of the poly(A) tail and translation activation. Recent evidence indicates a crucial role for CPE-binding protein in long-term facilitation in Aplysia californica and in hippocampal synaptic plasticity.
Abstract
Changes in gene expression are required for long-lasting synaptic plasticity and long-term memory in both invertebrates and vertebrates. Regulation of local protein synthesis allows synapses to control synaptic strength independently of messenger RNA synthesis in the cell body. Recent reports indicate that several biochemical signalling cascades couple neurotransmitter and neurotrophin receptors to translational regulatory factors in protein synthesis-dependent forms of synaptic plasticity and memory. In this review, we highlight these translational regulatory mechanisms and the signalling pathways that govern the expression of synaptic plasticity in response to specific types of neuronal stimulation.
Similar content being viewed by others
Main
The consolidation and storage of long-term memories, but not short-term memories, require macromolecular synthesis. At the cellular level, the synthesis of new macromolecules is required for long-lasting forms of synaptic plasticity, including LONG-TERM FACILITATION (LTF) in sensory neurons of the mollusc Aplysia californica and a type of LONG-TERM POTENTIATION (LTP) known as LATE-PHASE LTP (L-LTP) in hippocampal neurons (see Ref. 1 for a review). By contrast, shorter-lasting forms of synaptic plasticity, including SHORT-TERM FACILITATION (STF) in Aplysia and EARLY-PHASE LTP (E-LTP), require post-translational modifications of existing proteins, but not the synthesis of new macromolecules. Although significant progress has been made in understanding transcriptional regulation in long-lasting synaptic plasticity and long-term memory, this is not the only biochemical mechanism that controls gene expression. The regulation of messenger RNA (mRNA) transport, mRNA stability and protein synthesis also contributes to the control of gene expression. The regulation of protein synthesis is particularly intriguing because it allows cells to rapidly modulate the production of proteins without involving new transcription or mRNA transport. To understand the regulation of gene expression during long-lasting forms of synaptic plasticity and long-term memory, it is important to examine the regulation of mRNA translation. The reader is directed to other recent reviews for details about the regulation of transcription and mRNA transport and localization in neurons during periods of neuronal activity2,3.
Dendrites and dendritic spines contain polyribosomes, translation factors and mRNA2,4,5,6,7,8, indicating that mRNA is translated in synaptic locations. Consistent with this idea, dendrites can translate mRNA9, and new protein synthesis is induced in dendrites after exposure of cultured neurons to either brain-derived neurotrophic factor (BDNF)10 or activity blockade11,12. In addition, membrane depolarization, BDNF and glutamate receptor agonists induce the synthesis of new proteins in isolated dendrites and synaptoneurosomes13,14,15. There is also evidence that local protein synthesis is required for synaptic plasticity. In hippocampal area CA1, both BDNF-induced potentiation16 and metabotropic glutamate receptor (mGluR)-dependent LONG-TERM DEPRESSION (LTD)17 are blocked by protein-synthesis inhibitors, even when the pre- and postsynaptic pyramidal cell somas are excised from their dendrites. Presynaptic local protein synthesis is required for long-term facilitation (LTF) in Aplysia sensory neurons18 and for activity-dependent synaptic plasticity in Xenopus laevis nerve–muscle cultures19. Furthermore, LTP-inducing stimulation causes polyribosomes to move from dendritic shafts to dendritic spines with enlarged synapses20, which indicates that local protein synthesis might be involved in morphological changes that are associated with LTP. These findings indicate that protein synthesis is triggered at synaptic locations and that local translation is required for several forms of synaptic plasticity. However, until recently, little was known about the biochemical signalling mechanisms that couple neurotransmitter and neurotrophin receptors to translational regulation in synaptic plasticity and memory. In this review, we describe several biochemical mechanisms that regulate translation initiation and discuss examples of this type of regulation in synaptic plasticity and memory.
Regulation of translation initiation
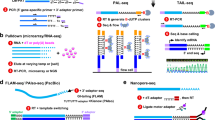
Protein synthesis is divided into three steps: initiation, elongation and termination. Initiation refers to the assembly of a translation-competent ribosome at the AUG start codon on an mRNA; translation elongation is the codon-dependent assembly of a polypeptide; and termination involves release of the completed protein when the ribosome reaches a termination codon. These three steps are facilitated by trans-acting proteins that are referred to, respectively, as eukaryotic initiation factors (eIF) (Box 1), elongation factors (eEF) and release factors (eRF). The initiation phase of protein synthesis (Fig. 1) can be further divided into three steps: first, the specific initiator methionyl–transfer RNA (Met–tRNAiMet) binds to the small (40S) ribosomal subunit to form a 43S pre-initiation complex; second, the 43S complex binds to an mRNA to form a 48S pre-initiation complex that scans the AUG start codon; and third, the large (60S) ribosomal subunit joins this complex to form an 80S ribosome. As might be expected for a complex biochemical process, the initiation phase of translation is a common target for regulation.
A binary complex of eukaryotic translation initiation factor 2 (eIF2) and GTP binds to methionyl–transfer RNA (Met–tRNAiMet), and the ternary complex associates with the 40S ribosomal subunit. The association of additional factors, such as eIF3 and eIF1A (1A), with the 40S subunit promotes ternary complex binding and generates a 43S pre-initiation complex. The cap-binding complex, which consists of eIF4E (4E), eIF4G and eIF4A (4A), binds to the 7-methyl-GTP (m7GTP) cap structure at the 5′ end of a messenger RNA (mRNA). eIF4G also binds to the poly(A)-binding protein (PABP), thereby bridging the 5′ and 3′ ends of the mRNA. This mRNA circularization and the ATP-dependent helicase activity of eIF4A are thought to promote the binding of the 43S pre-initiation complex to the mRNA, which produces a 48S pre-initiation complex. Following scanning of the ribosome to the AUG start codon, GTP is hydrolysed by eIF2, which triggers the dissociation of factors from the 48S complex and allows the eIF5B- and GTP-dependent binding of the large, 60S ribosomal subunit. Although the precise timing and requirements for the release of factors from the pre-initiation complexes are not clear, the 80S product of the pathway is competent for translation elongation and protein synthesis.
External cues are transduced to regulate protein synthesis by several diverse mechanisms, including post-translational modification of translation factors or binding of regulatory proteins to the 5′- or 3′-untranslated regions (UTRs) of specific mRNAs. To help to understand translational regulatory strategies, it is useful to review the translation pathway and the roles of the translation factors21,22. The translation factor eIF2, which is a G protein, binds to Met–tRNAiMet in a GTP-dependent manner. This ternary complex associates with the 40S subunit and other initiation factors, including eIF1, eIF1A, eIF3 and possibly eIF5, to form the 43S pre-initiation complex. The binding of the 43S complex to an mRNA is facilitated by the eIF4 family of translation factors. In addition to the AUG start codon, the 5′-7-methyl-GTP (m7GTP) cap and the 3′-poly(A) tail on mRNAs are also important in translation initiation. The cap-binding complex eIF4F consists of the m7GTP cap-binding protein eIF4E, the DEAD-box RNA helicase eIF4A and the scaffolding protein eIF4G. Interestingly, eIF4G also binds to the poly(A)-binding protein (PABP) and the 43S component eIF3. Through its interactions with both eIF4E (binding to the mRNA cap) and PABP (binding to the mRNA poly(A) tail), eIF4G effectively circularizes mRNAs. This mRNA circularization might similarly enhance eIF4F binding and 48S-complex formation, providing a mechanism for the synergistic enhancement of translation by these modifications23,24. The eIF4A component of eIF4F, with the initiation factor eIF4B, is thought to unwind secondary structures at the 5′ end of mRNAs. These mRNA-remodelling activities, in combination with the eIF4G–eIF3 interaction, are thought to promote the binding of the 43S complex to the mRNA, which forms the 48S pre-initiation complex.
After binding close to the 5′ end of the mRNA, the ribosomal complex migrates or scans down the mRNA and usually stops at the first AUG codon it encounters. Stable secondary RNA structures in the 5′ UTR or proteins that are bound to the 5′ UTR impede ribosome binding and scanning, thereby inhibiting translation23,25,26. eIF4A or other helicase activities might remodel the 5′ UTR to facilitate ribosomal binding and scanning. Recognition of the start codon triggers hydrolysis of GTP by eIF2, which leads to the release of factors from the 48S complex and allows the eIF5B- and GTP-dependent joining of the large ribosomal subunit. eIF2, which is now bound to GDP, is then released from the 48S complex. Similar to many G proteins, eIF2 has a higher affinity for GDP than for GTP, and the guanine-nucleotide exchange factor (GEF) eIF2B catalyses the exchange of GTP for GDP on eIF2, thereby recycling eIF2 for further rounds of translation initiation.
Translation factors and translational control mechanisms are downstream targets of several signalling pathways and are crucial during development and cellular stress responses. Many forms of translational control are homeostatic responses that alter general protein synthesis. However, reduced nutrient availability, oxidative stress, viral infection and misfolded proteins trigger inhibition of general protein synthesis, but stimulate translation of specific mRNAs. This gene-specific translational control depends on regulatory elements in the mRNA, such as upstream open reading frames (uORFs), RNA secondary structures or regulatory protein-binding sites22. So, mRNA specificity in translational control can be achieved through mRNA-specific binding proteins or through alterations to the general translational machinery (Box 2). Downregulation of general protein synthesis has been linked to two forms of mRNA-specific translational control: first, mRNAs that compete poorly for initiation factors and ribosomes are hypersensitive to small reductions in translational activity and their translation is preferentially inhibited; and second, translation of a specific class of mRNAs, including the uORF-containing GENERAL CONTROL NON-DEREPRESSIBLE 4 (GCN4) mRNA in yeast and the ACTIVATING TRANSCRIPTION FACTOR 4 (ATF4) mRNA in mammalian cells, is paradoxically increased when general translation is inhibited22. In the following sections, we review several mechanisms of translational control and evidence that they are linked to synaptic plasticity.
Phosphorylation of eIF2α
The conversion of inactive eIF2·GDP to active eIF2·GTP by eIF2B is regulated by phosphorylation (Fig. 2). eIF2 has three subunits (α, β and γ), and phosphorylation of eIF2α on serine at residue 51 (Ser51) converts eIF2 from a substrate to a competitive inhibitor of eIF2B27. Phosphorylation of eIF2α does not inhibit the general function of eIF2 — to deliver Met–tRNAiMet to the 40S subunit — but renders the protein defective in recycling. Most cells express more eIF2 than eIF2B, and phosphorylation of a fraction of the eIF2 is sufficient to inhibit eIF2B and block protein synthesis. The relative abundance of eIF2 and eIF2B in the nervous system has not been reported.
The eukaryotic translation initiation factor 2 (eIF2)–GTP binary complex binds to methionyl–transfer RNA (Met–tRNAiMet) and forms a ternary complex that then associates with the 40S ribosomal subunit. After start-codon recognition, GTP is hydrolysed by eIF2 and the binary eIF2–GDP complex is then released. The guanine nucleotide-exchange factor (GEF) eIF2B converts inactive eIF2–GDP to active eIF2–GTP, a process that is inhibited by phosphorylation (P) of the α-subunit of eIF2 on serine 51 by one of the four known eIF2α kinases. Phosphorylation of eIF2α converts eIF2 to a competitive inhibitor of eIF2B, and inhibition of eIF2B results in lowered levels of ternary complexes, which reduces general translation but increases translation of a specific class of messenger RNAs (mRNAs) with upstream open reading frames (uORFs)22. ATF4, activating transcription factor 4; C/EBP, CCAAT/enhancer-binding protein; GCN, general control non-derepressible; HRI, haem-regulated initiation factor 2α kinase; m7G, 7-methyl-GTP; PERK, eIF2α kinase 3; PKR, protein kinase-RNA regulated, interferon-inducible double-stranded RNA dependent.
Four kinases — PKR (protein kinase-RNA regulated), HRI (haem-regulated initiation factor 2α kinase), PERK (eIF2α kinase 3) and GCN2 (general control non-derepressible 2) — phosphorylate eIF2α on Ser51 (Table 1). PKR is activated by double-stranded RNA (dsRNA), HRI by low haem levels, PERK by endoplasmic reticulum (ER) stress and unfolded proteins in the ER, and GCN2 by amino-acid limitation. All four kinases are present in the brain. GCN2 mRNA is abundant in the mouse brain28,29, consistent with the restriction of GCN2 to the CNS during Drosophila melanogaster embryogenesis30. Phosphorylation of eIF2α is observed in the brain and increases significantly in neurons on reperfusion after ischaemia31. This eIF2α phosphorylation is probably mediated by PERK32, which is consistent with the finding that downregulation of eIF2α protects neurons during oxidative stress, perhaps by enhancing glutathione synthesis33. Surprisingly, mice that lack any one of the eIF2α kinases have only limited phenotypes, which indicates that the kinases perform redundant functions by responding to overlapping signals. PKR-knockout mice show elevated sensitivity to some viruses34,35; HRI-knockout mice have dysregulated globin protein synthesis in red blood cells36; PERK-knockout mice develop diabetes37,38, which is consistent with the ability of PERK mutations to cause insulin-dependent diabetes in patients with Wolcott–Rallison syndrome39; and GCN2-knockout mice fail to coordinate protein synthesis during amino-acid starvation and show increased lethality when they are deprived of the essential amino acid leucine40,41. Intriguingly, GCN2-knockout mice also have impaired regulation of phosphorylation of eIF4E-binding proteins (4E-BPs) and S6 kinase 1 (S6K1) (Ref. 41) — two other regulators of protein synthesis that will be discussed later.
Although the eIF2α kinases are generally described as stress-responsive regulators of general protein synthesis, activation of GCN2 in yeast by amino-acid starvation specifically stimulates the expression of GCN4 through regulated re-initiation at uORFs in the GCN4 mRNA42. Similarly, in mammalian cells, eIF2α phosphorylation stimulates translation of the mRNA that encodes the transcription factor ATF4 (Refs 43,44). So, eIF2α phosphorylation regulates both general and gene-specific protein synthesis. Phosphorylation of eIF2α during synaptic plasticity and memory has not been directly investigated, but treatment of cultured neurons with BDNF, which induces lasting plasticity, decreases eIF2α phosphorylation and enhances protein synthesis45. In addition, GCN2-knockout mice have altered synaptic plasticity and memory (N. Sonenberg, personal communication). These findings indicate that proper regulation of eIF2α phosphorylation is probably required for normal synaptic plasticity and memory.
Regulation by eIF2B
eIF2 function is modulated by the regulation of eIF2B, as well as being regulated by phosphorylation of eIF2α. The eIF2B complex consists of five polypeptides (α–ε), with eIF2Bε catalysing guanine nucleotide exchange on eIF2 (Ref. 27). The importance of eIF2B function in the brain is highlighted by the fact that mutations in each eIF2B subunit can cause leukoencephalopathy with vanishing white matter46. Phosphorylation of eIF2Bε by glycogen synthase kinase 3β (GSK3β) in vitro inhibits its activity47, and eIF2Bε is dephosphorylated in vivo after inactivation of GSK3β, which leads to increased eIF2B activity47,48. Although regulation of GSK3β has not been reported to be important in synaptic plasticity, treatment of neuronal cultures with BDNF results in decreased GSK3β activity due to increased phosphorylation, and this is correlated with an increase in eIF2B activity45. GSK3β also seems to be important in learning and memory. Mice that overexpress GSK3β show spatial learning deficits in the Morris water maze49 that have been attributed to the downstream effects of tau hyperphosphorylation. However, the learning deficits could also result from enhanced phosphorylation and inhibition of eIF2B, and the accompanying decrease in the levels of active, GTP-bound eIF2. Further studies are necessary to determine whether GSK3β regulates eIF2 activity during either synaptic plasticity or learning and memory.
Regulation by 4E-BPs
The integrity of the eIF4F cap-binding complex is modulated by 4E-BPs24. When not bound to 4E-BP, eIF4E readily associates with eIF4G to form the eIF4F complex and promote translation initiation. The 4E-BPs block eIF4F formation and inhibit protein synthesis by competing with eIF4G for binding to eIF4E. The binding of 4E-BPs to eIF4E is regulated by phosphorylation24: hypophosphorylated 4E-BPs bind to eIF4E and inhibit translation, whereas multi-site phosphorylation of 4E-BPs prevents their binding to eIF4E and allows eIF4F formation. Three 4E-BPs have been identified in mammals: 4E-BP1 is most prominent in adipose tissues and the pancreas, and 4E-BP1-knockout mice have less white adipose tissue50; 4E-BP3 is most abundant in the liver; and 4E-BP2 is the most abundant isoform in the brain, which contains little or no 4E-BP1 and 4E-BP3 (Ref. 50). Phosphorylation of 4E-BPs occurs in an ordered, hierarchical fashion. Threonine residues 37 and 46 (Thr37 and Thr46) are phosphorylated first, and this primes phosphorylation of Thr70 and Ser65 (Ref. 51). Phosphorylation of all these residues is required to block eIF4E binding. 4E-BP phosphorylation is regulated by the extracellular signal-regulated kinase (ERK), phosphoinositide 3-kinase (PI3K) and mammalian target of rapamycin (mTOR) signalling pathways24. Although phosphorylation of 4E-BP on the regulatory sites is sensitive to inhibitors of ERK, PI3K and mTOR, the identity of the kinase that phosphorylates each site has not been firmly established.
ERK is required for most forms of synaptic plasticity and memory52,53. Similarly, PI3K is required for several protein synthesis-dependent forms of synaptic plasticity and memory54,55,56,57. Rapamycin, an inhibitor of mTOR that decreases 4E-BP phosphorylation, inhibits synaptic plasticity in invertebrates, including LTF in Aplysia sensory neurons58 and at the crayfish neuromuscular junction55. In the hippocampus, mGluR-induced LTD59, mGluR-dependent depotentiation of LTP60, insulin-induced LTD61, BDNF-induced potentiation7 and L-LTP7 are all blocked by rapamycin. So, several forms of synaptic plasticity and memory require ERK, PI3K, mTOR and, presumably, phosphorylation of 4E-BP (Fig. 3). Consistent with this idea, emerging evidence indicates that proper regulation of 4E-BP is required for normal synaptic plasticity and memory. Treatment of hippocampal cultures with BDNF increases the phosphorylation of 4E-BP62, L-LTP-inducing stimulation is associated with increased phosphorylation of 4E-BP and an increase in eIF4F-complex formation, and 4E-BP2-knockout mice have altered synaptic plasticity and memory (J. Banko and E.K., unpublished observations). Therefore, proper regulation of 4E-BP by PI3K, ERK and mTOR is likely to be crucial for protein synthesis-dependent synaptic plasticity and memory.
Activation of group I metabotropic glutamate receptors (mGluRs) and NMDA (N-methyl-D-asparate) receptors (NMDARs) activate the mitogen-activated protein kinase kinase (MEK)/extracellular signal-regulated kinase (ERK) and phosphatidylinositol 3-kinase (PI3K) signalling pathways. Sequential activation of PI3K, phosphoinositide-dependent kinase 1 or 2 (PDK1/2), Akt and mammalian target of rapamycin (mTOR) results in activation of S6 kinase 1 (S6K1). Phosphorylation (P) of eIF4E-binding proteins (4E-BPs) by mTOR and other kinases induces the release of 4E-BP from eukaryotic translation initiation factor 4E (eIF4E) and results in the association of eIF4E with eIF4G and the formation of the active eIF4F (eIF4E·eIF4A·eIF4G) complex. eIF4F promotes messenger RNA (mRNA) binding to the 43S pre-initiation complex to form the 48S initiation complex. The eIF4F complex and the poly(A) tail act synergistically to stimulate mRNA translation. The ERK-dependent phosphorylation of both MAPK-interacting serine/threonine kinase 1 (Mnk1), which can phosphorylate eIF4E, and S6K1, which can phosphorylate ribosomal protein S6, is correlated with enhanced translation initiation. Of note, the signal transduction cascades depicted here are also activated by brain-derived neurotrophic factor (BDNF) in the hippocampus and cultured cortical neurons, and by serotonin in Aplysia californica sensory neurons. m7G, 7-methyl-GTP; PIP2, phosphatidylinositol-4,5-bisphosphate; PIP3, phosphatidylinositol-3,4,5-trisphosphate.
Phosphorylation of eIF4E
In addition to the regulation of eIF4F assembly by 4E-BP phosphorylation, the cap-binding protein eIF4E is a target for direct phosphorylation. Phosphorylation of eIF4E is stimulated by the ERK and p38 mitogen-activated protein kinase pathways, and correlates with increased translation rates in serum-stimulated cells63. The protein kinases MAPK-interacting serine/threonine kinase 1 and 2 (Mnk1 and Mnk2), which are phosphorylated and activated by ERK and p38, phosphorylate eIF4E on Ser209 (Refs 64–66). In mice that are deficient in both Mnk1 and Mnk2, phosphorylation of eIF4E is abolished67. Interestingly, both Mnk1 and Mnk2 bind to the C terminus of eIF4G, and phosphorylation of eIF4E by Mnk1 or Mnk2 seems to depend on the binding of both proteins to the scaffold protein eIF4G24,64. So, eIF4E phosphorylation by Mnk1 or Mnk2 is an indirect measure of eIF4F assembly. Phosphorylation of eIF4E reduces its cap-binding affinity68,69, which indicates that phosphorylation promotes eIF4E recycling after the ribosome has bound to an mRNA63. According to this model, dephosphorylated eIF4E binds to an mRNA cap, which then recruits eIF4F and the ribosome. Subsequent phosphorylation of eIF4E releases it from the cap and allows the ribosome to begin scanning. Despite the compelling nature of this model and the correlation of eIF4E phosphorylation with increased translation rates, mice that lack both Mnk1 and Mnk2 are viable and apparently normal67. However, synaptic plasticity and memory have not been analysed in these mice. In addition, and at odds with the lack of phenotype in the Mnk1/Mnk2-knockout mice, Drosophila that express a non-phosphorylatable eIF4E–S251A mutant, which corresponds to a Ser209 mutant in mammalian eIF4E, show impaired viability and growth, indicating that eIF4E phosphorylation is important for normal development70.
eIF4E phosphorylation might be important for synaptic plasticity and memory (Fig. 3). For example, Aplysia neurons treated with serotonin, which is required for LTF, have a p38-dependent increase in eIF4E phosphorylation71. In addition, ERK-dependent increases in the phosphorylation of eIF4E are induced in cultured neurons that are treated with BDNF62 and in the CA1 region of hippocampal slices treated with NMDA (N-methyl-D-aspartate)72. More recently, using dominant-negative MAPK/ERK kinase (MEK) mice that have decreased ERK activity, Kelleher and colleagues showed that L-LTP is associated with an ERK-dependent increase in the phosphorylation of eIF4E62. In further studies with the dominant-negative MEK mice, this same group showed that an ERK-dependent increase in the phosphorylation of eIF4E accompanies ASSOCIATIVE FEAR CONDITIONING, indicating that phosphorylation of eIF4E might also be involved in hippocampus-dependent memory formation62. Although there is no direct evidence that eIF4E phosphorylation is required for protein synthesis-dependent synaptic plasticity, these studies provide intriguing correlative data that link phosphorylation of eIF4E by Mnk1 through activation of either ERK or p38 to the regulation of translation initiation in synaptic plasticity and memory.
S6 and the S6 kinases
In addition to regulating 4E-BP phosphorylation and function, mTOR activation has been linked to translation elongation (Box 3) and mTOR directly phosphorylates and activates the p70 S6K73. S6K is also activated through the PI3K, phosphoinositide-dependent kinase 1 (PDK1) and ERK pathways74 (Fig. 3). S6K phosphorylates the ribosomal protein S6, an essential component of the small, 40S ribosomal subunit. The role of S6 and S6 phosphorylation in translational regulation is not well understood. S6 is located close to the mRNA- and tRNA-binding sites on the 40S subunit, and conditional deletion of S6 in the mouse liver impairs ribosome biogenesis and cell proliferation but not cell growth75. The role of S6 phosphorylation has been difficult to study owing to the presence of two S6Ks (S6K1 and S6K2) and additional kinases in the ERK pathway that phosphorylate S6 (Ref. 76). In general, S6 phosphorylation is enhanced in growth factor- and nutrient-stimulated cells and correlates with increased levels of translation73,74. The S6Ks and phosphorylation of S6 were previously implicated in the translational regulation of a specific class of mRNAs that encode components of the translational machinery, including eEF2, eEF1A, PABP and S6 (Ref. 77). These TERMINAL OLIGOPYRIMIDINE (TOP) mRNAs contain polypyrimidine tracts at the 5′ end, and their translation is upregulated by many of the conditions that stimulate S6 phosphorylation77. However, in cells that are deficient in both S6K1 and S6K2, the regulation of TOP mRNA translation is maintained76 and S6 is still phosphorylated, leaving open the possibility that S6 phosphorylation stimulates TOP mRNA translation76. Another substrate of the S6Ks is the translation initiation factor eIF4B78, a stimulator of the helicase eIF4A and of mRNA binding during translation initiation. It is not clear how, or whether, S6K phosphorylation modulates eIF4B function in protein synthesis.
Although the roles of S6K and S6 phosphorylation in translational regulation are not clear, the kinases have been implicated in synaptic plasticity and memory. For example, Aplysia synaptosomes treated with serotonin — a treatment that results in LTF in Aplysia neurons — show an increase in S6K1 activity, S6 phosphorylation and in levels of S6 (Ref. 79) and eEF2 (Ref. 80). In the CA1 region of the hippocampus, L-LTP-inducing stimulation results in an mTOR-dependent increase in S6K phosphorylation81. In addition, injection of the mouse hippocampus with a synthetic phospho-peptide to stimulate PI3K results in increased S6K phosphorylation and improved hippocampus-dependent memory82. Both L-LTP and associative fear conditioning are associated with an ERK-dependent increase in S6 phosphorylation in CA1 (Ref. 62). These findings indicate that the regulation of S6K and S6 is important for synaptic plasticity and memory. It will be of interest to determine whether the protein synthesis-dependent forms of synaptic plasticity and memory require an increase in the synthesis of ribosomal proteins and other components of the translation apparatus.
Regulation by the CPE-binding protein
In Xenopus oocytes, translation of a specific class of maternally inherited mRNAs is impaired, partly owing to a short poly(A) tail83. Translation of these dormant mRNAs is activated during the maturation of the oocyte or during early embryogenesis. The activation requires extension of the poly(A) tail, which depends on two RNA sequence elements in the 3′ UTR: the cytoplasmic polyadenylation element (CPE), a U-rich sequence that serves as a binding site for the CPE-binding protein (CPEB), and the sequence 5′-AAUAAA-3′, which binds to the cleavage and polyadenylation specificity factor (CPSF). In the oocyte, where translation of CPE-containing mRNAs is repressed (masked), CPEB binds to the 3′ UTR and recruits the protein Maskin83 (Fig. 4). Maskin contains an eIF4E-binding site and might act as a 4E-BP, blocking eIF4F formation (or binding) on the masked mRNA and thereby preventing its translation84. In this way, the CPEB–Maskin–eIF4E interaction could circularize a target mRNA and keep it in a translationally dormant state owing to its short poly(A) tail and impaired eIF4F binding (Fig. 4).
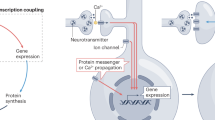
The 3′-untranslated region of a specific class of messenger RNA (mRNA) contains sequences that allow the binding of cytoplasmic polyadenylation element-binding protein (CPEB) and polyadenylation specificity factor (CPSF). Translation of transcripts that are bound to CPEB and its binding partner Maskin is inhibited, but can be de-repressed by extension of the poly (A) tail. NMDA (N-methyl-D-aspartate) receptor (NMDAR) activation results in calcium influx and activation of Aurora and/or calcium/calmodulin-dependent protein kinase II (CaMKII), which then phosphorylates (P) CPEB. This leads to the interaction between CPEB and CPSF and the subsequent recruitment of poly(A) polymerase (PAP) to lengthen the poly(A) tail. Poly(A)-binding protein (PABP) then binds to the extended poly(A) tail, interacts with eIF4G to circularize the mRNA, which releases eIF4E from Maskin, and results in enhanced translation of the CPE-containing mRNAs. m7G, 7-methyl-GTP; uORF, upstream open reading frame.
At the onset of maturation, CPEB is phosphorylated by a member of the aurora family of protein kinases83. Phosphorylation of CPEB increases its affinity for CPSF, thereby promoting binding of CPSF to the 3′ UTR of the dormant mRNAs (Fig. 4). CPSF then recruits poly(A) polymerase (PAP), which elongates the poly(A) tail on the mRNAs and creates additional binding sites for PABP. According to current models, PABP directly recruits eIF4G, which competes with Maskin for binding to eIF4E83. Enhanced eIF4G binding causes Maskin to dissociate and relieves translational repression of the dormant mRNAs. At the same time, formation of the PABP–eIF4G–eIF4E complex circularizes the mRNAs and promotes 43S-complex binding. So, translation of the targeted mRNAs is promoted by disruption of the CPEB–Maskin–eIF4E translation-silencing (masking) complex through enhanced binding of PABP to the elongated poly(A) tails and subsequent eIF4G recruitment (Fig. 4).
Recent studies indicate a crucial role for CPEB in synaptic plasticity. In Aplysia sensory neurons, LTF is correlated with a PI3K- and mTOR-dependent increase in CPEB, whereas downregulation of CPEB expression by antisense oligonucleotides blocks LTF85, which indicates that CPEB is required for this type of plasticity. LTF in Aplysia is also associated with an increase in actin mRNA polyadenylation that depends on cyclic AMP (cAMP)-dependent protein kinase86. CPEB is also required for hippocampal synaptic plasticity. LTP that is induced by theta-burst stimulation (TBS), E-LTP and captured LTP are reduced in CPEB1-knockout mice87. Interestingly, L-LTP that is induced with stronger stimulus protocols (for example, multiple trains of TBS and HFS) is unaltered in the CPEB1-knockout mice87. These findings indicate that CPEB1 is involved in LTP that is induced by weaker stimulation protocols but not in L-LTP. Other CPEB family members in the brain88 might be involved in L-LTP that is induced by strong stimulation protocols. Consistent with the proposed role of CPEB in synaptic plasticity, stimulation of cultured neurons with NMDA results in polyadenylation and translation of CPE-containing mRNAs in dendrites89. Although CPEB governs the translation of only a select group of mRNAs, it is noteworthy that one of these target mRNAs in the mouse brain encodes the α-subunit of calcium/calmodulin-dependent protein kinase II (α-CaMKII). Interestingly, α-CaMKII is rapidly synthesized after LTP-inducing stimulation90,91 and is necessary for LTP and memory92 (Box 1). One caveat to this model of CPEB-dependent translational regulation in synaptic plasticity is that no Maskin homologues have been identified in mammals. It is possible that distinct eIF4E-binding proteins might be functional homologues of Maskin, which are recruited by CPEB to compete with the eIF4E–eIF4G interaction, silence translation and generate dormant mRNAs.
Two extra features of CPEB indicate that CPEB-mediated translational regulation might be important in long-lasting synaptic plasticity and long-term memory. First, an amino-terminal extension of neuronal CPEB from Aplysia can be converted into a PRION-like state93. This autocatalytic mechanism in the mammalian hippocampus would allow CPEB to self-perpetuate at synapses, which provides a mechanism for the maintenance of long-lasting increases in protein synthesis during synaptic plasticity and long-term memory. Second, CPEB is regulated by CaMKII, a kinase that is crucial for both LTP and memory function92. CPEB is phosphorylated by CaMKII in vitro, and CPE-dependent translation that is induced by membrane depolarization also partially depends on CaMKII (Ref. 94). So, activation of CaMKII after LTP-inducing stimulation could stimulate CPE-dependent translation, including translation of α-CaMKII mRNA (Box 1), and promote a feedforward mechanism for maintaining increased protein synthesis during LTP and memory. Further studies are necessary to test the exciting hypothesis that persistent CPE-dependent translation is a molecular mechanism for long-lasting synaptic plasticity and long-term memory.
Concluding remarks
Studies in several model systems, including invertebrate and mammalian cell cultures, hippocampal slices and intact behaving animals, indicate that translational regulatory mechanisms are crucial for many forms of synaptic plasticity and memory. Although progress has been made in identifying signalling cascades that couple neurotransmitter and neurotrophin receptors to translation initiation regulatory factors, several important issues and questions remain to be considered.
One important question is whether protein synthesis-dependent synaptic plasticity in invertebrates and vertebrates involves the same translational regulatory mechanisms. Protein synthesis-dependent synaptic plasticity in vertebrates seems to be localized postsynaptically, whereas in invertebrates such as Aplysia it seems to be localized presynaptically. Presynaptic protein synthesis has not been convincingly shown in adult vertebrate systems; however, presynaptic protein synthesis is involved in growth-cone navigation in response to environmental stimuli in developing axons95. Similarly, within a particular neuron, it is not known whether the translational regulatory mechanisms are identical between axons, dendrites and the soma. So, although translational regulatory strategies seem to be conserved across species, key regulators such as the eIF2α and 4E-BP kinases might be spatially restricted, leading to the localized control of protein synthesis within the neuron.
Another important question concerns the spatial, temporal and magnitude differences in translational regulation during protein synthesis-dependent forms of synaptic plasticity and memory. The signalling pathways that couple various cell-surface receptors to translation initiation have several points of convergence and divergence, which would allow neurons to fine-tune translational regulation in response to a particular type of synaptic input. Careful characterization of the differences in the localization, duration and magnitude of the biochemical changes in translation initiation factors should provide important clues to the identity of the mRNAs that are translated in response to various types of input.
Traditionally, both LTP in the hippocampus and LTF in Aplysia have been described as having two distinct phases: a transient phase that involves post-translational modification of pre-existing proteins, and an enduring phase that requires de novo macromolecular synthesis. This enduring phase has distinct kinetic patterns of sensitivity to transcription and translation inhibitors: the former producing a delayed inhibition, and the latter impeding the initiation of the enduring plasticity62,96,97,98,99. These findings are consistent with the interesting possibility that local translation might be part of a retrograde signal that recruits nuclear transcription100. So, long-lasting synaptic plasticity might require initial induction of protein synthesis followed by induction of transcription to generate additional proteins. There are several examples of translation activation by particular patterns of stimulation that normally induce protein synthesis-independent, short-lasting plasticity58,101. What then is the function of translation activation? An intriguing possibility is that translation activation is a component of a tagging mechanism used by the activated synapse, whereby the synapse is marked for the consolidation of enduring synaptic plasticity. Although it is clear that translational activation does not account for all aspects of synaptic tagging96, it is required to mark the synapse for persistent functional58,101 and structural58 changes that underlie enduring plasticity. Finally, there are examples of translation-dependent, transcription-independent forms of synaptic plasticity. In both BDNF-induced potentiation16 and mGluR-dependent LTD17, local translation is necessary for the expression of synaptic plasticity. Despite these advances, it still is not known how the mRNAs are selected or how the encoded proteins are used to sustain the different types of synaptic plasticity (Box 4).
Much progress has been made in delineating biochemical signalling mechanisms that regulate translation initiation during synaptic plasticity and memory. Ongoing studies with diverse model systems — most notably knockout mice that are deficient in key translational regulatory factors — are revealing interesting links among the biochemical activities of translation factors, the electrophysiology of neurons and mouse behaviour. We anticipate that these and future studies will shed light on the molecular mechanisms that underlie protein synthesis-dependent synaptic plasticity and memory.
References
Kandel, E. R. The molecular biology of memory storage: a dialogue between genes and synapses. Science 294, 1030–1038 (2001).
Job, C. & Eberwine, J. Localization and translation of mRNA in dendrites and axons. Nature Rev. Neurosci. 2, 889–898 (2001).
West, A. E., Griffith, E. C. & Greenberg, M. E. Regulation of transcription factors by neuronal activity. Nature Rev. Neurosci. 3, 921–931 (2002).
Steward, O. & Levy, W. B. Preferential localization of polyribosomes under the base of dendritic spines in granule cells of the dentate gyrus. J. Neurosci. 2, 284–291 (1982).
Steward, O. Polyribosomes at the base of dendritic spines of central nervous system neurons — their possible role in synapse construction and modification. Cold Spring Harb. Symp. Quant. Biol. 48, 745–759 (1983).
Steward, O. & Schuman, E. M. Protein synthesis at synaptic sites on dendrites. Annu. Rev. Neurosci. 24, 299–325 (2001).
Tang, S. J. et al. A rapamycin-sensitive signaling pathway contributes to long-term synaptic plasticity in the hippocampus. Proc. Natl Acad. Sci. USA 99, 467–472 (2002). The first study to show that mTOR is required for hippocampal synaptic plasticity.
Inamura, N., Hoshino, S., Uchiumi, T., Nawa, H. & Takei, N. Cellular and subcellular distributions of translation initiation, elongation, and release factors in rat hippcampus. Brain Res. Mol. Brain Res. 111, 165–174 (2003).
Crino, P. B. & Eberwine, J. Molecular characterization of the dendritic growth cone: regulated mRNA transport and local protein synthesis. Neuron 17, 1173–1187 (1996). The first study to directly show that dendrites can translate mRNA.
Aakalu, G., Smith, W. B., Nguyen, N., Jiang, C. & Schuman, E. M. Dynamic visualization of local protein synthesis in hippocampal neurons. Neuron 30, 489–502 (2001).
Sutton, M. A., Wall, N. R., Aakalu, G. N. & Schuman, E. M. Regulation of dendritic protein synthesis by miniature synaptic events. Science 304, 1979–1983 (2004).
Ju, W. et al. Activity-dependent regulation of dendritic synthesis and trafficking of AMPA receptors. Nature Neurosci. 7, 244–253 (2004).
Weiler, I. J. et al. Fragile X mental retardation protein is translated near synapses in response to neurotransmitter activation. Proc. Natl Acad. Sci. USA 94, 5395–5400 (1997). Shows that stimulation of mGluRs results in the rapid translation of fragile X mental retardation protein near synapses.
Bagni, C., Mannucci, L., Dotti, C. G. & Amaldi, F. Chemical stimulation of synaptosomes modulates α-Ca2+/calmodulin-dependent protein kinase II mRNA association to polysomes. J. Neurosci. 20, RC76 (2000).
Yin, Y., Edelman, G. M. & Vanderklish, P. W. The brain-derived neurotrophic factor enhances synthesis of Arc in synaptoneurosomes. Proc. Natl Acad. Sci. USA 99, 2368–2373 (2002).
Kang, H. & Schuman, E. M. A requirement for local protein synthesis in neurotrophin-induced hippocampal synaptic plasticity. Science 273, 1402–1406 (1996). The first study to show that local protein synthesis is required for the potentiation of synpatic transmission in the hippocampus.
Huber, K. M., Kayser, M. S. & Bear, M. F. Role for rapid dendritic protein synthesis in hippocampal mGluR-dependent long-term depression. Science 288, 1254–1257 (2000). The first study to show that local protein synthesis is required for mGluR-LTD in the hippocampus.
Martin, K. C. et al. Synapse-specific, long-term facilitation of Aplysia sensory to motor synapses: a function for local protein synthesis in memory storage. Cell 91, 927–938 (1997). Shows that local protein synthesis is required for LTF in Aplysia.
Zhang, X. & Poo, M. M. Localized synaptic potentiation by BDNF requires local protein synthesis in the developing axon. Neuron 36, 675–688 (2002).
Ostroff, L. E., Fiala, J. C., Allwardt, B. & Harris, K. M. Polyribosomes redistribute from dendrtic shafts into spines with enlarged synapses during LTP in developing rat hippocampal slices. Neuron 35, 535–545 (2002).
Hershey, J. W. & Merrick, W. C. in Translational Control of Gene Expression (eds Sonenberg, N., Hershey, J. W. & Mathews, M. B.) 33–88 (Cold Spring Harbor Laboratory Press, Cold Spring Harbor, 2000).
Dever, T. E. Gene-specific regulation by general translation factors. Cell 108, 545–556 (2002).
Gingras, A. C., Raught, B. & Sonenberg, N. eIF4 initiation factors: effectors of mRNA recruitment to ribosomes and regulators of translation. Annu. Rev. Biochem. 68, 913–963 (1999).
Raught, B., Gingras, A. -C. & Sonenberg, N. in Translational Control of Gene Expression (eds Sonenberg, N., Hershey, J. W. & Mathews, M. B.) 245–294 (Cold Spring Harbor Laboratory Press, Cold Spring Harbor, 2000).
Kozak, M. Structural features in eukaryotic mRNAs that modulate the initiation of translation. J. Biol. Chem. 266, 19867–19870 (1991).
Wickens, M., Goodwin, E. B., Kimble, J., Strickland, S. & Hentze, M. W. in Translational Control of Gene Expression (eds Sonenberg, N., Hershey, J. W. & Mathews, M. B.) 295–370 (Cold Spring Harbor Laboratory Press, Cold Spring Harbor, 2000).
Hinnebusch, A. G. in Translational Control of Gene Expression (eds Sonenberg, N., Hershey, J. W. & Mathews, M. B.) 185–243 (Cold Spring Harbor Laboratory Press, Cold Spring Harbor, 2000).
Berlanga, J. J., Santoyo, J. & De Haro, C. Characterization of a mammalian homolog of the GCN2 eukaryotic initiation factor 2α kinase. Eur. J. Biochem. 265, 754–762 (1999).
Sood, R., Porter, A. C., Olsen, D., Cavener, D. R. & Wek, R. C. A mammalian homologue of GCN2 protein kinase important for translational control by phosphorylation of eukaryotic initiation factor-2α. Genetics 154, 787–801 (2000).
Santoyo, J., Alcalde, J., Mendez, R., Pulido, D. & de Haro, C. Cloning and characterization of a cDNA encoding a protein synthesis initiation factor-2α (eIF-2α) kinase from Drosophila melanogaster. J. Biol. Chem. 272, 12544–12550 (1997).
DeGracia, D. J., Neumar, R. W., White, B. C. & Krause, G. S. Global brain ischemia and reperfusion: modifications in eukaryotic initiation factors are associated with inhibition of translation initiation. J. Neurochem. 67, 2005–2012 (1996).
Kumar, R. et al. Brain ischemia and reperfusion activates the eukaryotic initiation factor 2α kinase, PERK. J. Neurochem. 77, 1418–1421 (2001).
Tan, S., Somia, N., Maher, P. & Schubert, D. Regulation of antioxidant metabolism by translation initiation factor 2α. J. Cell Biol. 152, 997–1006 (2001).
Balachandran, S. et al. Essential role for the dsRNA-dependent protein kinase PKR in innate immunity to viral infection. Immunity 13, 129–141 (2000).
Stojdl, D. F. et al. The murine double-stranded RNA-dependent protein kinase PKR is required for resistance to vesicular stomatitis virus. J. Virol. 74, 9580–9585 (2000).
Han, A. -P. et al. HRI is required for translational regulation and survival of erythroid precursors in iron deficiency. EMBO J. 20, 6909–6918 (2001).
Harding, H. P. et al. Diabetes mellitus and exocrine pancreatic dysfunction in Perk−/− mice reveals a role for translational control in secretory cell survival. Mol. Cell 7, 1153–1163 (2001).
Zhang, P. et al. The PERK eukaryotic initiation factor 2α kinase is required for the development of the skeletal system, postnatal growth, and the function and viability of the pancreas. Mol. Cell. Biol. 22, 3864–3874 (2002).
Delepine, M. et al. EIF2AK3, encoding translation initiation factor 2-α kinase 3, is mutated in patients with Wolcott–Rallison syndrome. Nature Genet. 25, 406–409 (2000).
Zhang, P. et al. The GCN2 eIF2α kinase is required for adaptation to amino acid deprivation in mice. Mol. Cell. Biol. 22, 6681–6688 (2002).
Anthony, T. et al. Preservation of liver protein synthesis during dietary leucine deprivation occurs at the expense of skeletal muscle mass in mice deleted for eIF2 kinase GCN2. J. Biol. Chem. 279, 36553–36561 (2004).
Hinnebusch, A. G. in Translational Control (eds Hershey, J. W., Mathews, M. B. & Sonenberg, N.) 199–244 (Cold Spring Harbor Laboratory Press, Cold Spring Harbor, New York, 1996).
Harding, H. P. et al. Regulated translation initiation controls stress-induced gene expression in mammalian cells. Mol. Cell 6, 1099–1108 (2000).
Vattem, K. M. & Wek, R. C. Reinitiation involving upstream ORFs regulates ATF4 mRNA translation in mammalian cells. Proc. Natl Acad. Sci. USA 101, 11269–11274 (2004).
Takei, N., Kawamura, M., Hara, K., Yonezawa, K. & Nawa, H. Brain-derived neurotrophic factor enhances neuronal translation by activating multiple initiation processes: comparison with the effects of insulin. J. Biol. Chem. 276, 42818–42825 (2001).
Leegwater, P. A. et al. Subunits of the translation initiation factor eIF2B are mutant in leukoencephalopathy with vanishing white matter. Nature Genet. 29, 383–388 (2001).
Welsh, G. I., Miller, C. M., Loughlin, A. J., Price, N. T. & Proud, C. G. Regulation of eukaryotic initiation factor eIF2B: glycogen synthase kinase-3 phosphorylates a conserved serine which undergoes dephosphorylation in response to insulin. FEBS Lett. 421, 125–130 (1998).
Kleijn, M. et al. Nerve and epidermal growth factor induce protein synthesis and eIF2B activation in PC12 cells. J. Biol. Chem. 273, 5536–5541 (1998).
Hernandez, F., Borrell, J., Guaza, C., Avila, J. & Lucas, J. J. Spatial learning deficit in transgenic mice that conditionally overexpress GSK-3β in the brain but do not form tau filaments. J. Neurochem. 83, 1529–1533 (2002).
Tsukiyama-Kohara, K. et al. Adipose tissue reduction in mice lacking the translational inhibitor 4E-BP1. Nature Med. 7, 1128–1132 (2001).
Gingras, A. -C. et al. Hierarchical phosphorylation of the translation inhibitor 4E-BP1. Genes Dev. 15, 2852–2864 (2001).
Thomas, G. M. & Huganir, R. L. MAPK cascade signalling and synaptic plasticity. Nature Rev. Neurosci. 5, 173–183 (2004).
Sweatt, J. D. Mitogen-activated protein kinases in synaptic plasticity and memory. Curr. Opin. Neurobiol. 14, 311–317 (2004).
Kelly, A. & Lynch, M. A. Long-term potentiation in dentate gyrus of the rat is inhibited by the phosphoinositide 3-kinase inhibitor wortmannin. Neuropharmacology 39, 643–651 (2000).
Beaumont, V., Zhong, N., Fletcher, R., Froemke, R. C. & Zucker, R. S. Phosphorylation and local presynaptic protein synthesis in calcium- and calcineurin-dependent induction of crayfish long-term facilitation. Neuron 32, 489–501 (2001).
Sanna, P. P. et al. Phosphatidylinositol 3-kinase is required for the expression but not for the induction or the maintenance of long-term potentiation in the hippocampal CA1 region. J. Neurosci. 22, 3359–3365 (2002).
Opazo, P., Watabe, A. M., Grant, S. G. & O'Dell, T. J. Phosphatidylinositol 3-kinase regulates the induction of long-term potentiation through extracellular signal-regulated kinase-independent mechanisms. J. Neurosci. 23, 3679–3688 (2003).
Casadio, A. et al. A transient, neuron-wide form of CREB-mediated long-term facilitation can be stabilized at specific synapses by local protein synthesis. Cell 99, 221–237 (1999).
Hou, L. & Klann, E. Activation of the phosphoinositide 3-kinase-Akt-mammalian target of rapamycin signaling pathway is required for metabotropic glutamate receptor-dependent long-term depression. J. Neurosci. 24, 6352–6361 (2004). Shows that stimulation of group I mGluRs results in activation of mTOR in dendrites and that mTOR is required for mGluR-dependent LTD.
Zho, W. M., You, J. L., Huang, C. C. & Hsu, K. S. The group I metabotropic glutamate receptor agonist (S)-3,5-dihydroxyphenylglycine induces a novel form of depotentiation in the CA1 region of the hippocampus. J. Neurosci. 22, 8838–8849 (2002).
Huang, C. -C., Lee, C. -C. & Hsu, K. -S. An investigation into signal transduction mechanisms involved in insulin-induced long-term depression in the CA1 region of the hippocampus. J. Neurochem. 89, 217–231 (2004).
Kelleher, R. J., Govindarajan, A., Jung, H. -Y., Kang, H. & Tonegawa, S. Translational control by MAPK signaling in long-term synaptic plasticity and memory. Cell 116, 1–20 (2004). The first study to show that ERK regulates translation during hippocampal synaptic plasticity and memory.
Scheper, G. & Proud, C. Does phosphorylation of the cap-binding protein eIF4E play a role in translation initiation? Eur. J. Biochem. 269, 5350–5359 (2002).
Pyronnet, S. et al. Human eukaryotic translation initiation factor 4G (eIF4G) recruits Mnk1 to phosphorylate eIF4E. EMBO J. 18, 270–279 (1999).
Waskiewicz, A. J. et al. Phosphorylation of the cap-binding protein eukaryotic translation initiation factor 4E by protein kinase Mnk1 in vivo. Mol. Cell. Biol. 19, 1871–1880 (1999).
Scheper, G., Morrice, N., Kleijn, M. & Proud, C. The mitogen-activated protein kinase signal-integrating kinase Mnk2 is a eukaryotic initiation factor 4E kinase with high levels of basal activity in mammalian cells. Mol. Cell. Biol. 21, 743–754 (2001).
Ueda, T., Watanabe-Fukunaga, R., Fukuyama, H., Nagata, S. & Fukunaga, R. Mnk2 and Mnk1 are essential for constitutive and inducible phosphorylation of eukaryotic initiation factor 4E but not for cell growth or development. Mol. Cell. Biol. 24, 6539–6549 (2004).
Scheper, G. C. et al. Phosphorylation of eukaryotic initiation factor 4E markedly reduces its affinity for capped mRNA. J. Biol. Chem. 277, 3303–3309 (2002).
Zuberek, J. et al. Phosphorylation of eIF4E attenuates its interaction with mRNA 5′ cap analogs by electrostatic repulsion: intein-mediated protein ligation strategy to obtain phosphorylated protein. RNA 9, 52–61 (2003).
Lachance, P. E., Miron, M., Raught, B., Sonenberg, N. & Lasko, P. Phosphorylation of eukaryotic translation initiation factor 4E is critical for growth. Mol. Cell. Biol. 22, 1656–1663 (2002).
Dyer, J. R. & Sossin, W. S. Regulation of eukaryotic initiation factor 4E phosphorylation in the nervous system of Aplysia californica. J. Neurochem. 75, 872–881 (2000).
Banko, J. L., Hou, L. & Klann, E. NMDA receptor activation results in PKA- and ERK-dependent Mnk1 activation and increased eIF4E phosphorylation in hippocampal area CA1. J. Neurochem. 91, 462–470 (2004).
Fumagalli, S. & Thomas, G. in Translational Control of Gene Expression (eds Sonenberg, N., Hershey, J. W. & Mathews, M. B.) 695–717 (Cold Spring Harbor Laboratory Press, Cold Spring Harbor, 2000).
Dufner, A. & Thomas, G. Ribosomal S6 kinase signaling and the control of translation. Exp. Cell Res. 253, 100–109 (1999).
Volarevic, S. et al. Proliferation, but not growth, blocked by conditional deletion of 40S ribosomal protein S6. Science, 288, 2045–2047 (2000).
Pende, M. et al. S6K1−/−/S6K2−/− mice exhibit perinatal lethality and rapamycin-sensitive 5′-terminal oligopyrimidine mRNA translation and reveal a mitogen-activated protein kinase-dependent S6 kinase pathway. Mol. Cell. Biol. 24, 3112–3124 (2004).
Meyuhas, O. & Hornstein, E. in Translational Control of Gene Expression (eds Sonenberg, N., Hershey, J. W. & Mathews, M. B.) 671–693 (Cold Spring Harbor Laboratory Press, Cold Spring Harbor, 2000).
Raught, B. et al. Phosphorylation of eucaryotic translation initiation factor 4B Ser422 is modulated by S6 kinases. EMBO J. 23, 1761–1769 (2004).
Khan, A., Pepio, A. M. & Sossin, W. S. Serotonin activates S6 kinase in a rapamycin-sensitive manner in Aplysia synaptosomes. J. Neurosci. 21, 382–391 (2001). Shows that the neurotransmitter serotonin can stimulate S6 kinase and TOP mRNA translation.
Carroll, M., Warren, O., Fan, X. & Sossin, W. S. 5-HT stimulates eEF2 dephosphorylation in a rapamycin-sensitive manner in Aplysia neurites. J. Neurochem. 90, 1464–1476 (2004).
Cammalleri, M. et al. Time-restricted role for dendritic activation of the mTOR-p70S6K pathway in the induction of late-phase long-term potentiation in the CA1. Proc. Natl Acad. Sci. USA 100, 14368–14373 (2003).
Dash, P. K., Mach, S. A., Moody, M. R. & Moore, A. N. Performance in long-term memory tasks is augmented by a phosphorylated growth factor receptor fragment. J. Neurosci. Res. 77, 205–216 (2004).
Mendez, R. & Richter, J. Translational control by CPEB: a means to the end. Nature Rev. Mol. Cell. Biol. 2, 521–529 (2001).
Stebbins-Boaz, B., Cao, Q., de Moor, C. H., Mendez, R. & Richter, J. D. Maskin is a CPEB-associated factor that transiently interacts with elF-4E. Mol. Cell 4, 1017–1027 (1999).
Si, K. et al. A neuronal isoform of CPEB regulates local protein synthesis and stabilizes synapse-specific long-term facilitation in Aplysia. Cell 115, 893–904 (2003). This study, together with reference 88, shows that CPEB is required for synaptic plasticity.
Liu, J. & Schwartz, J. H. The cytoplasmic polyadenylation element binding protein and polyadenylation of messenger RNAs in Aplysia neurons. Brain Res. 959, 68–76 (2003).
Alarcon, J. M. et al. Selective modulation of some forms of Schaffer collateral-CA1 synaptic plasticity in mice with a disruption of the CPEB-1 gene. Learn. Mem. 11, 318–327 (2004).
Theis, M., Si, K. & Kandel, E. R. Two previously undescribed members of the mouse CPEB family of genes and their inducible expression in the principal cell layers of the hippocampus. Proc. Natl Acad. Sci. USA 100, 9602–9607 (2003).
Huang, Y. -S., Jung, M. -Y., Sarkissian, M. & Richter, J. D. N-methyl-D-aspartate receptor signaling results in Aurora kinase-catalyzed CPEB phosphorylation and αCaMKII mRNA polyadenylation at synapses. EMBO J. 21, 2139–2148 (2002).
Ouyang, Y., Kantor, D. B., Harris, K. M., Schuman, E. M. & Kennedy, M. B. Visualization of the distribution of autophosphorylated calcium/calmodulin-dependent protein kinase II after tetanic stimulation in the CA1 area of the hippocampus. J. Neurosci. 17, 5416–5427 (1997). The first study to show that rapid dendritic synthesis of CaMKII occurs after delivery of LTP-inducing stimulation in the hippocampus.
Giovannini, M. G. et al. Mitogen-activated protein kinase regulates early phosphorylation and delayed expression of Ca2+/calmodulin-dependent protein kinase II in long-term potentiation. J. Neurosci. 21, 7053–7062 (2001).
Lisman, J., Schulman, H. & Cline, H. The molecular basis of CaMKII function in synaptic and behavioural memory. Nature Rev. Neurosci. 3, 175–190 (2002).
Si, K., Lindquist, S. & Kandel, E. R. A neuronal isoform of the Aplysia CPEB has prion-like properties. Cell 115, 879–891 (2003). The first study to show that CPEB has prion-like properties.
Atkins, C. M., Nozaki, N., Shigeri, Y. & Soderling, T. R. Cytoplasmic polyadenylation element binding protein-dependent protein synthesis is regulated by calcium/calmodulin-dependent protein kinase II. J. Neurosci. 24, 5193–5201 (2004).
Piper, M. & Holt, C. RNA translation in axons. Annu. Rev. Cell Dev. Biol. 20, 505–523 (2004).
Frey, U. & Morris, R. G. Synaptic tagging and long-term potentiation. Nature 385, 533–536 (1997).
Nguyen, P. V., Abel, T. & Kandel, E. R. Recruitment of a critical period of transcription for induction of a late phase of LTP. Science 265, 1104–1107 (1994).
Frey, U., Frey, S., Schollmeier, F. & Krug, M. Influence of actinomycin D, a RNA synthesis inhibitor, on long-term potentiation in rat hippocampal neurons in vivo and in vitro. J. Physiol. 490, 703–711 (1996).
Frey, U. & Morris, R. G. Synaptic tagging: implications for the late maintenance of hippocampal long-term potentiation. Trends Neurosci. 21, 181–188 (1998).
Martin, K. C. Synaptic tagging during synapse-specific long-term facilitation of Aplysia sensory-motor neurons. Neurobiol. Learn. Mem. 78, 489–497 (2002).
Barco, A., Alarcon, J. M. & Kandel, E. R. Expression of constitutively active CREB protein facilitates the late phase of long-term potentiation by enhancing synaptic capture. Cell 108, 689–703 (2002).
Hellen, C. U. & Sarnow, P. Internal ribosome entry sites in eukaryotic mRNA molecules. Genes Dev. 15, 1593–1612 (2001).
Dyer, J. et al. An activity-dependent switch to cap-independent translation triggered by eIF4E dephosphorylation. Nature Neurosci. 6, 219–220 (2003).
Huang, Y. -S., Carson, J. H., Barbarese, E. & Richter, J. D. Facilitation of dendritic mRNA transport by CPEB. Genes Dev. 17, 638–653 (2003).
Laggerbauer, B., Ostareck, D., Keidel, E. -M., Ostareck-Lederer, A. & Fischer, U. Evidence that fragile X mental retardation protein is a negative regulator of translation. Hum. Mol. Genet. 10, 329–338 (2001).
Li, Z. et al. The fragile X mental retardation protein inhibits translation via intereacting with mRNA. Nucleic Acids Res. 29, 2276–2283 (2001).
Zhang, Y. Q. et al. Drosophila fragile X-related gene regulates the MAP1B homolog Futsch to control synaptic structure and function. Cell 107, 591–603 (2001).
Jin, P. & Warren, S. T. New insights into fragile X syndrome: from molecules to neurobehaviors. Trends Biochem. Sci. 28, 152–158 (2003).
Antar, L. N., Afroz, R., Dictenberg, J. B., Carroll, R. C. & Bassell, G. J. Metabotropic glutamate receptor activation regulates fragile X mental retardation protein and Fmr1 mRNA localization differentially in dendrites and at synapses. J. Neurosci. 24, 2648–2655 (2004).
Feng, Y. et al. Fragile X mental retardation protein: nucleocytoplasmic shuttling and association with somatodendritic ribosomes. J. Neurosci. 17, 1539–1547 (1997).
Todd, P. K., Mack, K. J. & Malter, J. S. The fragile X mental retardation protein is required for type-I metabotropic glutamate receptor-dependent translation of PSD-95. Proc. Natl Acad. Sci. USA 100, 14374–14378 (2003).
Huber, K. M., Gallagher, S. M., Warren, S. T. & Bear, M. F. Altered synaptic plasticity in a mouse model of fragile X mental retardation. Proc. Natl Acad. Sci. USA 99, 7746–7750 (2002).
Godfraind, J. M. et al. Long-term potentiation in the hippocampus of fragile X knockout mice. Am. J. Med. Genet. 64, 246–251 (1996).
Paradee, W. et al. Fragile X mouse: strain effects of knockout phenotype and evidence suggesting deficient amygdala function. Neuroscience 94, 185–192 (1999).
Darnell, J. C. et al. Fragile X mental retardation protein targets G quartet mRNAs important for neuronal function. Cell 107, 489–499 (2001).
Schaefer, C., Bardoni, B., Moro, A., Bagni, C. & Mandel, J. L. The fragile X mental retardation protein binds specifically to its mRNA via a purine quartet motif. EMBO J. 20, 4803–4813 (2001).
Miyashiro, K. Y. et al. RNA cargoes associating with FMRP reveal deficits in cellular functioning in Fmr1 null mice. Neuron 37, 417–431 (2003).
Moldave, K. Eukaryotic protein synthesis. Annu. Rev. Biochem. 54, 1109–1149 (1985).
Ryazanov, A. G. & Davydova, E. K. Mechanism of elongation factor 2 (EF-2) inactivation upon phosphorylation. Phosphorylated EF-2 is unable to catalyze translocation. FEBS Lett. 251, 187–190 (1989).
Carlberg, U., Nilsson, A. & Nygard, O. Functional properties of phosphorylated elongation factor 2. Eur. J. Biochem. 191, 639–645 (1990).
Redpath, N. T., Price, N. T., Severinov, K. V. & Proud, C. G. Regulation of elongation factor-2 by multisite phosphorylation. Eur. J. Biochem. 213, 689–699 (1993).
Browne, G. J. & Proud, C. G. A novel mTOR-regulated phosphorylation site in elongation factor 2 kinase modulates the activity of the kinase and its binding to calmodulin. Mol. Cell. Biol. 24, 2986–2997 (2004).
Scheetz, A. J., Nairn, A. C. & Constantine-Paton, M. N-methyl-D-aspartate receptor activation and visual activity induce elongation factor-2 phosphorylation in amphibian tecta: a role for N-methyl-D-aspartate receptors in controlling protein synthesis. Proc. Natl Acad. Sci. USA 94, 14770–14775 (1997). This study, together with reference 125, shows that NMDA receptor activation regulates elongation factor 2 phosphorylation.
Marin, P. et al. Glutamate-dependent phosphorylation of elongation factor-2 and inhibition of protein synthesis in neurons. J. Neurosci. 17, 3445–3454 (1997).
Scheetz, A. J., Nairn, A. C. & Constantine-Paton, M. NMDA receptor-mediated control of protein synthesis at developing synapses. Nature Neurosci. 3, 211–216 (2000).
Chotiner, J. K., Khorasani, H., Nairn, A. C., O'Dell, T. J. & Watson, J. B. Adenylyl cyclase-dependent form of chemical long-term potentiation triggers translational regulation at the elongation step. Neuroscience 116, 743–752 (2003).
Dubnau, J. et al. The staufen/pumilio pathway is involved in Drosophila long-term memory. Curr. Biol. 13, 286–296 (2003).
Tang, S., Meulemans, D., Vazquez, L., Colaco, N. & Schuman, E. A role for a rat homolog of staufen in the transport of RNA to neuronal dendrites. Neuron 32, 463–475 (2001).
Macchi, P. et al. Barentsz, a new component of the Staufen-containing ribonucleoprotein particles in mammalian cells, interacts with Staufen in an RNA-dependent manner. J. Neurosci. 23, 5778–5788 (2003).
Kanai, Y., Dohmae, N. & Hirokawa, N. Kinesin transports RNA: isolation and characterization of an RNA-transporting granule. Neuron 43, 513–525 (2004).
Acknowledgements
We thank J. Richter and N. Sonenberg for communicating unpublished results, and J. Banko for helpful comments on the manuscript. E.K. is supported by the National Institutes of Health and the Fragile X (FRAXA) Research Foundation.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Related links
Related links
DATABASES
Entrez Gene
OMIM
FURTHER INFORMATION
Encyclopedia of Life Sciences
Protein phosphorylation and long-term synaptic plasticity
Glossary
- LONG-TERM FACILITATION
-
(LTF). A cellular analogue of long-term memory in Aplysia that can be induced at Aplysia sensory–motor synapses with multiple, spaced pulses of serotonin.
- LONG-TERM POTENTIATION
-
(LTP). A putative cellular model for memory that is manifested as a long-lasting increase in synaptic strength. LTP is most often induced with high-frequency stimulation of afferent inputs.
- LATE-PHASE LTP
-
(L-LTP). Transcription- and translation-dependent LTP that is typically induced with multiple, spaced trains of high-frequency stimulation. This type of LTP persists for more than 3 hours.
- SHORT-TERM FACILITATION
-
(STF). A cellular analogue of short-term memory in Aplysia that can be induced at Aplysia sensory–motor synapses with one pulse of serotonin.
- EARLY-PHASE LTP
-
(E-LTP). Transcription- and translation-independent LTP that is typically induced with one train of high-frequency stimulation. This type of LTP persists for 1–3 hours.
- LONG-TERM DEPRESSION
-
(LTD). A long-lasting decrease in synaptic strength that can be induced in hippocampal area CA1 by either low-frequency stimulation (NMDA receptor-dependent) or stimulation of group I metabotropic glutamate receptors.
- GENERAL CONTROL NON-DEREPRESSIBLE 4
-
(GCN4). A yeast transcription factor. Its expression is controlled by eIF2α phosphorylation, which regulates re-initiation at the upstream open reading frames in the GCN4 messenger RNA.
- ACTIVATING TRANSCRIPTION FACTOR 4
-
(ATF4). A mammalian transcription factor whose expression is regulated by eIF2α phosphorylation and upstream open reading frames, like GCN4 in yeast.
- INTERNAL RIBOSOMAL ENTRY SITE
-
(IRES). A messenger RNA (mRNA) sequence that directs 5′-cap-independent binding of the 40S subunit to an mRNA.
- ASSOCIATIVE FEAR CONDITIONING
-
An associative learning model in which the animal learns to associate a neutral conditioned stimulus (CS) with an aversive unconditioned stimulus (US). In many fear conditioning studies with rats and mice, the presentation of a harmless acoustic cue (CS) is paired with a mild foot shock (US) in a novel environment. At some point after training, the animals are tested for fear (usually freezing) in response to presentation of either the context (contextual fear conditioning) or the CS delivered in a novel context (cued fear conditioning). Both types of fear conditioning depend on the amygdala, whereas contextual fear conditioning also depends on the hippocampus.
- TERMINAL OLIGOPYRIMIDINES
-
(TOP). Messenger RNAs (mRNAs) that typically encode ribosomal proteins or translation factors. These mRNAs are translationally regulated through their 5′-terminal oligopyrimidine tracts.
- PRION
-
A protein that folds into two alternate conformations with different functions, wherein one conformation is self-perpetuated through conversion of the protein in the other confirmation.
Rights and permissions
About this article
Cite this article
Klann, E., Dever, T. Biochemical mechanisms for translational regulation in synaptic plasticity. Nat Rev Neurosci 5, 931–942 (2004). https://doi.org/10.1038/nrn1557
Issue Date:
DOI: https://doi.org/10.1038/nrn1557
This article is cited by
-
ERK/mTOR signaling may underlying the antidepressant actions of rapastinel in mice
Translational Psychiatry (2022)
-
Synaptic plasticity mechanisms behind TMS efficacy: insights from its application to animal models
Journal of Neural Transmission (2022)
-
Contribution of histone acetylation to the serotonin-mediated long-term synaptic plasticity in terrestrial snails
Journal of Comparative Physiology A (2022)
-
Mammalian/mechanistic target of rapamycin (mTOR) complexes in neurodegeneration
Molecular Neurodegeneration (2021)
-
Brain-specific suppression of AMPKα2 isoform impairs cognition and hippocampal LTP by PERK-mediated eIF2α phosphorylation
Molecular Psychiatry (2021)