Key Points
-
This Review covers the recent advances in synthetic biology and how these advances will affect the field of natural products.
-
There has been an emphasis on creating genetic parts, such as promoters, that generate precise levels of gene expression. The generation of large libraries of well-characterized parts and the development of biophysical and bioinformatic models to predict the behaviour of genetic parts in different organisms will aid in the transfer of biosynthetic gene clusters between hosts.
-
The capacity of DNA synthesis has exploded over the past decade and it is routine to synthesize the 20–100 kb required for a large gene cluster. In addition, new DNA assembly methods enable the rapid construction of different genetic part permutations or to substitute many genetic parts in a single step.
-
With regard to synthetic regulation, genetic circuits have been constructed that function as logic gates, timers, switches and oscillators. Sensors have also been developed that respond to many inducible inputs as well as metabolite levels. These could be incorporated into natural product pathways to control the timing of expression of different genes or to implement feedback in response to a toxic intermediate.
-
It is often desirable to make many simultaneous genomic changes. Methods such as CRISPR–Cas9 can target essentially any region of the genome and have been shown to function in many species, including several host species that are well suited for the industrial-scale production of small molecules.
Abstract
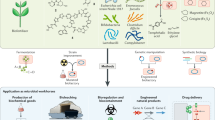
Bacterial genomes encode the biosynthetic potential to produce hundreds of thousands of complex molecules with diverse applications, from medicine to agriculture and materials. Accessing these natural products promises to reinvigorate drug discovery pipelines and provide novel routes to synthesize complex chemicals. The pathways leading to the production of these molecules often comprise dozens of genes spanning large areas of the genome and are controlled by complex regulatory networks with some of the most interesting molecules being produced by non-model organisms. In this Review, we discuss how advances in synthetic biology — including novel DNA construction technologies, the use of genetic parts for the precise control of expression and for synthetic regulatory circuits — and multiplexed genome engineering can be used to optimize the design and synthesis of pathways that produce natural products.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Newman, D. J. & Cragg, G. M. Natural products as sources of new drugs over the 30 years from 1981 to 2010. J. Nat. Prod. 75, 311–335 (2012).
Demain, A. L. Importance of microbial natural products and the need to revitalize their discovery. J. Ind. Microbiol. Biotechnol. 41, 185–201 (2014).
Davies, J. How to discover new antibiotics: harvesting the parvome. Curr. Opin. Chem. Biol. 15, 5–10 (2011).
Röttig, M. et al. NRPSpredictor2 — a web server for predicting NRPS adenylation domain specificity. Nucleic Acids Res. 39, W362–W367 (2011).
Kim, J. & Yi, G.-S. PKMiner: a database for exploring type II polyketide synthases. BMC Microbiol. 12, 169 (2012).
Cimermancic, P. et al. Insights into secondary metabolism from a global analysis of prokaryotic biosynthetic gene clusters. Cell 158, 412–421 (2014). This study provides a comparative analysis of 33,000 putative BGCs present in more than 1,000 sequenced bacterial and archaeal genomes. A large family of aryl polyene gene clusters was characterized as a result.
van Heel, A. J., de Jong, A., Montalbán-López, M., Kok, J. & Kuipers, O. P. BAGEL3: automated identification of genes encoding bacteriocins and (non-)bactericidal posttranslationally modified peptides. Nucleic Acids Res. 41, W448–W453 (2013).
Blin, K. et al. antiSMASH 2.0 — a versatile platform for genome mining of secondary metabolite producers. Nucleic Acids Res. 41, W204–W212 (2013).
Medema, M. H. et al. AntiSMASH: rapid identification, annotation and analysis of secondary metabolite biosynthesis gene clusters in bacterial and fungal genome sequences. Nucleic Acids Res. 39, W339–W346 (2011).
Weber, T. et al. antiSMASH 3.0 — a comprehensive resource for the genome mining of biosynthetic gene clusters. Nucleic Acids Res. 43, W237–W243 (2015).
Watve, M. G., Tickoo, R., Jog, M. M. & Bhole, B. D. How many antibiotics are produced by the genus Streptomyces? Arch. Microbiol. 176, 386–390 (2001).
Doroghazi, J. R. et al. A roadmap for natural product discovery based on large-scale genomics and metabolomics. Nat. Chem. Biol. 10, 963–968 (2014). This study describes a multi-parameter distance metric for comparing BGCs and applies this metric to organize more than 11,000 actinobacterial BGCs into families of gene clusters.
Galm, U. et al. In vivo manipulation of the bleomycin biosynthetic gene cluster in Streptomyces verticillus ATCC15003 revealing new insights into its biosynthetic pathway. J. Biol. Chem. 283, 28236–28245 (2008).
Piel, J. A polyketide synthase–peptide synthetase gene cluster from an uncultured bacterial symbiont of Paederus beetles. Proc. Natl Acad. Sci. USA 99, 14002–14007 (2002).
Galm, U. & Shen, B. Expression of biosynthetic gene clusters in heterologous hosts for natural product production and combinatorial biosynthesis. Expert Opin. Drug Discov. 1, 409–437 (2006).
Williams, G. J. Engineering polyketide synthases and nonribosomal peptide synthetases. Curr. Opin. Struct. Biol. 23, 603–612 (2013).
Pickens, L. B., Tang, Y. & Chooi, Y. H. Metabolic engineering for the production of natural products. Annu. Rev. Chem. Biomolecular Engineer. 2, 211–236 (2011).
Demain, A. L. & Adrio, J. L. Strain improvement for production of pharmaceuticals and other microbial metabolites by fermentation. Prog. Drug Res. 65, 252–289 (2008).
Voigt, C. A. Synthetic biology. ACS Synth. Biol. 1, 1–2 (2012).
Way, J. C., Collins, J. J. & Keasling, J. D. & Silver, P. A. Integrating biological redesign: where synthetic biology came from and where it needs to go. Cell 157, 151–161 (2014).
Medema, M. H., Breitling, R., Bovenberg, R. & Takano, E. Exploiting plug-and-play synthetic biology for drug discovery and production in microorganisms. Nat. Rev. Microbiol. 9, 131–137 (2011).
Luo, Y., Cobb, R. E. & Zhao, H. Recent advances in natural product discovery. Curr. Opin. Biotechnol. 30, 230–237 (2014).
Luo, Y. et al. Engineered biosynthesis of natural products in heterologous hosts. Chem. Soc. Rev. 44, 5265–5290 (2015).
Rutledge, P. J. & Challis, G. L. Discovery of microbial natural products by activation of silent biosynthetic gene clusters. Nat. Rev. Microbiol. 13, 509–523 (2015).
Keasling, J. D. Synthetic biology and the development of tools for metabolic engineering. Metab. Eng. 14, 189–195 (2012).
Fischbach, M. & Voigt, C. A. Prokaryotic gene clusters: a rich toolbox for synthetic biology. Biotechnol. J. 5, 1277–1296 (2010).
Endy, D. Foundations for engineering biology. Nature 438, 449–453 (2005).
Temme, K., Zhao, D. & Voigt, C. A. Refactoring the nitrogen fixation gene cluster from Klebsiella oxytoca. Proc. Natl Acad. Sci. USA 109, 7085–7090 (2012).
Chan, L. Y., Kosuri, S. & Endy, D. Refactoring bacteriophage T7. Mol. Syst. Biol. 1, 2005.0018 (2005).
Shao, Z. et al. Refactoring the silent spectinabilin gene cluster using a plug-and-play scaffold. ACS Synth. Biol. 2, 662–669 (2013).
Oßwald, C. et al. Modular construction of a functional artificial epothilone polyketide pathway. ACS Synth. Biol. 3, 759–772 (2012).
Luo, Y. et al. Activation and characterization of a cryptic polycyclic tetramate macrolactam biosynthetic gene cluster. Nat. Commun. 4, 2894 (2013).
Ajikumar, P. K. et al. Isoprenoid pathway optimization for Taxol precursor overproduction in Escherichia coli. Science 330, 70–74 (2010). This study showcases the approach of multivariate modular metabolic engineering for optimizing the biosynthesis of a Taxol precursor to 1 gram per litre in E. coli.
Paddon, C. J. et al. High-level semi-synthetic production of the potent antimalarial artemisinin. Nature 496, 528–532 (2013).
Mutalik, V. K. et al. Quantitative estimation of activity and quality for collections of functional genetic elements. Nat. Methods 10, 347–353 (2013).
Chen, Y.-J. et al. Characterization of 582 natural and synthetic terminators and quantification of their design constraints. Nat. Methods 10, 659–664 (2013).
Seghezzi, N., Amar, P., Koebmann, B., Jensen, P. R. & Virolle, M. J. The construction of a library of synthetic promoters revealed some specific features of strong Streptomyces promoters. Appl. Microbiol. Biotechnol. 90, 615–623 (2011).
Alper, H., Fischer, C., Nevoigt, E. & Stephanopoulos, G. Tuning genetic control through promoter engineering. Proc. Natl Acad. Sci. USA 102, 12678–12683 (2005).
Salis, H. M., Mirsky, E. A. & Voigt, C. A. Automated design of synthetic ribosome binding sites to control protein expression. Nat. Biotechnol. 27, 946–950 (2009).
Farasat, I. et al. Efficient search, mapping, and optimization of multi-protein genetic systems in diverse bacteria. Mol. Syst. Biol. 10, 731 (2014).
Nielsen, A. A. K., Segall-Shapiro, T. H. & Voigt, C. A. Advances in genetic circuit design: novel biochemistries, deep part mining, and precision gene expression. Curr. Opin. Chem. Biol. 17, 878–892 (2013).
Kosuri, S. et al. Composability of regulatory sequences controlling transcription and translation in Escherichia coli. Proc. Natl Acad. Sci. USA 110, 14024–14029 (2013).
Mutalik, V. K. et al. Precise and reliable gene expression via standard transcription and translation initiation elements. Nat. Methods 10, 354–360 (2013).
Cardinale, S. & Arkin, A. P. Contextualizing context for synthetic biology - identifying causes of failure of synthetic biological systems. Biotechnol. J. 7, 856–866 (2012).
Lou, C., Stanton, B., Chen, Y.-J., Munsky, B. & Voigt, C. A. Ribozyme-based insulator parts buffer synthetic circuits from genetic context. Nat. Biotechnol. 30, 1137–1142 (2012).
Bai, C. et al. Exploiting a precise design of universal synthetic modular regulatory elements to unlock the microbial natural products in Streptomyces. Proc. Natl Acad. Sci. USA 112, 12181–12186 (2015).
Davis, J. H., Rubin, A. J. & Sauer, R. T. Design, construction and characterization of a set of insulated bacterial promoters. Nucleic Acids Res. 39, 1131–1141 (2011).
Kalir, S. et al. Ordering genes in a flagella pathway by analysis of expression kinetics from living bacteria. Science 292, 2080–2083 (2001).
Temme, K. et al. Induction and relaxation dynamics of the regulatory network controlling the type III secretion system encoded within Salmonella pathogenicity Island 1. J. Mol. Biol. 377, 47–61 (2008).
Brophy, J. A. N. & Voigt, C. A. Principles of genetic circuit design. Nat. Methods 11, 508–520 (2014).
Liu, G., Chater, K. F., Chandra, G., Niu, G. & Tan, H. Molecular regulation of antibiotic biosynthesis in Streptomyces. Microbiol. Mol. Biol. Rev. 77, 112–143 (2013).
Chatterjee, A. et al. Convergent transcription in the butyrolactone regulon in Streptomyces coelicolor confers a bistable genetic switch for antibiotic biosynthesis. PLoS ONE 6, e21974 (2011).
Sherwood, E. J. & Bibb, M. J. The antibiotic planosporicin coordinates its own production in the actinomycete Planomonospora alba. Proc. Natl Acad. Sci. USA 110, E2500–E2509 (2013).
Chen, Y., Smanski, M. J. & Shen, B. Improvement of secondary metabolite production in Streptomyces by manipulating pathway regulation. Appl. Microbiol. Biotechnol. 86, 19–25 (2010).
Smanski, M. J., Peterson, R. M., Rajski, S. R. & Shen, B. Engineered Streptomyces platensis strains that overproduce antibiotics platensimycin and platencin. Antimicrob. Agents Chemother. 53, 1299–1304 (2009).
Tahlan, K. et al. Initiation of actinorhodin export in Streptomyces coelicolor. Mol. Microbiol. 63, 951–961 (2007).
Stevens, J. T. & Carothers, J. M. Designing RNA-based genetic control systems for efficient production from engineered metabolic pathways. ACS Synth. Biol. 4, 107–115 (2015).
Zhang, F., Carothers, J. M. & Keasling, J. D. Design of a dynamic sensor–regulator system for production of chemicals and fuels derived from fatty acids. Nat Biotechnol. 30, 354–359 (2012).
Kushwaha, M. & Salis, H. M. A portable expression resource for engineering cross-species genetic circuits and pathways. Nat. Commun. 6, 7832 (2015).
Solomon, K. V., Sanders, T. M. & Prather, K. L. J. A dynamic metabolite valve for the control of central carbon metabolism. Metab. Eng. 14, 661–671 (2012).
Brockman, I. M. & Prather, K. L. J. Dynamic knockdown of E. coli central metabolism for redirecting fluxes of primary metabolites. Metab. Eng. 28, 104–113 (2015).
Nieselt, K. et al. The dynamic architecture of the metabolic switch in Streptomyces coelicolor. BMC Genomics 11, 10 (2010).
Ellis, T., Wang, X. & Collins, J. J. Diversity-based, model-guided construction of synthetic gene networks with predicted functions. Nat. Biotechnol. 27, 465–471 (2009).
Gardner, T. S., Cantor, C. R. & Collins, J. J. Construction of a genetic toggle switch in Escherichia coli. Nature 403, 339–342 (2000).
Hasty, J., McMillen, D. & Collins, J. J. Engineered gene circuits. Nature 420, 224–230 (2002).
Qi, L. S. et al. Repurposing CRISPR as an RNA-guided platform for sequence-specific control of gene expression. Cell 152, 1173–1183 (2013). This study describes CRISPRi, which utilizes a catalytically inactive dCas9 protein to block gene expression by RNA polymerase.
Gilbert, L. A. et al. XCRISPR-mediated modular RNA-guided regulation of transcription in eukaryotes. Cell 154, 442–451 (2013). This study demonstrates that the CRISPR–dCas9 system can be used to activate gene expression by fusing transcriptional activator domains to dCas9.
Tong, Y., Charusanti, P., Zhang, L., Weber, T. & Lee, S. Y. CRISPR–Cas9 based engineering of actinomycetal genomes. ACS Synth. Biol. 4, 1020–1029 (2015).
Piatek, A. et al. RNA-guided transcriptional regulation in planta via synthetic dCas9-based transcription factors. Plant Biotechnol. J. 13, 578–589 (2015).
Lv, L., Ren, Y., Chen, J., Wu, Q. & Chen, G. Application of CRISPRi for prokaryotic metabolic engineering involving multiple genes, a case study: controllable P (3HB-co-4HB) biosynthesis. Metab. Eng. 29, 160–168 (2015).
Zalatan, J. G. et al. Engineering complex synthetic transcriptional programs with CRISPR RNA scaffolds. Cell 160, 339–350 (2014).
Zetsche, B., Volz, S. E. & Zhang, F. A split-Cas9 architecture for inducible genome editing and transcription modulation. Nat. Biotechnol. 33, 139–142 (2015).
Sánchez, C. et al. Combinatorial biosynthesis of antitumor indolocarbazole compounds. Proc. Natl Acad. Sci. USA 102, 461–466 (2005).
Menzella, H. G. et al. Combinatorial polyketide biosynthesis by de novo design and rearrangement of modular polyketide synthase genes. Nat. Biotechnol. 23, 1171–1176 (2005).
Kakule, T. B., Lin, Z. & Schmidt, E. W. Combinatorialization of fungal polyketide synthase–peptide synthetase hybrid proteins J. Am. Chem. Soc. 136, 17882–17890 (2014).
Nguyen, K. T. et al. Combinatorial biosynthesis of novel antibiotics related to daptomycin. Proc. Natl Acad. Sci. USA 103, 17462–17467 (2006).
Kosuri, S. & Church, G. M. Large-scale de novo DNA synthesis: technologies and applications. Nat. Methods 11, 499–507 (2014).
Ru, D. E., Schmidt, E. W. & Heemstra, J. R. Assessing the combinatorial potential of the RiPP cyanobactin tru pathway. ACS Synth. Biol. 4, 482–492 (2015). The authors use high-throughput mutagenesis and analytical chemistry to create more than 300 structural variants of the cyanobacterial RiPP, trunkamide. The data were analysed to establish rules for amino acid preference at various positions along the core scaffold.
Mitchell, D. A. et al. Structural and functional dissection of the heterocyclic peptide cytotoxin streptolysin S. J. Biol. Chem. 284, 13004–13012 (2009).
Smanski, M. J. et al. Functional optimization of gene clusters by combinatorial design and assembly. Nat. Biotechnol. 32, 1241–1249 (2014).
Appleton, E., Tao, J., Haddock, T. & Densmore, D. Interactive assembly algorithms for molecular cloning. Nat. Methods 11, 657–662 (2014).
Freestone, T. S. & Zhao, H. Combinatorial pathway engineering for optimized production of the anti-malarial FR900098. Biotechnol. Bioeng. 113, 384–392 (2015).
Biggs, B. W., De Paepe, B., Santos, C. N. S., De Mey, M. & Ajikumar, P. K. Multivariate modular metabolic engineering for pathway and strain optimization. Curr. Opin. Biotechnol. 29, 156–162 (2014).
Thodey, K., Galanie, S. & Smolke, C. D. A microbial biomanufacturing platform for natural and semisynthetic opioids. Nat. Chem. Biol. 10, 837–844 (2014).
Li, L. et al. A stepwise increase in pristinamycin II biosynthesis by Streptomyces pristinaespiralis through combinatorial metabolic engineering. Metab. Eng. 29, 12–25 (2015).
Zhao, S. et al. Improvement of catechin production in Escherichia coli through combinatorial metabolic engineering. Metab. Eng. 28, 43–53 (2015).
Du, Y.-L. & Ryan, K. S. Expansion of bisindole biosynthetic pathways by combinatorial construction. ACS Synth. Biol. 4, 682–688 (2015).
Chemler, J. a. et al. Evolution of efficient modular polyketide synthases by homeologous recombination. J. Am. Chem. Soc. 137, 10603–10609 (2015).
Smanski, M. J. et al. Expression of the platencin biosynthetic gene cluster in heterologous hosts yielding new platencin congeners. J. Nat. Prod. 75, 2158–2167 (2012).
Wang, B., Kitney, R. I., Joly, N. & Buck, M. Engineering modular and orthogonal genetic logic gates for robust digital-like synthetic biology. Nat. Commun. 2, 508 (2011).
Hajimorad, M., Gray, P. R. & Keasling, J. D. A framework and model system to investigate linear system behavior in Escherichia coli. J. Biol. Eng. 5, 3 (2011).
Moser, F. et al. Genetic circuit performance under conditions relevant for industrial bioreactors. ACS Synth. Biol. 1, 555–564 (2012).
Siegl, T., Tokovenko, B., Myronovskyi, M. & Luzhetskyy, A. Design, construction and characterisation of a synthetic promoter library for fine-tuned gene expression in actinomycetes. Metab. Eng. 19, 98–106 (2013).
Sarrion-Perdigones, A. et al. GoldenBraid 2.0: a comprehensive DNA assembly framework for plant synthetic biology. Plant Physiol. 162, 1618–1631 (2013).
Zhang, Y., Perry, K., Vinci, V. & Powell, K. Genome shuffling leads to rapid phenotypic improvement in bacteria. Nature 415, 5–7 (2002).
Gibson, D. G. et al. Creation of a bacterial cell controlled by a chemically synthesized genome. Science 329, 52–56 (2010).
Warner, J. R., Reeder, P. J., Karimpour-Fard, A., Woodruff, L. B. A. & Gill, R. T. Rapid profiling of a microbial genome using mixtures of barcoded oligonucleotides. Nat. Biotechnol. 28, 856–862 (2010).
Isaacs, F. J. et al. Precise manipulation of chromosomes in vivo enables genome-wide codon replacement. Science 333, 348–353 (2011).
Wang, H. H. et al. Genome-scale promoter engineering by coselection MAGE. Nat. Methods 9, 591–593 (2012).
Wang, H. H. et al. Programming cells by multiplex genome engineering and accelerated evolution. Nature 460, 894–898 (2009).
Wang, H. H. & Church, G. M. Multiplexed genome engineering and genotyping methods: applications for synthetic biology and metabolic engineering. Methods Enzymol. 498, 409–426 (2011).
Wang, H. H. et al. Multiplexed in vivo his-tagging of enzyme pathways for in vitro single-pot multienzyme catalysis. ACS Synth. Biol. 1, 43–52 (2012).
Bonde, M. T. et al. Direct mutagenesis of thousands of genomic targets using microarray-derived oligonucleotides. ACS Synth. Biol. 4, 17–22 (2015). In this study, the authors develop a workflow for the integration of microarray-based DNA synthesis with MAGE and apply their approach to introduce T7 promoters upstream of 2,500 operons in the E. coli genome.
Binder, S., Siedler, S., Marienhagen, J., Bott, M. & Eggeling, L. Recombineering in Corynebacterium glutamicum combined with optical nanosensors: a general strategy for fast producer strain generation. Nucleic Acids Res. 41, 6360–6369 (2013).
Dicarlo, J. E. et al. Yeast oligo-mediated genome engineering (YOGE). ACS Synth. Biol. 2, 741–749 (2013).
Sandoval, N. R. et al. Strategy for directing combinatorial genome engineering in Escherichia coli. Proc. Natl Acad. Sci. USA 109, 10540–10545 (2012).
Belhaj, K., Chaparro-Garcia, A., Kamoun, S. & Nekrasov, V. Plant genome editing made easy: targeted mutagenesis in model and crop plants using the CRISPR/Cas system. Plant Methods 9, 39 (2013).
Cobb, R. E., Wang, Y. & Zhao, H. High-efficiency multiplex genome editing of Streptomyces species using an engineered CRISPR/Cas system. ACS Synth. Biol. 4, 723–728 (2014). This is the first study to demonstrate CRISPR–Cas9 genome editing in Streptomyces spp.; the authors achieve up to 100% editing efficiency for the deletion of single genes or whole gene clusters.
Cong, L. et al. Multiplex genome engineering using CRISPR/Cas systems. Science 339, 819–823 (2013).
Jiang, W., Bikard, D., Cox, D., Zhang, F. & Marraffini, L. A. RNA-guided editing of bacterial genomes using CRISPR–Cas systems. Nat. Biotechnol. 31, 233–239 (2013).
Jakočiūnas, T. et al. Multiplex metabolic pathway engineering using CRISPR/Cas9 in Saccharomyces cerevisiae. Metab. Eng. 28, 213–222 (2015).
Huang, H., Zheng, G., Jiang, W., Hu, H. & Lu, Y. One-step high-efficiency CRISPR/Cas9-mediated genome editing in Streptomyces. Acta Biochim. Biophys. Sin. (Shanghai). 47, 231–243 (2015).
Konermann, S. et al. Genome-scale transcriptional activation by an engineered CRISPR–Cas9 complex. Nature 517, 583–588 (2015).
Doudna, J. A. & Charpentier, E. The new frontier of genome engineering with CRISPR–Cas9. Science 346, 1–9 (2014).
Bao, Z., Cobb, R. E. & Zhao, H. Accelerated genome engineering through multiplexing. Wiley Interdiscip. Rev. Syst. Biol. Med. 8, 5–21 (2015).
Chen, C., Fenk, L. a. & De Bono, M. Efficient genome editing in Caenorhabditis elegans by CRISPR-targeted homologous recombination. Nucleic Acids Res. 41, e193 (2013).
Michener, J. K., Thodey, K., Liang, J. C. & Smolke, C. D. Applications of genetically-encoded biosensors for the construction and control of biosynthetic pathways. Metab. Eng. 14, 212–222 (2012).
An, G. H., Bielich, J., Auerbach, R. & Johnson, E. A. Isolation and characterization of carotenoid hyperproducing mutants of yeast by flow cytometry and cell sorting. Biotechnology 9, 70–73 (1991).
Cho, E. J., Lee, J.-W. & Ellington, A. D. Applications of aptamers as sensors. Annu. Rev. Anal. Chem. (Palo Alto. Calif.). 2, 241–264 (2009).
Wachsmuth, M., Findeiss, S., Weissheimer, N., Stadler, P. F. & Morl, M. De novo design of a synthetic riboswitch that regulates transcription termination. Nucleic Acids Res. 41, 2541–2551 (2012).
Weigand, J. E. & Suess, B. Tetracycline aptamer-controlled regulation of pre-mRNA splicing in yeast. Nucleic Acids Res. 35, 4179–4185 (2007).
Michener, J. K. & Smolke, C. D. High-throughput enzyme evolution in Saccharomyces cerevisiae using a synthetic RNA switch. Metab. Eng. 14, 306–316 (2012).
Weigand, J. E. et al. Screening for engineered neomycin riboswitches that control translation initiation. RNA 14, 89–97 (2008).
Schoukroun-Barnes, L. R., Wagan, S. & White, R. J. Enhancing the analytical performance of electrochemical RNA aptamer-based sensors for sensitive detection of aminoglycoside antibiotics. Anal. Chem. 86, 1131–1137 (2014).
Farjami, E. et al. RNA aptamer-based electrochemical biosensor for selective and label-free analysis of dopamine. Anal. Chem. 85, 121–128 (2013).
Chen, J., Fang, Z., Liu, J. & Zeng, L. A simple and rapid biosensor for ochratoxin A based on a structure-switching signaling aptamer. Food Control 25, 555–560 (2012).
Gossen, M. & Bujard, H. Tight control of gene expression in mammalian cells by tetracycline-responsive promoters. Proc. Natl Acad. Sci. USA 89, 5547–5551 (1992).
Grkovic, S., Hardie, K. M., Brown, M. H. & Skurray, R. A. Interactions of the QacR multidrug-binding protein with structurally diverse ligands: implications for the evolution of the binding pocket. Biochemistry 42, 15226–15236 (2003).
Ramos, J. L. et al. The TetR family of transcriptional repressors. Microbiol. Mol. Biol. Rev. 69, 326–356 (2005).
Mohn, W. W., Garmendia, J., Galvao, T. C. & De Lorenzo, V. Surveying biotransformations with à la carte genetic traps: translating dehydrochlorination of lindane (γ-hexachlorocyclohexane) into lacZ-based phenotypes. Environ. Microbiol. 8, 546–555 (2006).
Mustafi, N., Grünberger, A., Kohlheyer, D., Bott, M. & Frunzke, J. The development and application of a single-cell biosensor for the detection of L-methionine and branched-chain amino acids. Metab. Eng. 14, 449–457 (2012).
Tang, S. Y. & Cirino, P. C. Design and application of a mevalonate-responsive regulatory protein. Angew. Chem. Int. Ed. Engl. 50, 1084–1086 (2011).
Tang, S. Y. et al. Screening for enhanced triacetic acid lactone production by recombinant Escherichia coli expressing a designed triacetic acid lactone reporter. J. Am. Chem. Soc. 135, 10099–10103 (2013).
Buskirk, A. R., Ong, Y.-C., Gartner, Z. J. & Liu, D. R. Directed evolution of ligand dependence: small-molecule-activated protein splicing. Proc. Natl Acad. Sci. USA 101, 10505–10510 (2004).
Peck, S. H., Chen, I. & Liu, D. R. Directed evolution of a small-molecule-triggered intein with improved splicing properties in mammalian cells. Chem. Biol. 18, 619–630 (2011).
DeLoache, W. C. et al. An enzyme-coupled biosensor enables (S)-reticuline production in yeast from glucose. Nat. Chem. Biol. 11, 465–471 (2015).
Gandía-Herrero, F. & García-Carmona, F. Biosynthesis of betalains: yellow and violet plant pigments. Trends Plant Sci. 18, 334–343 (2013).
Meyer, A. et al. Optimization of a whole-cell biocatalyst by employing genetically encoded product sensors inside nanolitre reactors. Nat. Chem. 7, 673–678 (2015).
Pfleger, B. F., Pitera, D. J., Newman, J. D., Martin, V. J. J. & Keasling, J. D. Microbial sensors for small molecules: development of a mevalonate biosensor. Metab. Eng. 9, 30–38 (2007).
Bertels, F., Merker, H. & Kost, C. Design and characterization of auxotrophy-based amino acid biosensors. PLoS ONE 7, e41349 (2012).
Bugni, T. S. et al. Marine natural product libraries for high-throughput screening and rapid drug discovery. J. Nat. Prod. 71, 1095–1098 (2008).
Kotula, J. W. et al. Programmable bacteria detect and record an environmental signal in the mammalian gut. Proc. Natl Acad. Sci. USA 111, 4838–4843 (2014).
Claesen, J. & Fischbach, M. A. Synthetic microbes as drug delivery systems. ACS Synth. Biol. 4, 358–364 (2015).
Fischbach, M. A., Bluestone, J. A. & Lim, W. A. Cell-based therapeutics: the next pillar of medicine. Sci. Transl. Med. 5, 179ps7 (2013).
Qian, P.-Y., Xu, Y. & Fusetani, N. Natural products as antifouling compounds: recent progress and future perspectives. Biofouling 26, 223–234 (2010).
Yoshino, T. & Matsunaga, T. Development of efficient expression system for protein display on bacterial magnetic particles. Biochem. Biophys. Res. Commun. 338, 1678–1681 (2005).
Dayan, F. E., Cantrell, C. L. & Duke, S. O. Natural products in crop protection. Bioorg. Med. Chem. 17, 4022–4034 (2009).
Zimmermann, M. & Fischbach, M. A. A family of pyrazinone natural products from a conserved nonribosomal peptide synthetase in Staphylococcus aureus. Chem. Biol. 17, 925–930 (2010).
Smanski, M. J. et al. Dedicated ent-kaurene and ent-atiserene synthases for platensimycin and platencin biosynthesis. Proc. Natl Acad. Sci. USA 108, 13498–13503 (2011).
Sudek, S. et al. Identification of the putative bryostatin polyketide synthase gene cluster from 'Candidatus Endobugula sertula', the uncultivated microbial symbiont of the marine bryozoan Bugula neritina. J. Nat. Prod. 70, 67–74 (2007).
Wong, F. T. & Khosla, C. Combinatorial biosynthesis of polyketides — a perspective. Curr. Opin. Chem. Biol. 16, 117–123 (2012).
Medema, M. H. Minimum information about a biosynthetic gene cluster. Nat. Chem. Biol. 11, 625–631 (2015).
Carlson, R. Time for new DNA synthesis and sequencing cost curves. Synthesis [online], (2014).
Gustafsson, C., Govindarajan, S. & Minshull, J. Codon bias and heterologous protein expression. Trends Biotechnol. 22, 346–353 (2004).
Rodríguez-García, A., Combes, P., Pérez-Redondo, R., Smith, M. C. A. & Smith, M. C. M. Natural and synthetic tetracycline-inducible promoters for use in the antibiotic-producing bacteria Streptomyces. Nucleic Acids Res. 33, e87 (2005).
Stanton, B. C. et al. Systematic transfer of prokaryotic sensors and circuits to mammalian cells. ACS Synth. Biol. 19, 880–891 (2014).
Selle, K. & Barrangou, R. Harnessing CRISPR–Cas systems for bacterial genome editing. Trends Microbiol. 23, 225–232 (2015).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Glossary
- Polyketide
-
One of a family of natural products that share a common biosynthetic pathway through the decarboxylative condensation of substituted malonyl-CoA-derived extender units and acyl-CoA starter units on polyketide synthase enzymes.
- Non-ribosomal peptide
-
One of a family of natural products that share a common biosynthetic pathway through the condensation of proteogenic or non-proteogenic amino acids on modular non-ribosomal peptide synthetase enzymes.
- Terpene
-
One of a family of natural products that share a common biosynthetic pathway through the polymerization of branched five-carbon isoprene units and cyclization by terpene synthases.
- Ribosomally synthesized and post-translationally modified peptide
-
(RiPP). One of a family of natural products, including the lanthipeptides, bacteriocins, and thiazole-modified or oxazole-modified microcins, that share a common biosynthetic pathway through the translation of an mRNA-encoded core peptide and subsequent modification.
- Random chemical mutagenesis
-
A process by which cells or organisms are exposed to chemical mutagens to introduce mutations at random locations in the genome.
- 5′ untranslated region
-
(5′ UTR). The untranslated region of an mRNA transcript that is upstream of the start codon. The sequence of the 5′ UTR can influence translation initiation and mRNA stability.
- Bicistrons
-
Transcripts that contain two coding DNA sequences (CDSs). For translational control, the first CDS encodes a short, non-functional peptide and is located immediately upstream of the ribosome binding site for the second CDS.
- Actinorhodin
-
A benzoisochromanequinone polyketide pigment produced by Streptomyces coelicolor.
- para-aminostyrene
-
An industrially relevant vinyl aromatic monomer with applications in materials and biomedicine.
- Undecylprodigiosin
-
A tripyrrole polyketide pigment produced by Streptomyces coelicolor.
- Gibson assembly
-
A restriction-enzyme-independent method for the joining of several DNA fragments in a single isothermal reaction.
- Pristinamycin II
-
One of two structurally unrelated chemical components of the clinical antibiotic pristinamycin. Pristinamycin II is a depsipeptide antibiotic produced by Streptomyces pristinaespiralis.
- Catechin
-
A plant flavonoid with antioxidant properties.
- Indolocarbazoles
-
A family of five-ring heterocyclic aromatic compounds that share a common biosynthetic pathway from two tryptophan molecules.
- Platencin
-
An antibiotic produced by Streptomyces platensis containing an amino-dihydroxybenzoic acid moiety fused to a modified diterpene core.
- Shunt metabolites
-
Chemically modified intermediates of a biosynthetic pathway that can no longer proceed through the biosynthetic pathway.
- Tylosin
-
A glycosylated macrolide antibiotic produced by Streptomyces venezuelae.
- Okazaki fragments
-
Short, newly synthesized single-stranded DNA oligomers that are formed on the lagging template strand during DNA replication.
- Lycopene
-
A symmetrical tetraterpene pigment formed by the tail-to-tail condensation of two molecules of geranylgeranyl diphosphate.
- T7 RNA polymerase promoters
-
Short DNA sequences that are recognized by T7 RNA polymerase to initiate transcription.
Rights and permissions
About this article
Cite this article
Smanski, M., Zhou, H., Claesen, J. et al. Synthetic biology to access and expand nature's chemical diversity. Nat Rev Microbiol 14, 135–149 (2016). https://doi.org/10.1038/nrmicro.2015.24
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrmicro.2015.24
This article is cited by
-
Epigallocatechin-3-gallate and cancer: focus on the role of microRNAs
Cancer Cell International (2023)
-
Phenotypically complex living materials containing engineered cyanobacteria
Nature Communications (2023)
-
A portable regulatory RNA array design enables tunable and complex regulation across diverse bacteria
Nature Communications (2023)
-
Synthetic genetic oscillators demonstrate the functional importance of phenotypic variation in pneumococcal-host interactions
Nature Communications (2023)
-
Design of a redox-proficient Escherichia coli for screening terpenoids and modifying cytochrome P450s
Nature Catalysis (2023)