Abstract
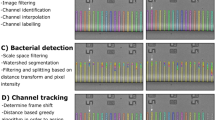
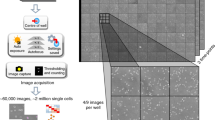
Single-cell microscopy is a powerful tool for studying gene functions using strain libraries, but it suffers from throughput limitations. Here we describe the Strain Library Imaging Protocol (SLIP), which is a high-throughput, automated microscopy workflow for large strain collections that requires minimal user involvement. SLIP involves transferring arrayed bacterial cultures from multiwell plates onto large agar pads using inexpensive replicator pins and automatically imaging the resulting single cells. The acquired images are subsequently reviewed and analyzed by custom MATLAB scripts that segment single-cell contours and extract quantitative metrics. SLIP yields rich data sets on cell morphology and gene expression that illustrate the function of certain genes and the connections among strains in a library. For a library arrayed on 96-well plates, image acquisition can be completed within 4 min per plate.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Dörr, T. et al. Differential requirement for PBP1a and PBP1b in in vivo and in vitro fitness of Vibrio cholerae. Infect. Immun. 82, 2115–2124 (2014).
Monds, R.D. et al. Systematic perturbation of cytoskeletal function reveals a linear scaling relationship between cell geometry and fitness. Cell Rep. 9, 1528–1537 (2014).
Yao, Z., Kahne, D. & Kishony, R. Distinct single-cell morphological dynamics under beta-lactam antibiotics. Mol. Cell 48, 705–712 (2012).
Zambrano, M.M. & Kolter, R. GASPing for life in stationary phase. Cell 86, 181–184 (1996).
Helaine, S. et al. Internalization of Salmonella by macrophages induces formation of nonreplicating persisters. Science 343, 204–208 (2014).
Baba, T. et al. Construction of Escherichiacoli K-12 in-frame, single-gene knockout mutants: the Keio collection. Mol. Syst. Biol. 2, 2006.0008 (2006).
Bogomolnaya, L.M. et al. Identification of novel factors involved in modulating motility of Salmonella enterica serotype typhimurium. PLoS One 9, e111513 (2014).
Wang, H.H. et al. Programming cells by multiplex genome engineering and accelerated evolution. Nature 460, 894–898 (2009).
Peters, J.M. et al. A comprehensive, CRISPR-based functional analysis of essential genes in bacteria. Cell 165, 1493–1506 (2016).
Nichols, R.J. et al. Phenotypic landscape of a bacterial cell. Cell 144, 143–156 (2011).
Young, J.W. et al. Measuring single-cell gene expression dynamics in bacteria using fluorescence time-lapse microscopy. Nat. Protoc. 7, 80–88 (2012).
Edelstein, A.D. et al. Advanced methods of microscope control using μManager software. J. Biol. Methods 1, e10 (2014).
Kitagawa, M. et al. Complete set of ORF clones of Escherichia coli ASKA library (a complete set of E. coli K-12 ORF archive): unique resources for biological research. DNA Res. 12, 291–299 (2006).
Kuwada, N.J., Traxler, B. & Wiggins, P.A. Genome-scale quantitative characterization of bacterial protein localization dynamics throughout the cell cycle. Mol. Microbiol. 95, 64–79 (2015).
Wang, P. et al. Robust growth of Escherichia coli. Curr. Biol. 20, 1099–1103 (2010).
Gefen, O., Fridman, O., Ronin, I. & Balaban, N.Q. Direct observation of single stationary-phase bacteria reveals a surprisingly long period of constant protein production activity. Proc. Natl. Acad. Sci. 111, 556–561 (2014).
Ursell, T. et al. Rapid, precise quantification of bacterial cellular dimensions across a genomic-scale knockout library. BMC Biol. (in the press).
Paintdakhi, A. et al. Oufti: an integrated software package for high-accuracy, high-throughput quantitative microscopy analysis. Mol. Microbiol. 99, 767–777 (2016).
Ducret, A., Quardokus, E.M. & Brun, Y.V. MicrobeJ, a tool for high throughput bacterial cell detection and quantitative analysis. Nat. Microbiol. 1, 16077 (2016).
Bacus, J.W. & Hernicz, R.S. Dual color camera microscope and methodology for cell staining and analysis. US Patent (1991).
Chang, F. & Huang, K.C. How and why cells grow as rods. BMC Biol. 12, 54 10.1186/s12915-014-0054-8 (2014).
Sezonov, G., Joseleau-Petit, D. & D'Ari, R. Escherichia coli physiology in Luria-Bertani broth. J. Bacteriol. 189, 8746–8749 (2007).
Sliusarenko, O., Heinritz, J., Emonet, T. & Jacobs-Wagner, C. High-throughput, subpixel precision analysis of bacterial morphogenesis and intracellular spatio-temporal dynamics. Mol. Microbiol. 80, 612–627 (2011).
Kjeldgaard, N., Maaløe, O. & Schaechter, M. The transition between different physiological states during balanced growth of Salmonella typhimurium. Microbiology 19, 607–616 (1958).
Gitai, Z., Dye, N.A., Reisenauer, A., Wachi, M. & Shapiro, L. MreB actin-mediated segregation of a specific region of a bacterial chromosome. Cell 120, 329–341 (2005).
Acknowledgements
The authors thank members of the Huang lab, especially M. Anjur-Dietrich, M. Rajendram, and A. Aranda-Diaz, for insightful discussions and beta-testing the SLIP software. This work was supported by a Stanford Agilent Fellowship and a Stanford Interdisciplinary Graduate Fellowship (to H.S.), a Stanford Graduate Fellowship (to A.C.), a Siebel Scholars Graduate Fellowship (to T.K.L.), funding through a National Institutes of Health (NIH) Biotechnology Training Grant (to T.K.L.), NIH Director's New Innovator Award DP2OD006466 (to K.C.H.), and National Science Foundation CAREER Award MCB-1149328 (to K.C.H.).
Author information
Authors and Affiliations
Contributions
H.S., A.C., T.K.L., and K.C.H. designed the research; H.S., A.C., and T.K.L. performed the research; H.S. and K.C.H. analyzed the data; and H.S. and K.C.H. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Photographs of agar pad and sample preparation.
(a) Melted agar is poured onto the bottom surface of a Singer PlusPlate and spread evenly by gently tilting and shaking the plate. (b) A flat agar surface is generally achieved when the plate is left on the benchtop to solidify without disturbance. The agar layer is ~2 mm in thickness. (c, d) Bacterial cultures are transferred onto an agar pad using a 96-well replicator pin. (e) After the agar pad absorbs all the liquid from the cultures, a large glass cover slip is used to cover the agar surface. (f) Immersion oil is applied evenly onto the cover slip for oil-immersion objectives.
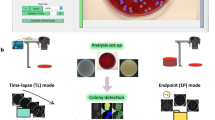
Supplementary Figure 2 Optimization of TTL setup.
(a) Imaging time for a 5x5 grid of images for one strain scales linearly with camera exposure time. Reducing exposure time can expedite image acquisition. (b) Total imaging time for a 5x5 grid is dependent on stage acceleration time. In our configuration, an acceleration time of 40-50 ms is optimal.
Supplementary Figure 3 Variation in mean cellular dimensions calculated from individual fields of view is small.
The coefficient of variation of mean cell width (a) and length (b) across the fields of view from one of the wells in the experiment shown in Fig. 3a was ~3%. Data points are mean ± standard deviation for n > 30, horizontal solid lines are the mean value for cells across all fields of view, and dashed lines are ±5% of the mean. Position #7 happened to have no cells and thus is not included in the plot.
Supplementary information
Supplementary Data
Source code for SLIP software. (ZIP 1596 kb)
Rights and permissions
About this article
Cite this article
Shi, H., Colavin, A., Lee, T. et al. Strain Library Imaging Protocol for high-throughput, automated single-cell microscopy of large bacterial collections arrayed on multiwell plates. Nat Protoc 12, 429–438 (2017). https://doi.org/10.1038/nprot.2016.181
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2016.181
This article is cited by
-
Previously uncharacterized rectangular bacterial structures in the dolphin mouth
Nature Communications (2023)
-
Construction and characterization of a genome-scale ordered mutant collection of Bacteroides thetaiotaomicron
BMC Biology (2022)
-
Imaging-based screens of pool-synthesized cell libraries
Nature Methods (2021)
-
Strain-level epidemiology of microbial communities and the human microbiome
Genome Medicine (2020)
-
High-throughput time-resolved morphology screening in bacteria reveals phenotypic responses to antibiotics
Communications Biology (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.