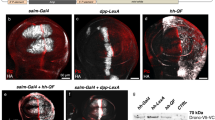
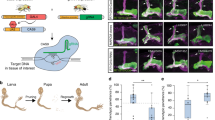
Abstract
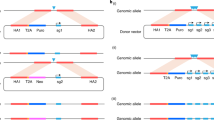
Overexpression screens can be used to explore gene function in Drosophila melanogaster, but to demonstrate their full potential, comprehensive and systematic collections of fly strains are required. Here we provide a protocol for high-throughput cloning of Drosophila open-reading frames (ORFs) that are regulated by upstream activation sequences (UAS sites); the resulting GAL4-inducible UAS-ORF plasmid library is then used to generate Drosophila strains by ΦC31 integrase–mediated site-specific integration. We also provide details for FLP/FRT-mediated in vivo exchange of epitope tags (or regulatory regions) in the ORF library strains, which further extends the potential applications of the library. These transgenic UAS-ORF strains are a useful resource to complement and validate genetic experiments performed with loss-of-function mutants and RNA interference (RNAi) lines. The duration of the complete protocol strongly depends on the number of ORFs required, but embryos can be injected and balanced fly stocks can be established within ∼7–8 weeks for a few genes.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
References
Dietzl, G. et al. A genome-wide transgenic RNAi library for conditional gene inactivation in Drosophila. Nature 448, 151–156 (2007).
Miklos, G.L. & Rubin, G.M. The role of the genome project in determining gene function: insights from model organisms. Cell 86, 521–529 (1996).
Rorth, P. A modular misexpression screen in Drosophila detecting tissue-specific phenotypes. Proc. Natl. Acad. Sci. USA 93, 12418–12422 (1996).
Toba, G. et al. The gene search system. A method for efficient detection and rapid molecular identification of genes in Drosophila melanogaster. Genetics 151, 725–737 (1999).
Bellen, H.J. et al. The Drosophila gene disruption project: progress using transposons with distinctive site specificities. Genetics 188, 731–743 (2011).
Xu, R. et al. A large-scale functional approach to uncover human genes and pathways in Drosophila. Cell Res. 18, 1114–1127 (2008).
Bischof, J. et al. A versatile platform for creating a comprehensive UAS-ORFeome library in Drosophila. Development 140, 2434–2442 (2013).
Schertel, C. et al. Systematic screening of a Drosophila ORF library in vivo uncovers Wnt/Wg pathway components. Dev. Cell 25, 207–219 (2013).
Taxis, C., Stier, G., Spadaccini, R. & Knop, M. Efficient protein depletion by genetically controlled deprotection of a dormant N-degron. Mol. Syst. Biol. 5, 267 (2009).
Hudry, B., Viala, S., Graba, Y. & Merabet, S. Visualization of protein interactions in living Drosophila embryos by the bimolecular fluorescence complementation assay. BMC Biol. 9, 5 (2011).
Prelich, G. Gene overexpression: uses, mechanisms, and interpretation. Genetics 190, 841–854 (2012).
Gibson, T.J., Seiler, M. & Veitia, R.A. The transience of transient overexpression. Nat. Methods 10, 715–721 (2013).
Venken, K.J., Simpson, J.H. & Bellen, H.J. Genetic manipulation of genes and cells in the nervous system of the fruit fly. Neuron 72, 202–230 (2011).
Brown, J.B. et al. Diversity and dynamics of the Drosophila transcriptome. Nature 10.1038/nature12962 (2014).
Yu, C. et al. Development of expression-ready constructs for generation of proteomics libraries. Methods Mol. Biol. 723, 257–272 (2011).
Cavener, D.R. Comparison of the consensus sequence flanking translational start sites in Drosophila and vertebrates. Nucleic Acids Res. 15/4, 1353–1361 (1987).
DeAngelis, M.M., Wang, D.G. & Hawkins, T.L. Solid-phase reversible immobilization for the isolation of PCR products. Nucleic Acids Res. 23, 4742–4743 (1995).
Massouras, A., Decouttere, F., Hens, K. & Deplancke, B. WebPrInSeS: automated full-length clone sequence identification and verification using high-throughput sequencing data. Nucleic Acids Res. 38, W378–W384 (2010).
Stadler, C. et al. Immunofluorescence and fluorescent-protein tagging show high correlation for protein localization in mammalian cells. Nat. Methods 10, 315–323 (2013).
Giaever, G. et al. Functional profiling of the Saccharomyces cerevisiae genome. Nature 418, 387–391 (2002).
Groth, A.C., Fish, M., Nusse, R. & Calos, M.P. Construction of transgenic Drosophila by using the site-specific integrase from phage φC31. Genetics 166, 1775–1782 (2004).
Bateman, J.R., Lee, A.M. & Wu, C.T. Site-specific transformation of Drosophila via φC31 integrase-mediated cassette exchange. Genetics 173, 769–777 (2006).
Venken, K.J., He, Y., Hoskins, R.A. & Bellen, H.J. P[acman]: a BAC transgenic platform for targeted insertion of large DNA fragments in D. melanogaster. Science 314, 1747–1751 (2006).
Bischof, J., Maeda, R.K., Hediger, M., Karch, F. & Basler, K. An optimized transgenesis system for Drosophila using germ-line-specific φC31 integrases. Proc. Natl. Acad. Sci. USA 104, 3312–3317 (2007).
Fish, M.P., Groth, A.C., Calos, M.P. & Nusse, R. Creating transgenic Drosophila by microinjecting the site-specific φC31 integrase mRNA and a transgene-containing donor plasmid. Nat. Protoc. 2, 2325–2331 (2007).
Schlake, T. & Bode, J. Use of mutated FLP recognition target (FRT) sites for the exchange of expression cassettes at defined chromosomal loci. Biochemistry 33, 12746–12751 (1994).
Dahman, C. Drosophila. Methods and Protocols Vol. 420 (Humana Press, 2008).
Meyer, M. & Kircher, M. Illumina sequencing library preparation for highly multiplexed target capture and sequencing. Cold Spring Harb. Protoc. 2010 /10.1101/pdb.prot5448 (2010).
Gloor, G.B. et al. Type I repressors of P element mobility. Genetics 135, 81 (1993) 95.
Saka, Y., Hagemann, A.I., Piepenburg, O. & Smith, J.C. Nuclear accumulation of Smad complexes occurs only after the midblastula transition in Xenopus. Development 134, 4209–4218 (2007).
Acknowledgements
We gratefully acknowledge J. Taipale, who co-initiated the pilot library project. We thank E. Furger and C. Schertel for substantial contributions to molecular cloning, library construction and overexpression analysis; C. Bastos, E. Escher, A. McLeod, S. Miettinen and N. Wang for technical assistance; J.-P. Vincent for FRT details; R. Baumgartner for BiFC reagents; and K. Hens and B. Deplancke for some specific pGW-HA.attB expression clones. This work was supported by the National Center of Competence in Research 'Frontiers in Genetics', the Swiss National Science Foundation, the Kanton of Zürich, the European Research Council and the Scottish Universities Life Sciences Alliance and by the UK Biotechnology and Biological Sciences Research Council (BB/J006424/1).
Author information
Authors and Affiliations
Contributions
K.B. conceived of and coordinated the project, M.B. coordinated and supervised the molecular cloning and data analysis and J.B. coordinated and supervised the creation of the fly library. The experiments were performed by J.B., E.M.S. and M.B., who together wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Bischof, J., Sheils, E., Björklund, M. et al. Generation of a transgenic ORFeome library in Drosophila. Nat Protoc 9, 1607–1620 (2014). https://doi.org/10.1038/nprot.2014.105
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2014.105
This article is cited by
-
Retromer deficiency in Tauopathy models enhances the truncation and toxicity of Tau
Nature Communications (2022)
-
A versatile toolkit for CRISPR-Cas13-based RNA manipulation in Drosophila
Genome Biology (2020)
-
Dysbindin links presynaptic proteasome function to homeostatic recruitment of low release probability vesicles
Nature Communications (2018)
-
Starvation-Induced Dietary Behaviour in Drosophila melanogaster Larvae and Adults
Scientific Reports (2015)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.