Abstract
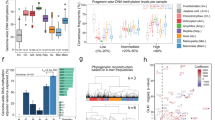
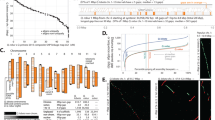
Little is known about patterns of genic DNA methylation across the plant kingdom or about the evolutionary processes that shape them. To characterize gene-body methylation (gbM) within exons, we have gathered single-base resolution methylome data that span the phylogenetic breadth of land plants. We find that a basal land plant, Marchantia polymorpha, lacks any evident signal of gbM within exons, but conifers have high levels of both CG and CHG (where H is A, C or T) methylation in expressed genes. To begin to understand the evolutionary forces that shape gbM, we first tested for correlations in methylation levels across orthologues1,2. Genic CG methylation levels, but not CHG or CHH levels, are correlated across orthologues for species as distantly related as ferns and angiosperms. Hence, relative levels of CG methylation are a consistent property across genes, even for species that diverged ∼400 million years ago3,4. In contrast, genic CHG methylation correlates with genome size, suggesting that the host epigenetic response to transposable elements also affects genes. Altogether, our data indicate that the evolutionary forces acting on DNA methylation vary substantially across species, genes and methylation contexts.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Takuno, S. & Gaut, B. S. Gene body methylation is conserved between plant orthologs and is of evolutionary consequence. Proc. Natl Acad. Sci. USA 110, 1797–1802 (2013).
Seymour, D. K., Koenig, D., Hagmann, J., Becker, C. & Weigel, D. Evolution of DNA methylation patterns in the Brassicaceae is driven by differences in genome organization. PLoS Genet. 10, e1004785 (2014).
Schneider, H. et al. Ferns diversified in the shadow of angiosperms. Nature 428, 553–557 (2004).
Magallon, S., Hilu, K. W. & Quandt, D. Land plant evolutionary timeline: gene effects are secondary to fossil constraints in relaxed clock estimation of age and substitution rates. Am. J. Bot. 100, 556–573 (2013).
Law, J. A. & Jacobsen, S. E. Establishing, maintaining and modifying DNA methylation patterns in plants and animals. Nature Rev. Genet. 11, 204–220 (2010).
Slotkin, R. K. & Martienssen, R. Transposable elements and the epigenetic regulation of the genome. Nature Rev. Genet. 8, 272–285 (2007).
Cokus, S. J. et al. Shotgun bisulphite sequencing of the Arabidopsis genome reveals DNA methylation patterning. Nature 452, 215–219 (2008).
Lister, R. et al. Highly integrated single-base resolution maps of the epigenome in Arabidopsis. Cell 133, 523–536 (2008).
Greaves, I. K. et al. Trans chromosomal methylation in Arabidopsis hybrids. Proc. Natl Acad. Sci. USA 109, 3570–3575 (2012).
Diez, C. M., Roessler, K. & Gaut, B. S. Epigenetics and plant genome evolution. Curr. Opin. Plant Biol. 18C, 1–8 (2013).
Zemach, A., McDaniel, I. E., Silva, P. & Zilberman, D. Genome-wide evolutionary analysis of eukaryotic DNA methylation. Science 328, 916–919 (2010).
Bird, A. P. DNA methylation and the frequency of CpG in animal DNA. Nucleic Acids Res. 8, 1499–1504 (1980).
Rabinowicz, P. D. et al. Differential methylation of genes and repeats in land plants. Genome Res. 15, 1431–1440 (2005).
Neale, D. B. et al. Decoding the massive genome of loblolly pine using haploid DNA and novel assembly strategies. Genome Biol. 15, R59 (2014).
Schmitz, R. J. et al. Epigenome-wide inheritance of cytosine methylation variants in a recombinant inbred population. Genome Res. 23, 1663–1674 (2013).
Regulski, M. et al. The maize methylome influences mRNA splice sites and reveals widespread paramutation-like switches guided by small RNA. Genome Res. 23, 1651–1662 (2013).
The Amborella genome and the evolution of flowering plants. Science 342, 1241089 (2013).
Moissiard, G. et al. Transcriptional gene silencing by Arabidopsis microrchidia homologues involves the formation of heteromers. Proc. Natl Acad. Sci. USA 111, 7474–7479 (2014).
Zhang, X. et al. Genome-wide high-resolution mapping and functional analysis of DNA methylation in Arabidopsis. Cell 126, 1189–1201 (2006).
Alonso, C., Perez, R., Bazaga, P. & Herrera, C. M. Global DNA cytosine methylation as an evolving trait: phylogenetic signal and correlated evolution with genome size in angiosperms. Front Genet. 6, 4 (2015).
Tenaillon, M. I., Hollister, J. D. & Gaut, B. S. A triptych of the evolution of plant transposable elements. Trends Plant Sci. 15, 471–478 (2010).
Symonds, M. R. E. & Blomberg, S. P. in Modern phylogenetic comparative methods and their application in evolutionary biology (ed. Garamszegi, L. Z. ) 105–130 (Springer, 2014).
Zemach, A. et al. The Arabidopsis nucleosome remodeler DDM1 allows DNA methyltransferases to access H1-containing heterochromatin. Cell 153, 193–205 (2013).
Gent, J. I. et al. Accessible DNA and relative depletion of H3K9me2 at maize loci undergoing RNA-directed DNA methylation. Plant Cell 26, 4903–4917 (2014).
Miura, A. et al. An Arabidopsis jmjC domain protein protects transcribed genes from DNA methylation at CHG sites. EMBO J. 28, 1078–1086 (2009).
Ji, L., Neumann, D. A. & Schmitz, R. J. Crop epigenomics: identifying, unlocking, and harnessing cryptic variation in crop genomes. Mol. Plant (2015).
Jones, P. A. Functions of DNA methylation: islands, start sites, gene bodies and beyond. Nature Rev. Genet. 13, 484–492 (2012).
Kim, M. Y. & Zilberman, D. DNA methylation as a system of plant genomic immunity. Trends Plant Sci. 19, 320–326 (2014).
Coleman-Derr, D. & Zilberman, D. Deposition of histone variant H2A.Z within gene bodies regulates responsive genes. PLoS Genet. 8, e1002988 (2012).
Roudier, F., Teixeira, F. K. & Colot, V. Chromatin indexing in Arabidopsis: an epigenomic tale of tails and more. Trends Genet. 25, 511–517 (2009).
Schiex, T., Gouzy, J., Moisan, A. & de Oliveira, Y. FrameD: a flexible program for quality check and gene prediction in prokaryotic genomes and noisy matured eukaryotic sequences. Nucleic Acids Res. 31, 3738–3741 (2003).
Salzberg, S. L., Delcher, A. L., Kasif, S. & White, O. Microbial gene identification using interpolated Markov models. Nucleic Acids Res. 26, 544–548 (1998).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Krueger, F. & Andrews, S. R. Bismark: a flexible aligner and methylation caller for Bisulfite-Seq applications. Bioinformatics 27, 1571–1572 (2011).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nature Methods 9, 357–359 (2012).
Finet, C., Timme, R. E., Delwiche, C. F. & Marletaz, F. Multigene phylogeny of the green lineage reveals the origin and diversification of land plants. Curr. Biol. 20, 2217–2222 (2010).
Lartillot, N., Lepage, T. & Blanquart, S. PhyloBayes 3: a Bayesian software package for phylogenetic reconstruction and molecular dating. Bioinformatics 25, 2286–2288 (2009).
Huelsenbeck, J. P. & Suchard, M. A. A nonparametric method for accommodating and testing across-site rate variation. Syst. Biol. 56, 975–987 (2007).
Lartillot, N. & Philippe, H. A Bayesian mixture model for across-site heterogeneities in the amino-acid replacement process. Mol. Biol. Evol. 21, 1095–1109 (2004).
Bennett, M. D. & Leitch, I. J. Plant DNA C-values database (release 6.0). (2012).
O'Brien, K. P., Remm, M. & Sonnhammer, E. L. Inparanoid: a comprehensive database of eukaryotic orthologs. Nucleic Acids Res. 33, D476–D480 (2005).
Acknowledgements
The authors would like to thank J. Bousquet (University of Laval, Canada), J. Der (Cal State Fullerton, USA), C. Depamphilis (Penn State University, USA), K. Ishizaki (Kobe University, Japan), T. Kohchi (Kyoto University, Japan), C. Langley (University of California Davis, USA) and D. Stevenson (New York Botanical Garden, USA) for plant DNA and tissue. R. Gaut (University of Californa Irvine, USA) generated the BSseq libraries; C. Finet (University of Lyon, France) shared the alignment of amino acid sequences from his study; A. Delcher (John Hopkins University, USA) provided Glimmer software. J. A. Fawcett (Graduate University for Advanced Studies, Japan), A. Bousios (University of Sussex, UK), S. Hug, K. Roessler, Q. Liu and A. Gonzalez-Gonzalez provided helpful suggestions. S.T. is supported by an internal grant from SOKENDAI and JSPS Grant-in-Aid for Young Scientists (B) (grant no. 15K18585); J.-H.R. is supported by the National Natural Science Foundation of China (grant no. 31330008) and B.S.G. is supported by NSF Grant IOS-1542703.
Author information
Authors and Affiliations
Contributions
J.-H.R. and B.S.G. conceived of the project; S.T. and J.-H.R. analysed data; S.T., J.-H.R. and B.S.G. wrote the paper. All authors approved the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
About this article
Cite this article
Takuno, S., Ran, JH. & Gaut, B. Evolutionary patterns of genic DNA methylation vary across land plants. Nature Plants 2, 15222 (2016). https://doi.org/10.1038/nplants.2015.222
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/nplants.2015.222
This article is cited by
-
The pattern of alternative splicing and DNA methylation alteration and their interaction in linseed (Linum usitatissimum L.) response to repeated drought stresses
Biological Research (2023)
-
DDM1-mediated gene body DNA methylation is associated with inducible activation of defense-related genes in Arabidopsis
Genome Biology (2023)
-
The methylation landscape of giga-genome and the epigenetic timer of age in Chinese pine
Nature Communications (2023)
-
The Torreya grandis genome illuminates the origin and evolution of gymnosperm-specific sciadonic acid biosynthesis
Nature Communications (2023)
-
DNA Methylation is Involved in Sex Determination in Spinach
Biochemical Genetics (2023)