Abstract
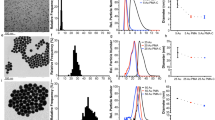
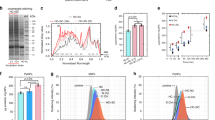
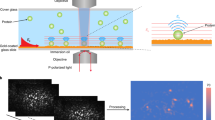
Nanoparticles in a biological milieu are known to form a sufficiently long-lived and well-organized ‘corona’ of biomolecules to confer a biological identity to the particle. Because this nanoparticle–biomolecule complex interacts with cells and biological barriers, potentially engaging with different biological pathways, it is important to clarify the presentation of functional biomolecular motifs at its interface. Here, we demonstrate that by using antibody-labelled gold nanoparticles, differential centrifugal sedimentation and various imaging techniques it is possible to identify the spatial location of proteins, their functional motifs and their binding sites. We show that for transferrin-coated polystyrene nanoparticles only a minority of adsorbed proteins exhibit functional motifs and the spatial organization appears random, which is consistent, overall, with a stochastic and irreversible adsorption process. Our methods are applicable to a wide array of nanoparticles and can offer a microscopic molecular description of the biological identity of nanoparticles.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Monopoli, M. P., Åberg, C., Salvati, A. & Dawson, K. A. Biomolecular coronas provide the biological identity of nanosized materials. Nature Nanotech. 7, 779–786 (2012).
Cedervall, T. et al. Understanding the nanoparticle–protein corona using methods to quantify exchange rates and affinities of proteins for nanoparticles. Proc. Natl Acad. Sci. USA 104, 2050–2055 (2007).
Nel, A. E. et al. Understanding biophysicochemical interactions at the nano–bio interface. Nature Mater. 8, 543–557 (2009).
Tenzer, S. et al. Rapid formation of plasma protein corona critically affects nanoparticle pathophysiology. Nature Nanotech. 8, 772–781 (2013).
Hellstrand, E. et al. Complete high-density lipoproteins in nanoparticle corona. FEBS J. 276, 3372–3381 (2009).
Wan, S. et al. The ‘sweet’ side of the protein corona: effects of glycosylation on nanoparticle–cell interactions. ACS Nano 9, 2157–2166 (2015).
Casals, E., Pfaller, T., Duschl, A., Oostingh, G. J. & Puntes, V. Time evolution of the nanoparticle protein corona. ACS Nano 4, 3623–3632 (2010).
Maiorano, G. et al. Effects of cell culture media on the dynamic formation of protein–nanoparticle complexes and influence on the cellular response. ACS Nano 4, 7481–7491 (2010).
Walkey, C. D. & Chan, W. C. W. Understanding and controlling the interaction of nanomaterials with proteins in a physiological environment. Chem. Soc. Rev. 41, 2780–2799 (2012).
Walczyk, D., Baldelli Bombelli, F., Monopoli, M. P., Lynch, I. & Dawson, K. A. What the cell ‘sees’ in bionanoscience. J. Am. Chem. Soc. 132, 5761–5768 (2010).
Fleischer, C. & Payne, C. K. Secondary structure of corona proteins determines the cell surface receptors used by nanoparticles. J. Phys. Chem. B 118, 14017–14026 (2014).
Treuel, L. et al. Impact of protein modification on the protein corona on nanoparticles and nanoparticle–cell interactions. ACS Nano 8, 503–513 (2014).
Zensi, A. et al. Albumin nanoparticles targeted with Apo E enter the CNS by transcytosis and are delivered to neurones. J. Control. Release 137, 78–86 (2009).
Deng, Z. J., Liang, M., Monteiro, M., Toth, I. & Minchin, R. F. Nanoparticle-induced unfolding of fibrinogen promotes Mac-1 receptor activation and inflammation. Nature Nanotech. 6, 39–44 (2011).
Jiang, X. et al. Quantitative analysis of the protein corona on FePt nanoparticles formed by transferrin binding. J. R. Soc. Interface 7, S5–S13 (2010).
Martel, J. et al. Comprehensive proteomic analysis of mineral nanoparticles derived from human body fluids and analyzed by liquid chromatography–tandem mass spectrometry. Anal. Biochem. 418, 111–125 (2011).
Sund, J., Alenius, H., Vippola, M., Savolainen, K. & Puustinen, A. Proteomic characterization of engineered nanomaterial–protein interactions in relation to surface reactivity. ACS Nano 5, 4300–4309 (2011).
Mortimer, G. M. et al. Cryptic epitopes of albumin determine mononuclear phagocyte system clearance of nanomaterials. ACS Nano 8, 3357–3366 (2014).
Milani, S., Baldelli Bombelli, F., Pitek, A. S., Dawson, K. A. & Rädler, J. Reversible versus irreversible binding of transferrin to polystyrene nanoparticles: soft and hard corona. ACS Nano 6, 2532–2541 (2012).
Pitek, A. S. et al. Transferrin coated nanoparticles: study of the bio–nano interface in human plasma. PLoS ONE 7, e40685 (2012).
Slot, J. W. & Geuze, H. J. A new method of preparing gold probes for multiple-labeling cytochemistry. Eur. J. Cell Biol. 38, 87–93 (1985).
Hancock, J. F. & Prior, I. A. Electron microscopic imaging of Ras signaling domains. Methods 37, 165–172 (2005).
Cheng, Y., Zak, O., Aisen, P., Harrison, S. C. & Walz, T. Structure of the human transferrin receptor–transferrin complex. Cell 116, 565–576 (2004).
Mason, A. B. et al. A loop in the N-lobe of human serum transferrin is critical for binding to the transferrin receptor as revealed by mutagenesis, isothermal titration calorimetry, and epitope mapping. J. Mol. Recogn. 22, 521–529 (2009).
Carney, R. P. et al. Determination of nanoparticle size distribution together with density or molecular weight by 2D analytical ultracentrifugation. Nature Commun. 2, 335 (2011).
Wang, J., Tian, S., Petros, R. A., Napier, M. E. & DeSimone, J. M. The complex role of multivalency in nanoparticles targeting the transferrin receptor for cancer therapies. J. Am. Chem. Soc. 132, 11306–11313 (2010).
Hansen, J. P. & McDonald, I. R. Theory of Simple Liquids 3rd edn (Academic Press, 2006).
Dawson, K. A. The glass paradigm for colloidal glasses, gels, and other arrested states driven by attractive interactions. Curr. Opin. Colloid Interface Sci. 7, 218–227 (2002).
Salvati, A. et al. Transferrin-functionalized nanoparticles lose their targeting capabilities when a biomolecule corona adsorbs on the surface. Nature Nanotech. 8, 137–143 (2013).
Berman, H. M. et al. The protein data bank. Nucleic Acids Res. 28, 235–242 (2000).
Pettersen, E. F. et al. UCSF chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Acknowledgements
This work was supported by a Science Foundation Ireland (SFI) Principal Investigator Award (agreement no. 12/IA/1422). P.M.K. acknowledges the Irish Research Council for funding under the Embark scheme (RS/2011/106). The authors also acknowledge the small collaborative project NanoTransKinetics (NMP4-2010-EU-US-266737) funded by the European Union Seventh Framework Programme. This work was based on work supported by NanoSolutions (NMP-L6-2012-309329) and NAMDIATREAM (Nanotechnological Toolkit for Multi-modal Disease Diagnostics and Treatment Monitoring; NMP4-LA-2010-246479), both funded by the European Commission Seventh Framework Programme. The authors acknowledge support from Mass Spectrometry Resource and Conway Core Technologies, based at the Conway Institute of Biomolecular and Biomedical Research (University College Dublin), and the Advanced Microscopy Laboratory at the Centre for Research on Adaptive Nanostructures and Nanodevices (Trinity College Dublin) for access to advanced electron microscopy facilities.
Author information
Authors and Affiliations
Contributions
P.M.K. designed and performed the experiments, collected, analysed and interpreted the data, and contributed to writing the manuscript. C.Å. constructed a theoretical framework to describe the experimental results and created the pairwise distributions, interpreted the data and contributed to writing the manuscript. E.P. contributed to the biological interaction experiments. A.O'C. performed the advanced electron microscopy analysis. J.C. contributed to TEM imaging. J.F. contributed to particle counting and image analysis. Z.K. contributed to the design of the experiments, oversaw the acquisition and interpretation of the microscopy images, and contributed to writing the manuscript. K.A.D. conceived and designed the experiments, interpreted the data and wrote the paper. All authors discussed the results and commented on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary information
Supplementary information (PDF 4891 kb)
Supplementary information
Supplementary Movie 1 (WMV 4070 kb)
Rights and permissions
About this article
Cite this article
Kelly, P., Åberg, C., Polo, E. et al. Mapping protein binding sites on the biomolecular corona of nanoparticles. Nature Nanotech 10, 472–479 (2015). https://doi.org/10.1038/nnano.2015.47
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nnano.2015.47
This article is cited by
-
Passive, active and endogenous organ-targeted lipid and polymer nanoparticles for delivery of genetic drugs
Nature Reviews Materials (2023)
-
Predicting protein function and orientation on a gold nanoparticle surface using a residue-based affinity scale
Nature Communications (2022)
-
Current understanding of biological identity at the nanoscale and future prospects
Nature Nanotechnology (2021)
-
Unusual zymogen activation patterns in the protein corona of Ca-zeolites
Nature Catalysis (2021)
-
Nanoscale characterization of the biomolecular corona by cryo-electron microscopy, cryo-electron tomography, and image simulation
Nature Communications (2021)