Abstract
Evidence has begun to emerge for microRNAs as regulators of synaptic signaling, specifically acting to control postsynaptic responsiveness during synaptic transmission. In this report, we provide evidence that Drosophila melanogaster miR-1000 acts presynaptically to regulate glutamate release at the synapse by controlling expression of the vesicular glutamate transporter (VGlut). Genetic deletion of miR-1000 led to elevated apoptosis in the brain as a result of glutamatergic excitotoxicity. The seed-similar miR-137 regulated VGluT2 expression in mouse neurons. These conserved miRNAs share a neuroprotective function in the brains of flies and mice. Drosophila miR-1000 showed activity-dependent expression, which might serve as a mechanism to allow neuronal activity to fine-tune the strength of excitatory synaptic transmission.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Hébert, S.S. et al. Loss of microRNA cluster miR-29a/b-1 in sporadic Alzheimer's disease correlates with increased BACE1/β-secretase expression. Proc. Natl. Acad. Sci. USA 105, 6415–6420 (2008).
Delay, C., Mandemakers, W. & Hebert, S.S. MicroRNAs in Alzheimer's disease. Neurobiol. Dis. 46, 285–290 (2012).
Mouradian, M.M. MicroRNAs in Parkinson's disease. Neurobiol. Dis. 46, 279–284 (2012).
Gascon, E. & Gao, F.B. Cause or effect: misregulation of microRNA pathways in neurodegeneration. Front. Neurosci. 6, 48 (2012).
Haramati, S. et al. miRNA malfunction causes spinal motor neuron disease. Proc. Natl. Acad. Sci. USA 107, 13111–13116 (2010).
Gehrke, S., Imai, Y., Sokol, N. & Lu, B. Pathogenic LRRK2 negatively regulates microRNA-mediated translational repression. Nature 466, 637–641 (2010).
Kim, J. et al. A microRNA feedback circuit in midbrain dopamine neurons. Science 317, 1220–1224 (2007).
Karres, J.S., Hilgers, V., Carrera, I., Treisman, J. & Cohen, S.M. The conserved microRNA miR-8 tunes atrophin levels to prevent neurodegeneration in Drosophila. Cell 131, 136–145 (2007).
Liu, N. et al. The microRNA miR-34 modulates ageing and neurodegeneration in Drosophila. Nature 482, 519–523 (2012).
Kosik, K.S. The neuronal microRNA system. Nat. Rev. Neurosci. 7, 911–920 (2006).
McNeill, E. & Van Vactor, D. MicroRNAs Shape the Neuronal Landscape. Neuron 75, 363–379 (2012).
Schratt, G.M. et al. A brain-specific microRNA regulates dendritic spine development. Nature 439, 283–289 (2006).
Saba, R. et al. Dopamine-regulated microRNA MiR-181a controls GluA2 surface expression in hippocampal neurons. Mol. Cell. Biol. 32, 619–632 (2012).
Harraz, M.M., Eacker, S.M., Wang, X., Dawson, T.M. & Dawson, V.L. MicroRNA-223 is neuroprotective by targeting glutamate receptors. Proc. Natl. Acad. Sci. USA 109, 18962–18967 (2012).
Hu, Z. et al. Neurexin and neuroligin mediate retrograde synaptic inhibition in C. elegans. Science 337, 980–984 (2012).
Cohen, J.E., Lee, P.R., Chen, S., Li, W. & Fields, R.D. MicroRNA regulation of homeostatic synaptic plasticity. Proc. Natl. Acad. Sci. USA 108, 11650–11655 (2011).
Kim, D.S. et al. Bilateral enhancement of excitation via up-regulation of vesicular glutamate transporter subtype 1, not subtype 2, immunoreactivity in the unilateral hypoxic epilepsy model. Brain. Res. 1055, 122–130 (2005).
Touret, M., Parrot, S., Denoroy, L., Belin, M.F. & Didier-Bazes, M. Glutamatergic alterations in the cortex of genetic absence epilepsy rats. BMC Neurosci. 8, 69 (2007).
Helton, T.D., Otsuka, T., Lee, M.C., Mu, Y. & Ehlers, M.D. Pruning and loss of excitatory synapses by the parkin ubiquitin ligase. Proc. Natl. Acad. Sci. USA 105, 19492–19497 (2008).
Blandini, F. An update on the potential role of excitotoxicity in the pathogenesis of Parkinson's disease. Funct. Neurol. 25, 65–71 (2010).
Raudensky, J. & Yamamoto, B.K. Effects of chronic unpredictable stress and methamphetamine on hippocampal glutamate function. Brain Res. 1135, 129–135 (2007).
Weng, R., Chen, Y.W., Bushati, N., Cliffe, A. & Cohen, S.M. Recombinase-mediated cassette exchange provides a versatile platform for gene targeting: knockout of miR-31b. Genetics 183, 399–402 (2009).
Bilen, J. & Bonini, N.M. Drosophila as a model for human neurodegenerative disease. Annu. Rev. Genet. 39, 153–171 (2005).
Kretzschmar, D., Hasan, G., Sharma, S., Heisenberg, M. & Benzer, S. The swiss cheese mutant causes glial hyperwrapping and brain degeneration in Drosophila. J. Neurosci. 17, 7425–7432 (1997).
Daniels, R.W., Gelfand, M.V., Collins, C.A. & DiAntonio, A. Visualizing glutamatergic cell bodies and synapses in Drosophila larval and adult CNS. J. Comp. Neurol. 508, 131–152 (2008).
Reisberg, B. et al. Memantine in moderate-to-severe Alzheimer's disease. N. Engl. J. Med. 348, 1333–1341 (2003).
Rothman, S.M. & Olney, J.W. Excitotoxicity and the NMDA receptor–still lethal after eight years. Trends Neurosci. 18, 57–58 (1995).
Ibáñez-Ventoso, C., Vora, M. & Driscoll, M. Sequence relationships among C. elegans, D. melanogaster and human microRNAs highlight the extensive conservation of microRNAs in biology. PLoS ONE 3, e2818 (2008).
Liguz-Lecznar, M. & Skangiel-Kramska, J. Vesicular glutamate transporters (VGLUTs): the three musketeers of glutamatergic system. Acta Neurobiol. Exp. (Warsz.) 67, 207–218 (2007).
Boulland, J.L. et al. Expression of the vesicular glutamate transporters during development indicates the widespread corelease of multiple neurotransmitters. J. Comp. Neurol. 480, 264–280 (2004).
Takamori, S. VGLUTs: 'exciting' times for glutamatergic research? Neurosci. Res. 55, 343–351 (2006).
Smrt, R.D. et al. MicroRNA miR-137 regulates neuronal maturation by targeting ubiquitin ligase mind bomb-1. Stem Cells 28, 1060–1070 (2010).
Daniels, R.W., Miller, B.R. & DiAntonio, A. Increased vesicular glutamate transporter expression causes excitotoxic neurodegeneration. Neurobiol. Dis. 41, 415–420 (2011).
Mark, K.A., Quinton, M.S., Russek, S.J. & Yamamoto, B.K. Dynamic changes in vesicular glutamate transporter 1 function and expression related to methamphetamine-induced glutamate release. J. Neurosci. 27, 6823–6831 (2007).
Tordera, R.M., Pei, Q. & Sharp, T. Evidence for increased expression of the vesicular glutamate transporter, VGLUT1, by a course of antidepressant treatment. J. Neurochem. 94, 875–883 (2005).
Willemsen, M.H. et al. Chromosome 1p21.3 microdeletions comprising DPYD and MIR137 are associated with intellectual disability. J. Med. Genet. 48, 810–818 (2011).
Geekiyanage, H. & Chan, C. MicroRNA-137/181c regulates serine palmitoyltransferase and in turn amyloid β, novel targets in sporadic Alzheimer's disease. J. Neurosci. 31, 14820–14830 (2011).
Schizophrenia Psychiatric Genome-Wide Association Study (GWAS) Consortium. Genome-wide association study identifies five new schizophrenia loci. Nat. Genet. 43, 969–976 (2011).
Kwon, E., Wang, W. & Tsai, L.H. Validation of schizophrenia-associated genes CSMD1, C10orf26, CACNA1C and TCF4 as miR-137 targets. Mol. Psychiatry 18, 11–12 (2013).
Gong, W.J. & Golic, K.G. Ends-out, or replacement, gene targeting in Drosophila. Proc. Natl. Acad. Sci. USA 100, 2556–2561 (2003).
Chen, Y.W., Weng, R. & Cohen, S.M. Protocols for use of homologous recombination gene targeting to produce microRNA mutants in Drosophila. Methods Mol. Biol. 732, 99–120 (2011).
Brennecke, J., Hipfner, D.R., Stark, A., Russell, R.B. & Cohen, S.M. bantam encodes a developmentally regulated microRNA that controls cell proliferation and regulates the pro-apoptotic gene hid in Drosophila. Cell 113, 25–36 (2003).
Acknowledgements
We thank G. Wright (Imaging Facility, Institute of Molecular Biology) and C. Libedinsky for help with image analysis, K. Ryoo (Korea Institute of Science and Technology) for help with lentiviral depletion assays, S. Tsuda for help with the electrophysiology set-up, S. Aw and L. Foo for discussion, and E. Lee for advice on statistics. K.J. Tan and S. Lim provided technical support. This work was supported by IMCB (SC laboratory) and Nanyang Technological University and University of Warwick (A.T. laboratory), as well as Competitive Research Program (CRP) funds from National Research Foundation, Singapore, and by the World Class Institute (WCI) Program of the National Research Foundation of Korea (NRF) funded by the Ministry of Education, Science and Technology of Korea (MEST) (NRF Grant Number: WCI 2009-003) (G.J.A. laboratory).
Author information
Authors and Affiliations
Contributions
P.V. performed all experiments. M.-R.A. and A.T. contributed to the mouse lentiviral injection experiments. P.V., G.J.A. and S.M.C. designed the experiments and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
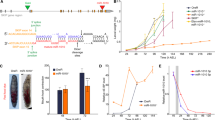
Supplementary Figure 1 Strategy for targeted knockout of mir-1000
miR-1000 is located in the first intron of the msi gene, but represents an independent transcription unit, transcribed from the opposite strand of the DNA. miR-1000 deletion mutants were generated by targeted homologous recombination, as described (ref 41). Left and right homology arms (blue bars) were amplified by PCR from genomic DNA (diagram not to scale). The 4436bp left arm was amplified with: LF-ACGAAGGTAACCATATCTCAGACC, LR-TTAACATTGTTGAGCGTCATTAGC; the 4199bp right arm was amplified with: RF-CAGCACTTTATTCAATTCACTTGG and RR-AATACTGTGGCGACTTTAGGTAGG.
Following recombination, 141 bp spanning the miRNA hairpin was deleted and replaced with a mini-white cassette (red). The presence of the intron-containing mini-white marker is expected to disrupt msi function. The mini-white marker in the miR-1000 allele is flanked by loxP sites (purple triangles) and was excised by crossing the mutant to heat shock-CRE flies. After CRE mediated excision, a single LoxP site is left in the msi intron. Genetic complementation tests showed that msi function was not compromised in the mini-white–excised miR-1000 mutant strains (miR-1000KO1), or in the mini-white–excised miR-1000 RMCE alleles (miR-1000KO2 depicted in Fig 1c). Deletion of the miRNA in the various mutants was confirmed by genomic DNA PCR and by microRNA quantitative-PCR. All genetic tests were done using flies carrying two independently generated alleles or an allele in trans to a chromosomal deletion (Df(3R)Exel6203) removing the miR-1000 locus.
Supplementary Figure 2 Embryo hatching, pupariation and adult eclosion rate
(a) Embryo hatching rate for the indicated genotypes. KO1 indicates the mini-white excised simple deletion described in Figure S1. KO2 indicates the mini-white excised RMCE deletion allele. Rescue indicates animals homozygous for the miRNA recue allele in the RMCE background (depicted in Fig 1c). Data represent the average of three independent experiments with n = 150 embryos each. Error bars: ± s.d. p-value of all mutant combination (KO1/Df, KO2/Df and KO1/KO2) is < 0.01 when compared to control and rescue.
(b) Percent pupariation indicates completion of larval development to reach the pupal stage (black bars). Percent eclosion indicates survival of the pupa to adult (grey bars). Data represent the average of three independent experiments with n = 50 larvae each. Error bar: ± s.d.
Supplementary Figure 3 Vacuole formation in the mir-1000 mutant brain
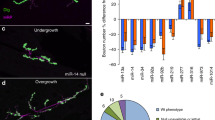
Confocal microscope images of 10D old adult brains from miR-1000KO1/miR-1000KO2 mutant flies, labeled with antibody to Discs large protein (Dlg). Left panel: overview. Olfactory lobes and mushroom bodies are prominently labeled. Center and right panels show magnified views of olfactory lobes. Arrows indicate vacuoles. Scale bars = 10 μm.
Supplementary Figure 4 Oxidative stress sensitivity in mir-1000 mutant
Flies were reared on medium supplemented with 2.5% H2O2. KO1/KO2 mutant flies (n = 100), Rescue denotes homozygous RMCE rescue allele (n = 100), KO2/+ heterozygous control (n = 100), yw control (n = 80). p < 0.0001 comparing mutant and rescue and comparing mutant with each control (Kaplan Meier analysis using the log rank test).
Supplementary Figure 5 mRNA expression for predicted mir-1000 targets
miR-1000 targets were predicted using Targetscan (www.targetscan.org). Selected targets were tested by quantitative real-time PCR for changes in RNA level comparing KO1/KO2 mutant, control and RMCE rescue mutant brains. Data were normalized to Ribosomal Protein rp49 and to the yw control and represent the average of three independent experiment ± s.d.
Only VGlut showed the expected pattern for a functional target: upregulation in mutant that is restored toward normal in the rescued mutant.
Nplp1: neuropeptide-like precursor 1
Pdk1: Phosphoinositide-dependent kinase 1
Rogdi: a leucine zipper protein involved in learning and memory.
NMDA1: N-methyl-D-aspartate receptor-associated protein, the Drosophila NMDA-type Glutamate receptor orthologue.
slgA: a proline catabolic enzyme which has been reported to affect locomotion.
VGlut: Vesicular Glutamate transporter
EAAT2: Excitatory Amino Acid Transporter 2
Supplementary Figure 6 Rescue by reduced AMPA-type glutamate receptor activity
S6a: Effect of removing one copy of the GluR1 gene on lifespan
Longevity of miR-1000 mutant (KO1/KO2) was improved by removing one copy of the AMPA-type Glutamate receptor gene GluR1 (PBac(WH)Glu-RIf05411). Kaplan Meier analysis using Log rank test: p < 0.0001. The GluR1 allele was recombined onto the KO2 mutant chromosome.
S6b: Effect of removing one copy of the GluR1 gene on climbing.
Climbing behavior of miR-1000 mutant (KO1/KO2) was improved by removing one copy of the AMPA-type Glutamate receptor gene GluRI.
S6c: Effect of removing one copy of the GluR1 gene on apoptosis
Quantification of apoptotic cells in 10 day-old brains of the indicated genotypes. KO2, GluRI is the homozygous double mutant chromosome. GluR1 is the homozygous GluR1 mutant. ANOVA: p < 0.0001 comparing KO2/KO2 with KO2, GluR1 and the other genotypes. control n = 10 brains; n ≥ 14 other genotypes
Supplementary Figure 7 Rescue by reduced metabotropic glutamate receptor mGluR activity
(a) Effect of removing one copy of the, mGluRA on lifespan
Longevity of miR-1000 mutant (KO1/KO2) was improved by removing one copy of the metabotropic Glutamate receptor gene mGluRA. p < 0.0001 comparing KO1/KO2 mutant with KO1/KO2 mutant carrying one copy of the mGluRA mutant (Log rank test).
(b) Effect of removing one copy of the mGluRA on climbing
Climbing behaviour of miR-1000 mutant was improved by removing one copy of mGluRA gene. p < 0.001 comparing KO1/KO2 mutant with KO1/KO2 carrying one copy of mGluRA mutant.
(c) Effects of RNAi mediated depletion of mGluR
Two different UAS-RNAi transgenes were used to deplete the metabotropic Glutamate receptor mGluR. A Gal4 insertion at the mGluRA locus was used to direct transgene expression. mGluR-RNAi-1: P(GD707)v1793, mGluR-RNAi-2: P(GD707)v1794. RNAi transgenes were recombined onto the KO2 chromosome. Gal4 driven RNAi expression significantly improved lifespan: p < 0.0001 comparing KO1/KO2;mGluR-GAL4/+ with and without mGluR-RNAi-1 or mGluR-RNAi-2 (log rank test). Note the partial rescue from the UAS-RNAi, KO2 recombinants as homozygotes without Gal4. This was due to leaky expression of the RNAi transgenes (panel S7d).
(d) RNAi transgene efficiency
Leaky expression of the mGluRRNAi transgenes shown by quantitative PCR.
mGluR mRNA was reduced by both transgenes when present in 2 copies without a Gal4 driver. Gal4 directed expression of the RNAi transgenes further reduced mGluR levels. Data were normalized to rp49 and to the yw control flies.
(e) Effect of RNAi-mediated depletion mGluR on climbing
Climbing behavior of miR-1000 (KO1/KO2) was improved by RNAi-mediated depletion of the metabotropic Glutamate receptor mGluR. Note the partial rescue from leaky expression of the mGluRRNAi transgenes. Addition of Gal4 provided a modest further improvement. Data: average of at least four independent experiments. ANOVA: p < 0.0001 comparing KO1/KO2;mGluR-GAL4 with KO2, mGluRRNAi 1/KO1; mGluR-GAL4 and KO2, mGluRRNAi 2/KO1;mGluR-GAL4.
(f) Effect of reducing mGluR on apoptosis
Quantification of apoptotic cells in 10 day-old brains of the indicated genotypes. ANOVA: p < 0.0001 comparing KO2/KO2 with all the other genotypes. Control n = 10 brains; n ≥ 14 other genotypes. This data set was collected in the same experiment as the data shown in Fig S6c. The control and KO2/KO2 samples are the same.
(g) Effect of reducing GluRIIA and GluRIIB on climbing
Left: KO1/KO2 mutant carrying one copy of Df(2L)Exel8016, which removes GluRIIA and GluRIIB. Data represent the average of 3 independent experiments. t-test: ns.
Supplementary Figure 8 mEJPs in mutants with reduced glutamate receptor activity
(a) Representative traces of spontaneous mEJPs at the NMJ from larvae of genotypes miR-1000 mutant (KO1/KO2), with reduced NMDAR1 expression (NMDAR, KO2/KO1), with reduced GluR1 expression (GluR1, KO2/KO1) and with reduced mGluRA expression (KO2/KO1; mGluR).
(b) Cumulative probability distribution of mEJP amplitudes from genotypes indicated in A.
(c) Median amplitude of mEJPs of the mutant (KO1/KO2) compared to mutant with reduced levels of receptors as in A. Significant rescue in mEJP amplitude was observed in miR-1000 mutant with reduced mGluR level (ANOVA: p = 0.004 comparing KO1/KO2 with KO1/KO2;mGluR).
(d) mEJP frequency of genotypes as in A. ANOVA: differences not significant.
Supplementary Figure 9 Regulation of the mouse VGluT2 3′ UTR by Mir-137
Luciferase activity of a reporter containing the mouse VGlut2 3′UTR. Expression of mouse miR-137 or fly miR-1000 reduced activity of the intact reporter (p = 0.0006 comparing the intact 3′ UTR to the control for miR-137; p = 0.0037 for miR-1000). Regulation was lost when the miR-137 site was mutated (p = 0.0006 comparing the intact and mutant 3′ UTR for miR-137; p = 0.0064 for miR-1000; The difference between control and mutant 3′ UTRs were not significant). The mutations introduced into the miR-137 sites in the mouse UTR are shown in red. Data represent the average of 5 independent experiments ± s.d. Data were analyzed by ANOVA using the Holm-Sidak multiple comparison test.
Supplementary Figure 11 mir-1000 and VGlut expression in the visual system
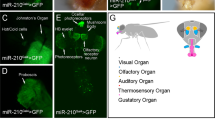
(a) Optical section showing miR-1000 expression using the GFP knock-in allele (green) and VGlut protein (red) in the retina of adult eye. Scale bar = 50 µm.
(b) Optical section showing miR-1000 expression (green) and VGlut-GAL4 driving UAS-CD8-RFP (red) in the lamina and medulla layers of the adult optic lobe. The lamina receives axonal projections from photoreceptors R1-R6 and the medulla receives projections from photoreceptor R7-R8. Scale bar = 50 µm.
(c) Optical section showing miR-1000 expression (green) and VGlut-GAL4 (red) in medulla and lobula layers of the adult optic lobe. Scale bar = 50 µm.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–11 (PDF 1197 kb)
Rights and permissions
About this article
Cite this article
Verma, P., Augustine, G., Ammar, MR. et al. A neuroprotective role for microRNA miR-1000 mediated by limiting glutamate excitotoxicity. Nat Neurosci 18, 379–385 (2015). https://doi.org/10.1038/nn.3935
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nn.3935
This article is cited by
-
A neuroprotective role of Ufmylation through Atg9 in the aging brain of Drosophila
Cellular and Molecular Life Sciences (2023)
-
Prediction of P-tau/Aβ42 in the cerebrospinal fluid with blood microRNAs in Alzheimer’s disease
BMC Medicine (2021)
-
The conserved microRNA miR-210 regulates lipid metabolism and photoreceptor maintenance in the Drosophila retina
Cell Death & Differentiation (2021)
-
The conserved microRNA miR-34 regulates synaptogenesis via coordination of distinct mechanisms in presynaptic and postsynaptic cells
Nature Communications (2020)
-
MicroRNAs as modulators of longevity and the aging process
Human Genetics (2020)