Abstract
The subsurface biosphere is largely unexplored and contains a broad diversity of uncultured microbes1. Despite being one of the few prokaryotic lineages that is cosmopolitan in both the terrestrial and marine subsurface2–4, the physiological and ecological roles of SAGMEG (South-African Gold Mine Miscellaneous Euryarchaeal Group) Archaea are unknown. Here, we report the metabolic capabilities of this enigmatic group as inferred from genomic reconstructions. Four high-quality (63–90% complete) genomes were obtained from White Oak River estuary and Yellowstone National Park hot spring sediment metagenomes. Phylogenomic analyses place SAGMEG Archaea as a deeply rooting sister clade of the Thermococci, leading us to propose the name Hadesarchaea for this new Archaeal class. With an estimated genome size of around 1.5 Mbp, the genomes of Hadesarchaea are distinctly streamlined, yet metabolically versatile. They share several physiological mechanisms with strict anaerobic Euryarchaeota. Several metabolic characteristics make them successful in the subsurface, including genes involved in CO and H2 oxidation (or H2 production), with potential coupling to nitrite reduction to ammonia (DNRA). This first glimpse into the metabolic capabilities of these cosmopolitan Archaea suggests they are mediating key geochemical processes and are specialized for survival in the subsurface biosphere.
Similar content being viewed by others
South-African Gold Mine Miscellaneous Euryarchaeal Group (SAGMEG) Archaea thrive in a broad range of anoxic subsurface environments and temperature regimes (4 to 80 °C)5. They were first identified in the terrestrial subsurface and have since been shown to be broadly distributed in marine sediments2,3. To begin to resolve their ecological roles and sample a broad diversity within the group, we reconstructed genomic bins from the White Oak River (WOR, North Carolina) estuary and from Yellowstone National Park (YNP) hot spring sediments (Supplementary Fig. 1). Two genomic bins were reconstructed (using emergent self-organizing map (ESOM) mapping of tetranucleotide signatures, see Supplementary Information for additional details) from the WOR estuary sediment (DG-33 and DG-33–1, ∼63% complete) from the anoxic methane-rich layer (53–54 cmbsf (centimetres below surface))6. Two additional genomic bins (YNP_N21 and YNP_45, 80% and 90% complete, respectively) were obtained from a microaerophilic (0.45 mg per litre O2) YNP hot spring (68.8 °C, pH 8.6). This spring is relatively high in sulphate (25 ppm) and low in sulphide (0.27 ppm) and nitrogen species (nitrate and nitrite were below detection, ammonium was 1.25 × 10−4 ppm). All four genomic bins were manually curated (see Methods) to remove potential contaminant sequences (Table 1). Based on the size and completeness of the bins we estimated the average size of all these genomes to be around 1.5 Mbp. The average gene size is relatively small (716 (DG-33-1), 688 (DG-33), 821 (bin YNP_45) and 820 (YNP_N21) bp) compared to the average prokaryote (∼920 bp7). These observations indicate that SAGMEG genomes have been subjected to genome streamlining, possibly as a result of nutrient limitation8.
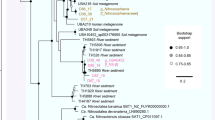
Phylogenetic identification of the genomic bins based on 16S rRNA genes (excluding DG-33-1, for which no such gene was recovered) revealed that they represent distinct lineages within the SAGMEG clade (Fig. 1). Both of the YNP bins are closely related to clones that were previously recovered from Obsidian Pool at YNP5, while WOR bin DG-33 is similar to rRNA sequences from subseafloor sediments (Fig. 1). rRNA gene phylogeny places SAGMEG Archaea as a basal lineage within the Euryarchaeota. We were not able to resolve the branching position of SAGMEG within the Euryarchaeota solely on the basis of rRNA phylogenies. The placement of the group is also inconsistent in the literature4,9. Therefore, to investigate the phylogenetic position of SAGMEG within the Archaea in more detail, we constructed phylogenetic trees based on concatenated ribosomal proteins (Supplementary Fig. 2), and more comprehensively using 48 phylogenetically conserved markers (Fig. 2 and Supplementary Fig. 3). These analyses revealed that SAGMEG Archaea form a distinct clade within the Euryarchaeota that represents a sister group to the class Thermococci. To reflect their affiliation with the subsurface biosphere, we propose that they be given a new class name, ‘Hadesarchaea’.
Bin DG-33-1 lacked a gene long enough to accurately resolve its phylogeny. Taxa labelled in blue are from the terrestrial subsurface, orange are from the oceanic subsurface, green are from deep-sea sediments, purple are from shallow aquatic sediments, and red are from terrestrial hot springs. The phylogeny was generated based on 1,682 unambiguous characters using the RAxML (maximum likelihood) method with gamma rate distribution model, in the ARB software package47. Bootstrap values were generated using UFBoot48 with 10,000 replicates; those >95 are designated with filled circles and those >70 are shown as open circles. The numbers of sequences present in the collapsed groups are labelled on the wedges.
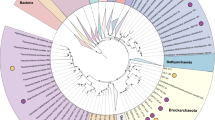
Bayesian phylogeny of Hadesarchaea (SAGMEG) and other representative Archaeal phyla using 48 concatenated arCOGs (a total of 9,982 unambiguous characters). The tree was constructed from two independent Phylobayes chains after running for more than 4,000 generations until convergence (maxdiff of 0.15) and the consensus tree was obtained after burn-in of the first 2,500 generations and subsampling every tenth tree. Both maximum likelihood bootstrap (top) and Bayesian posterior probability (below) values are shown. The numbers of sequences present in the collapsed groups are labelled on the wedges. Bacteria were used as an outgroup (not shown).
The first hints into the metabolic capabilities of the Hadesarchaea came from analyses of the stable isotopic composition of Archaeal cells (Bathyarchaeota, Thermoplasmatales, SAGMEG and Lokiarchaeota) in deep-sea sediments, which indicate consistently that all members of this Archaeal community incorporate organic carbon, suggesting a heterotrophic lifestyle3. Comparative genomic analyses of the obtained genome bins reveal that Hadesarchaea share several carbon pathways with other Euryarchaeota members. We could not identify genes encoding methyl-CoM reductase, indicating that Hadesarchaea are probably not involved in methanogenesis in a conventional manner. However, they do have a variety of genes for carbon metabolism present in methanogens and other Euryarchaeal lineages. All four Hadesarchaeal genome bins lack most of the genes for the forward and reverse tricarboxylic acid (TCA) cycle (or the glycoxylate bypass) for the production of acetyl-CoA. Instead, they contain genes for the C1 pathway (the carbon monoxide dehydrogenase pathway), which is typically used by methanogens and numerous other anaerobes for CO2 fixation or production10.
The YNP genomic bins (YNP_45 and YNP_N21) lack several genes for the CO dehydrogenase pathway, including those encoding formylmethanofuran-tetrahydromethanopterin N-formyltransferase (ftr), methenyltetrahydromethanopterin cyclohydrolase (mch) and methylenetetrahydromethanopterin dehydrogenase (mtd), while the WOR bin DG-33 encodes all three (Fig. 3). However, bin YNP_N21 has a mch gene, suggesting that the absence of these genes may be a result of the incomplete draft assemblies that were obtained. Alternatively, both YNP genomic bins have D-3-phosphoglycerate dehydrogenase, phosphoserine phosphatase and glycine hydroxymethyltransferase, which may be used to generate 5,10-methylene-THMPT from glycerate 3-phosphate (as illustrated for bin YNP_45 in Fig. 3). This is referred to as the glycine pathway, and is thought to be an ancestral form of carbon fixation11. However, it is possible that these C1-pathway genes are used in reverse for CO2 production12.
Black arrows represent genes that were identified in bin YNP_45 and YNP_N21. Grey arrows are those that were not found in the YNP bins, but are present in WOR bin DG-33-1. DNRA, Denitrification and dissimilatory nitrite reduction to ammonia; FDH, format dehydrogenase; Hdr, membrane-type heterodisulphide hydrogenases; F420, coenzyme F420; V, complex V ATPase; COX, carbon-monoxide dehydrogenase; Ni,FeIII, Ni,Fe-dependent hydrogenases; Nit, nitrite reductase; Hyd, cytoplasmic hydrogenases; Nuo, NADH dehydrogenases; Sdh, succinate dehydrogenases; Znu, zinc transporter; Liv, a branched amino-acid transporter; Mal and Msm, monosaccharide transporters; PST, phosphate transporter.
The most complete genome bins YNP_45 and YNP_N21 lack the key glycolytic hexokinase, glucokinase (pfkC) and pyruvate kinase (pyk) genes, resulting in an incomplete glycolytic pathway. They do have genes encoding pyruvate:ferredoxin oxidoreductases (porA) and fructose-1,6-bisphosphatases (fbp), indicating that these Archaea contain a complete pathway for gluconeogenesis. With the exception of bin DG-33, all of the genomic bins contain a gene encoding a ribulose biphosphate carboxylase (RubisCO) homologue (Fig. 3). To confirm that the genes are related to known RubisCOs, we generated a phylogenetic tree of these proteins (Supplementary Fig. 4). This analysis revealed that the Hadesarchaeal RubisCOs form a distinct clade within Archaeal RubisCO type III proteins, which were recently shown to be involved in AMP metabolism in other Euryarchaea13. However, the genomes of previously described Archaea lacked Calvin–Benson–Bassham (CBB) cycles, which is the pathway for carbon fixation using RubisCO13. Interestingly, YNP_45 and YNP_N21 have near-complete CBB pathways, only lacking two CBB genes: phosphoribulokinase (prk) and fructose-1,6-bisphosphatase II (glpX). The ability to fix inorganic carbon opens up a new source of carbon in the deep subsurface, where dissolved organic carbon has been shown to be extremely low14,15 but inorganic carbon is readily available16.
Acetate and hydrogen constitute an important link between respiration and fermentative processes in the subsurface17. Hydrogen is believed to be a primary substrate for microbial growth in the subsurface and in hot springs5,18,19. Hadesarchaea constitute a significant proportion of microbial communities in hydrogen-rich hot springs at YNP5, and all genome bins contain genes encoding acetyl-CoA synthetase for the conversion of acetate into acetyl-CoA. In addition, these bins also contain genes encoding several hydrogenases, including Ni-dependent hydrogenase, Ni,Fe hydrogenase and coenzyme F420 hydrogenase. Ni,Fe hydrogenases are capable of both H2 consumption and production, and all the Hadesarchaea genomic bins contain the genes encoding the small and large subunits, G-components, as well as maturation factors. It is possible that the Ni,Fe hydrogenases, F420-hydrogen dehydrogenase and cytochrome b5 genes (also present in all four genomic bins) are used to derive energy by electron transfer from H2, as demonstrated in other Euryarchaeota20. We did not identify any Fe-hydrogenase encoding genes, which are generally assumed to be associated with H2 production and have only been identified in bacteria20, in the Hadesarchaeal genome bins.
Interestingly, some of these hydrogenases have been demonstrated to be involved in sulphur reduction (sulphidogenesis)21 and sulphide oxidation22. The YNP genomic bins YNP_45 and YNP_N21 contain phylogenetically distinct genes that are related to sulphohydrogenase (Supplementary Fig. 5). They also encode the nuoB, nuoH and ndhI genes of the NADH:quinone oxidoreductase, which are essential for transferring electrons from NADH to S0 (ref. 21). As these two Archaeal bins are associated with a hot sulphur spring environment, it seems likely that these genes have a role in sulphur cycling. Cultured species in the phylogenetically related sister group Thermococci are commonly shown to be reducing S0 to sulphide23.
Carbon monoxide is a common gas in hydrothermal environments24 and is produced by thermal decomposition of organic matter or as a side product of some anaerobes25. It is therefore likely to occur widely in sediments and the subsurface. It has been estimated that after sulphate reduction, CO oxidation is the most energetically favourable anaerobic reaction in the deep terrestrial subsurface15. In addition to carbon monoxide (cox) genes, the YNP_45 Hadesarchaea bin also has distinct genes for catalytic nickel-containing CO dehydrogenase (CooS) and the iron sulphur subunits (CooF), suggesting that they are able to oxidize CO and H2O to CO2 and H2 as an energy source. The other bins are missing cooS, but have genes with sequence similarity to cooF present in Thermococcus spp. Phylogenetic analyses of these proteins show they are affiliated with ‘clade E’ (Supplementary Fig. 6)12 and related to other methanogenic Euryarchaeota. The CooF sequences of Thermococcus spp, which also belong to this clade, have the ability to couple anaerobic CO oxidation to hydrogen production using cooS genes26. If the Hadesarchaea are deriving energy from the oxidation of CO, then the C1 pathway and hydrogenases are probably producing CO2 and H227.
CO oxidation coupled to anaerobic nitrate reduction is an energetically favourable reaction28 that has previously been demonstrated in bacteria29. Potentially, Hadesarchaea may be coupling CO oxidation to the reduction of nitrite to ammonia. We identified a distinct nitrite reductase in bin YNP_N21 (Supplementary Figs 7). Electron acceptors such as nitrite are generally thought to be depleted in the subsurface16, but it has been shown that the deep terrestrial subsurface contains considerable concentrations of nitrate that may be derived from radiolytic sources30. Thus, it seems feasible that these nitrite reductases can be used to derive energy.
The current understanding of the diversity, biology and ecology of the Archaea is very limited, when considering how few of the known phyla have been cultured or genomically explored, especially considering that many of these enigmatic lineages are predominant in nature. In the current study, we have obtained the first genomic sequences of a widespread group related to Euryarchaeota4, here named Hadesarchaea. Inference of their metabolic capabilities indicates that they are probably deriving energy from the oxidation of carbon monoxide, which may be coupled to H2O or nitrite reduction in the terrestrial subsurface. They contain a variety of central carbon metabolic (C1 pathway) genes found in methanogens, which may be used for carbon fixation. The reconstruction of these novel genomes expands our knowledge of the evolutionary histories and metabolic potential of these widespread Archaea. Their metabolic features encoded in relatively streamlined genomes make them prominent members of the deep dark subsurface biosphere.
Methods
Sample collection and processing
The estuary genomic libraries were obtained as described previously31. Six 1 m plunger cores were collected from a water depth of ∼1.5 m in three mid-estuary locations (two cores per site) of WOR, North Carolina, in October 2010 (site 1 at 34°44.592 N, 77°07.435 W; site 2 at 34°44.482 N, 77°07.404 W; site 3 at 34°44.141 N, 77°07298 W). Cores were stored at 4 °C overnight and processed 24 h after sampling6. Each core was sectioned into 2 cm intervals. From each site, one core was subsampled for geochemical analyses, and the other core was subsampled for DNA extractions. The sulphate, sulphide and methane profiles of these cores indicated a distinct sulphur and methane-cycling zonation. The sulphate-reducing zone (SRZ) started in sediment layers where sulphate was abundant and sulphide just started to accumulate (8–12 cm). The methane-producing zone (MPZ) was characterized by accumulating methane (a single sample from site 1 of this zone was taken at 52–54 cm (ref. 6). DNA was extracted from these sediments using the UltraClean Mega Soil DNA Isolation Kit (MoBio), using 6 g of sediment, and stored at −80 °C until use.
Hot spring sediment samples from YNP were collected from a hot spring in Lower Culex Basin (from a site named as 10Y13 at 44°34.23 N, 110°47.41 W) as described previously32. This uncharacterized hot spring has a temperature around 70 °C and pH of 8.6, and the hot spring sediments appeared light brown in colour. Sediment samples were taken using 50 ml falcon tubes, preserved in sucrose lysis buffer and kept at −20 °C until further use. DNA used for the metagenomic library was extracted from ∼500 mg of sediment using cetyltrimethyl ammonium bromide (CTAB)/phenol/chloroform method33.
Single-cell genomics
Cell fractions were obtained from the Yellowstone hot spring sediment sample using Nycodenz density gradient centrifugation34, and sent to the Bigelow Laboratory for Ocean Sciences for flow sorting and amplification to generate single-cell amplified genomes (SAGs) as described previously32. A 384-well microtitre plate was then screened for Archaea using a LightCycler 480 qPCR instrument (Roche) with SYBR Green I Master kit and utilizing an Archaea-specific primer pair, A519F (5′-CAGCMGCCGCGGTAA-3′) and A1041R (5′-GGCCATGCACCWCCTCTC-3′). A SAG positive for the Archaeal 16S rRNA gene sequence was selected (identified by sequencing its 16S rDNA using the Sanger sequencing method), a sequencing library was prepared using NexteraXT library preparation kit (Illumina), and the library was paired-end sequenced (2 × 300 bp) on a MiSeq instrument (Illumina). 16S rRNA amplification and Sanger sequencing of the multiple displacement amplication (MDA)-amplified DNA from the cell yielded one phylotype belonging to SAGMEG. After sequencing, no contaminants were detected based on BLASTn comparison of contigs with the NCBI nt database. The number of high-quality sequences generated from the MiSeq run was 3,260,839, representing a total of 0.5 Gbp of sequence data, which were then assembled with SPAdes version 3.5.035 using the following parameters: --sc --careful -k 21,33,55,75,101,125.
Genomic assembly, binning and annotation
Illumina sequencing adapters from the read pairs (2 × 150 bp) were removed from both YNP and WOR libraries using Scythe (https://github.com/vsbuffalo/scythe), and low-quality reads (Q < 30) were trimmed with Sickle (https://github.com/najoshi/sickle). Reads with three or more Ns or with average quality score of less than Q20 and a length <50 bps were removed. Trimmed, screened, paired-end Illumina reads from WOR were assembled using IDBA-UD36 with the following parameters (--pre_correction --mink 55 --maxk 95 --step 10 --seed_kmer 55). To maximize assembly, reads from different sites were co-assembled. The WOR assembly was generated from high-quality reads (378,027,948, average read length 124 bp and average insert 284 bp) of site 1 (52–54 cm) from WOR as described previously31.
The YNP hot spring assembly consisted of 666,428,020 high-quality paired-end Illumina reads representing a total of 86.4 Gbp of sequence data with an estimated insert size of 337 bp. Assembly was performed using the IDBA-UD assembler36 with the following parameter: --maxk 124. Contigs of genes of particular interest were checked for chimaeras by looking for dips in coverage within read mappings. Initial binning of the assembled fragments was performed using tetra-nucleotide frequency signatures using 5 kbp fragments of the contigs37. The ESOMs were manually delineated and curated based on clusters within the map (as shown in Supplementary Figs 8 and 9). Coverage was enhanced by recruiting reads (from each individual library/sample) to scaffolds by BLASTN (bitscore >75), which was then normalized to the number of reads from each library.
The accuracy of the binning was then assessed by checking the genomic bins to which each of the 5 kbp subportions of the contigs were assigned. If the contig was >15 kbp then the contig was assigned to the bin that contained the majority (>50%) of its 5 kbp subportions. The completeness, contamination and strain heterogeneity of the genomes within bins were then estimated using CheckM38. Some of the fragments were identified as contaminants based on their phylogenetic placement, and were manually removed. Closely related variants of a lineage were retained in a bin. Binning was also manually curated based on GC content, top blast hits and mate-pairings.
Examination of fragments identified as contaminants in the YNP_N21 bin revealed that CheckM38 might be inflating contamination levels. CheckM identifies multiple copies of genes and then calls the fragment they are on as contaminants. We believe in the case of bin YNP_N21 that gene duplications are indeed apparent in the genome. For example, ornithine carbamoyltransferase was identified to be present on four different large (12–71 kb) contigs that are all clearly Archaeal and have tetranucleotide signature binning, blast taxa matching and GC content consistent with other Hadesarchaea. This suggests that these are multiple copies of that gene in the genome, and not contaminants. CheckM also identified seven copies in the bin as a gene on five different contigs; four of these hypothetical are present on two contigs (two on each of the contigs). These sequences are considerably different (only 26–35% similarity at the protein level) and have unique hits to genes in GenBank, including hypotheticals, diguanylate cyclases and dihydroorotate dehydrogenases. The estimate that this bin has 10% contamination is thus potentially overestimated. Removal of four relatively low-coverage contigs (NODE_10_length_48481_cov_4.44764_ID_19, NODE_22_length_22948_cov_6.32857_ID_43, NODE_32_length_12588_cov_4.53992_ID_63 and NODE_35_length_10923_cov_3.69392_ID_69) resulted in the reduction of the double copy markers from six to one. However, examination of these contigs revealed they are definitely Archaeal and related to other Hadesarchaea bins present in this study. We therefore decided to leave them in the bin and have flagged them as potential contaminants in the public databases.
Co-assembly of the Yellowstone SAGMEG (YNP_N21) SAG and metagenomic bin was performed by first extracting metagenomic reads mapped against the YNP_N21 bin contigs using bwa39 and samtools40. This resulted in an improved assembly compared with that obtained just from the SAG reads (Supplementary Table 1 and Supplementary Fig. 10). The read coverage in contigs was calculated using bedtools41. Extracted reads were then co-assembled with SAG reads using SPAdes (version 3.5.0)35 with the same parameters as the single-cell genome assembly. Resulting contigs larger than 5 kbp were manually checked for contamination and removed.
Genes were called using Prodigal42. Initially gene functions were assigned by comparison to the KEGG database43. The called functions were finalized by comparisons with functional databases, including PFAM44 and arCOG45. Proteins with important ecological or geochemical relevance were selected for protein alignments using MUSCLE46 and phylogenetic comparisons with top BLAST hits in NCBI.
Phylogenetic analyses
The 16S rRNA gene trees were generated by running the RAxML (v2.2.1) tool implemented within the ARB software package (v2.5b)47. Bootstrap values were generated using UFBoot48 with 10,000 replicates. The concatenated ribosomal protein tree was generated using 16 syntenic genes that have been shown to undergo limited lateral gene transfer (rpL2, 3, 4, 5, 6, 14, 15, 16, 18, 22, 24 and rpS3, 8, 10, 17, 19)49. Amino acid alignments of the individual ribosomal proteins were generated using MUSCLE46 and trimmed using BMGE50 (with the following settings: -m BLOSUM30 –g 0.5). The curated alignments were then concatenated for phylogenetic analyses, generated by running the RAxML (v2.2.1) tool implemented within the ARB software package (v2.5b)47.
A total of 48 arCOGs (Archaeal cluster of orthologous genes45) previously determined to be universally present in single copies51 were identified and extracted from the four SAGMEG genome bins using PSI-BLAST51. The 48 reference arCOGs and identified orthologues from the genome bins were aligned using MAFFT (mafft-linsi option)52. Alignments were trimmed for poorly aligned regions and gaps using the BMGE tool50 (with the following settings: -m BLOSUM30 –g 0.5) and concatenated before phylogenomic inferences were perfomed using both maximum likelihood and Bayesian methods. Maximum likelihood analysis was performed using IQ-Tree53 with the C60+LG mixture model for 1,000 ultra-rapid bootstraps and Bayesian inference was carried out using Phylobayes54 with the CAT+GTR+Г mixture model of amino acid evolution for about 4,000 generations until convergence (maxdiff of 0.15, burnin of 2,500 generations and subsampling every tenth tree) for two of the independent chains out of the four.
Accession numbers
The genomes supporting the results of this article are available in NCBI Genbank under the BioProjectID PRJNA298487 (LQMP00000000 and LQMQ00000000 for the YNP genomic bins), and BioProjectID PRJNA270657 (LQHI00000000 and LQHJ00000000 for the WOR genomic bins).
References
Teske, A. & Sorensen, K. B. Uncultured Archaea in deep marine subsurface sediments: have we caught them all? ISME J. 2, 3–18 (2008).
Parkes, R. J. et al. Deep sub-seafloor prokaryotes stimulated at interfaces over geological time. Nature 436, 390–394 (2005).
Biddle, J. F. et al. Heterotrophic Archaea dominate sedimentary subsurface ecosystems off Peru. Proc. Natl Acad. Sci. USA 103, 3846–3851 (2006).
Takai, K., Moser, D. P., DeFlaun, M., Onstott, T. C. & Fredrickson, J. K. Archaeal diversity in waters from deep South African gold mines. Appl. Environ. Microbiol. 67, 5750–5760 (2001).
Spear, J. R., Walker, J. J., McCollom, T. M. & Pace, N. R. Hydrogen and bioenergetics in the Yellowstone geothermal ecosystem. Proc. Natl Acad. Sci. USA 102, 2555–2560 (2005).
Lazar, C. S. et al. Environmental controls on intragroup diversity of the uncultured benthic Archaea of the miscellaneous Crenarchaeotal group lineage naturally enriched in anoxic sediments of the White Oak River estuary (North Carolina, USA). Environ. Microbiol. 17, 2228–2238 (2015).
Baker, B. J. et al. Enigmatic, ultrasmall, uncultivated Archaea. Proc. Natl Acad. Sci. USA 107, 8806–8811 (2010).
Giovannoni, S. J. et al. Genome streamlining in a cosmopolitan oceanic bacterium. Science 309, 1242–1245 (2005).
Offre, P., Spang, A. & Schleper, C. Archaea in biogeochemical cycles. Annu. Rev. Microbiol. 67, 437–457 (2013).
Vornolt, J., Kunow, J., Stetter, K. & Thauer, R. Enzymes and coenzymes of the carbon monoxide dehydrogenase pathway for autotrophic CO2 fixation in Archaeoglobus lithotrophicus and the lack of carbon monoxide dehydrogenase in the heterotrophic A. profundus. Arch. Microbiol. 163, 112–118 (1995).
Braakman, R. & Smith, E. The emergence and early evolution of biological carbon-fixation. PLoS Comput. Biol. 8, e1002455 (2012).
Techtmann, S., Colman, A. S., Lebedinsky, A. V., Sokolova, T. G. & Robb, F. T. Evidence for horizontal gene transfer of anaerobic carbon monoxide dehydrogenases. Front. Microbiol. 3, 132 (2012).
Sato, T., Atomi, H. & Imanaka, T. Archaeal type III RuBisCOs function in a pathway for AMP metabolism. Science 315, 1003–1006 (2007).
Onstott, T. C. et al. The origin and age of biogeochemical trends in deep fracture water of the Witwatersrand Basin, South Africa. Geomicrobiol. J. 23, 369–414 (2006).
Magnabosco, C. et al. A metagenomic window into carbon metabolism at 3 km depth in Precambrian continental crust. ISME J. http://dx.doi.org/10.1038/ismej.2015.150 (2015).
D'Hondt, S. et al. Distributions of microbial activities in deep subseafloor sediments. Science 306, 2216–2221 (2004).
Oremland, R. S. & Polcin, S. Methanogenesis and sulfate reduction: competitive and noncompetitive substrates in estuarine sediments. Appl. Environ. Microbiol. 44, 1270–1276 (1982).
Stevens, T. O. & McKinley, J. P. Lithoautotrophic microbial ecosystems in deep basalt aquifers. Science 270, 450–455 (1995).
Wrighton, K. C. et al. Fermentation, hydrogen, and sulfur metabolism in multiple uncultivated bacterial phyla. Science 337, 1661–1665 (2012).
Vignais, P. M. & Billoud, B. Occurrence, classification, and biological function of hydrogenases: an overview. Chem. Rev. 107, 4206–4272 (2007).
Schut, G. J., Bridger, S. L. & Adams, M. W. W. Insights into the metabolism of elemental sulfur by the hyperthermophilic Archaeon Pyrococcus furiosus: characterization of a coenzyme a-dependent NAD(P)H sulfur oxidoreductase. J. Bacteriol. 189, 4431–4441 (2007).
Ma, K., Schicho, R. N., Kelly, R. M. & Adams, M. W. Hydrogenase of the hyperthermophile Pyrococcus furiosus is an elemental sulfur reductase or sulfhydrogenase: evidence for a sulfur-reducing hydrogenase ancestor. Proc. Natl Acad. Sci. USA 90, 5341–5344 (1993).
Kanai, T. et al. Distinct physiological roles of the three [NiFe]-hydrogenase orthologs in the hyperthermophilic Archaeon Thermococcus kodakarensis. J. Bacteriol. 193, 3109–3116 (2011).
Kochetkova, T. et al. Anaerobic transformation of carbon monoxide by microbial communities of Kamchatka hot springs. Extremophiles 15, 319–325 (2011).
Sokolova, T. G. et al. Diversity and ecophysiological features of thermophilic carboxydotrophic anaerobes. FEMS Microbiol. Ecol. 68, 131–141 (2009).
Lim, J. K., Kang, S. G., Lebedinsky, A. V., Lee, J.-H. & Lee, H. S. Identification of a novel class of membrane-bound [NiFe]-hydrogenases in Thermococcus onnurineus NA1 by in silico analysis. Appl. Environ. Microbiol. 76, 6286–6289 (2010).
Sipma, J., Lens, P. N. L., Stams, A. J. M. & Lettinga, G. Carbon monoxide conversion by anaerobic bioreactor sludges. FEMS Microbiol. Ecol. 44, 271–277 (2003).
Frunzke, K. & Meyer, O. Nitrate respiration, denitrification, and utilization of nitrogen sources by aerobic carbon monoxide-oxidizing bacteria. Arch. Microbiol. 154, 168–174 (1990).
King, G. M. Nitrate-dependent anaerobic carbon monoxide oxidation by aerobic CO-oxidizing bacteria. FEMS Microbiol. Ecol. 56, 1–7 (2006).
Silver, B. J. et al. The origin of NO3− and N2 in deep subsurface fracture water of South Africa. Chem. Geol. 294–295, 51–62 (2012).
Baker, B. J., Lazar, C., Teske, A. & Dick, G. J. Genomic resolution of linkages in carbon, nitrogen, and sulfur cycling among widespread estuary sediment bacteria. Microbiome 3, 14 (2015).
Saw, J. H. et al. Exploring microbial dark matter to resolve the deep archaeal ancestry of eukaryotes. Phil. Trans. R. Soc. Lond. B 370, 20140328 (2015).
Mitchell, K. & Takacs-Vesbach, C. A comparison of methods for total community DNA preservation and extraction from various thermal environments. J. Ind. Microbiol. Biotechnol. 35, 1139–1147 (2008).
Dodsworth, J. A. et al. Single-cell and metagenomic analyses indicate a fermentative and saccharolytic lifestyle for members of the OP9 lineage. Nature Commun. 4, 1854 (2013).
Bankevich, A. et al. SPAdes: a new genome assembly algorithm and its applications to single-cell sequencing. J. Comput. Biol. 19, 455–477 (2012).
Peng, Y., Leung, H. C. M., Yiu, S. M. & Chin, F. Y. L. IDBA-UD: a de novo assembler for single-cell and metagenomic sequencing data with highly uneven depth. Bioinformatics 28, 1420–1428 (2012).
Dick, G. J. et al. Community-wide analysis of microbial genome sequence signatures. Genome Biol. 10, R85 (2009).
Parks, D. H., Imelfort, M., Skennerton, C. T., Hugenholtz, P. & Tyson, G. W. CheckM: assessing the quality of microbial genomes recovered from isolates, single cells, and metagenomes. Genome Res. 25, 1043–1055 (2015).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Hyatt, D. et al. Prodigal: prokaryotic gene recognition and translation initiation site identification. BMC Bioinformatics 11, 119 (2010).
Moriya, Y., Itoh, M., Okuda, S., Yoshizawa, A. C. & Kanehisa, M. KAAS: an automatic genome annotation and pathway reconstruction server. Nucleic Acids Res. 35, W182–W185 (2007).
Finn, R. D. et al. Pfam: the protein families database. Nucleic Acids Res. 42, D222–D230 (2014).
Makarova, K. S., Wolf, Y. I. & Koonin, E. V. Archaeal clusters of orthologous genes (arCOGs): an update and application for analysis of shared features between thermococcales, methanococcales, and methanobacteriales. Life 5, 818–840 (2015).
Edgar, R. C. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 32, 1792–1797 (2004).
Ludwig, W. et al. ARB: a software environment for sequence data. Nucleic Acids Res. 32, 1363–1371 (2004).
Minh, B. Q., Nguyen, M. A. T. & von Haeseler, A. Ultrafast approximation for phylogenetic bootstrap. Mol. Biol. Evol. 30, 1188–1195 (2013).
Sorek, R. et al. Genome-wide experimental determination of barriers to horizontal gene transfer. Science 318, 1449–1452 (2007).
Criscuolo, A. & Gribaldo, S. BMGE (Block Mapping and Gathering with Entropy): a new software for selection of phylogenetic informative regions from multiple sequence alignments. BMC Evol. Biol. 10, 210 (2010).
Altschul, S. F., Gish, W., Miller, W., Myers, E. W. & Lipman, D. J. Basic local alignment search tool. J. Mol. Biol. 215, 403–410 (1990).
Katoh, K., Misawa, K., Kuma, K. & Miyata, T. MAFFT: a novel method for rapid multiple sequence alignment based on fast Fourier transform. Nucleic Acids Res. 30, 3059–3066 (2002).
Nguyen, L. T., Schmidt, H. A., von Haeseler, A. & Minh, B. Q. IQ-TREE: a fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol. Biol. Evol. 32, 268–274 (2015).
Lartillot, N., Rodrigue, N., Stubbs, D. & Richer, J. PhyloBayes MPI: phylogenetic reconstruction with infinite mixtures of profiles in a parallel environment. System. Biol. 62, 611–615 (2013).
Acknowledgements
The sequencing of the WOR sediment metagenome was funded by the US Department of Energy. The Joint Genome Institute is supported by the Office of Science of the US Department of Energy (contract no. DE-AC02-05CH11231 to B.J.B.). The sequencing of the YNP sediment metagenome was performed by the National Genomics Infrastructure sequencing platforms at the Science for Life Laboratory at Uppsala University, a national infrastructure supported by the Swedish Research Council (VR-RFI) and the Knut and Alice Wallenberg Foundation. The authors thank the Uppsala Multidisciplinary Center for Advanced Computational Science (UPPMAX) at Uppsala University and the Swedish National Infrastructure for Computing (SNIC) at the PDC Center for High-Performance Computing for providing computational resources. Sampling in the WOR estuary was supported by the European Research Council (ERC advanced grant no. 247153-DARCLIFE to K.-U.H.). A.T. was supported by the Center for Dark Energy Biosphere Investigations (C-DEBI). The authors thank D.R. Colman and C. Takacs-Vesbach (University of New Mexico) for collecting the YNP sediment samples used in the current study (under permit #YELL-2010-SCI-5344), and the Yellowstone Center for Resources for their assistance and for facilitating this research. The authors thank G. Reboul for his contributions to the initial stage of this project. This work was supported by a grant of the European Research Council (ERC starting grant no. 310039-PUZZLE_CELL) and the Swedish Foundation for Strategic Research (FFL12-0024 to T.J.G.E.), and by a Marie Curie IIF grant by the European Union (331291-Darkcells to J.H.S.).
Author information
Authors and Affiliations
Contributions
B.J.B., C.S.L., A.P.T., and T.J.G.E. conceived the study. B.J.B., J.H.S. and A.E.L. analysed the genomic data. K.-U.H. provided materials. B.J.B., J.H.S., A.P.T. and T.J.G.E. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Table 1, Figures 1-11 and References (PDF 10291 kb)
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article's Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Baker, B., Saw, J., Lind, A. et al. Genomic inference of the metabolism of cosmopolitan subsurface Archaea, Hadesarchaea. Nat Microbiol 1, 16002 (2016). https://doi.org/10.1038/nmicrobiol.2016.2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/nmicrobiol.2016.2
This article is cited by
-
Influence of Geochemistry in the Tropical Hot Springs on Microbial Community Structure and Function
Current Microbiology (2023)
-
Brockarchaeota, a novel archaeal phylum with unique and versatile carbon cycling pathways
Nature Communications (2021)
-
New approaches for archaeal genome-guided cultivation
Science China Earth Sciences (2021)
-
Contrasting bacterial and archaeal distributions reflecting different geochemical processes in a sediment core from the Pearl River Estuary
AMB Express (2020)
-
Microbial dark matter filling the niche in hypersaline microbial mats
Microbiome (2020)