Abstract
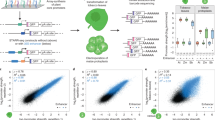
Plant synthetic biology promises immense technological benefits, including the potential development of a sustainable bio-based economy through the predictive design of synthetic gene circuits. Such circuits are built from quantitatively characterized genetic parts; however, this characterization is a significant obstacle in work with plants because of the time required for stable transformation. We describe a method for rapid quantitative characterization of genetic plant parts using transient expression in protoplasts and dual luciferase outputs. We observed experimental variability in transient-expression assays and developed a mathematical model to describe, as well as statistical normalization methods to account for, this variability, which allowed us to extract quantitative parameters. We characterized >120 synthetic parts in Arabidopsis and validated our method by comparing transient expression with expression in stably transformed plants. We also tested >100 synthetic parts in sorghum (Sorghum bicolor) protoplasts, and the results showed that our method works in diverse plant groups. Our approach enables the construction of tunable gene circuits in complex eukaryotic organisms.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Kiani, S. et al. CRISPR transcriptional repression devices and layered circuits in mammalian cells. Nat. Methods 11, 723–726 (2014).
Rinaudo, K. et al. A universal RNAi-based logic evaluator that operates in mammalian cells. Nat. Biotechnol. 25, 795–801 (2007).
Gardner, T.S., Cantor, C.R. & Collins, J.J. Construction of a genetic toggle switch in Escherichia coli. Nature 403, 339–342 (2000).
You, L., Cox, R.S. III, Weiss, R. & Arnold, F.H. Programmed population control by cell-cell communication and regulated killing. Nature 428, 868–871 (2004).
Steeves, T.A. & Sussex, I.M. Patterns in Plant Development 2nd edn. (Cambridge Univ. Press, 1989).
Lucks, J.B., Qi, L., Whitaker, W.R. & Arkin, A.P. Toward scalable parts families for predictable design of biological circuits. Curr. Opin. Microbiol. 11, 567–573 (2008).
Slusarczyk, A.L., Lin, A. & Weiss, R. Foundations for the design and implementation of synthetic genetic circuits. Nat. Rev. Genet. 13, 406–420 (2012).
Kim, J., Klein, P.G. & Mullet, J.E. Synthesis and turnover of photosystem II reaction center protein D1. Ribosome pausing increases during chloroplast development. J. Biol. Chem. 269, 17918–17923 (1994).
Asai, T. et al. MAP kinase signalling cascade in Arabidopsis innate immunity. Nature 415, 977–983 (2002).
Mewes, H.W. et al. Overview of the yeast genome. Nature 387, 7–65 (1997).
Klein, R.M., Wolf, E.D., Wu, R. & Sanford, J.C. High-velocity microprojectiles for delivering nucleic acids into living cells. Nature 327, 70–73 (1987).
Giniger, E., Varnum, S.M. & Ptashne, M. Specific DNA-binding of Gal4, a positive regulatory protein of yeast. Cell 40, 767–774 (1985).
Schnarr, M., Oertelbuchheit, P., Kazmaier, M. & Grangerschnarr, M. DNA-binding properties of the LexA repressor. Biochimie 73, 423–431 (1991).
Wang, S., Chang, Y., Guo, J. & Chen, J.-G. Arabidopsis Ovate Family Protein 1 is a transcriptional repressor that suppresses cell elongation. Plant J. 50, 858–872 (2007).
Wang, S. et al. Arabidopsis ovate family proteins, a novel transcriptional repressor family, control multiple aspects of plant growth and development. PLoS One 6, e23896 (2011).
Ikeda, M. & Ohme-Takagi, M. A novel group of transcriptional repressors in Arabidopsis. Plant Cell Physiol. 50, 970–975 (2009).
Ohta, M., Matsui, K., Hiratsu, K., Shinshi, H. & Ohme-Takagi, M. Repression domains of class II ERF transcriptional repressors share an essential motif for active repression. Plant Cell 13, 1959–1968 (2001).
Odell, J.T., Nagy, F. & Chua, N. Identification of DNA sequences required for activity of the cauliflower mosaic virus 35S promoter. Nature 313, 810–812 (1985).
Sanger, M., Daubert, S. & Goodman, R. Characteristics of a strong promoter from figwort mosaic virus: comparison with the analogous 35S promoter from cauliflower mosaic virus and the regulated mannopine synthase promoter. Plant Mol. Biol. 14, 433–443 (1990).
Shaw, C.H., Carter, G. & Watson, M. A functional map of the nopaline synthase promoter. Nucleic Acids Res. 12, 7831–7846 (1984).
Aoyama, T. & Chua, N. A glucocorticoid-mediated transcriptional induction system in transgenic plants. Plant J. 11, 605–612 (1997).
Zuo, J., Niu, Q. & Chua, N. An estrogen receptor-based transactivator XVE mediates highly inducible gene expression in transgenic plants. Plant J. 24, 265–273 (2000).
Alon, U. An Introduction to Systems Biology: Design Principles of Biological Circuits (Chapman & Hall/CRC, 2007).
Mascarenhas, D., Mettler, I., Pierce, D. & Lowe, H. Intron-mediated enhancement of heterologous gene expression in maize. Plant Mol. Biol. 15, 913–920 (1990).
Padidam, M. & Cao, Y. Elimination of transcriptional interference between tandem genes in plant cells. Biotechniques 31, 328–330, 332–324 (2001).
Depicker, A., Stachel, S., Dhaese, P., Zambryski, P. & Goodman, H.M. Nopaline synthase: transcript mapping and DNA sequence. J. Mol. Appl. Genet. 1, 561–573 (1982).
Engler, C., Gruetzner, R., Kandzia, R. & Marillonnet, S. Golden gate shuffling: a one-pot DNA shuffling method based on type IIs restriction enzymes. PLoS One 4, e5553 (2009).
Antunes, M.S. et al. A synthetic de-greening gene circuit provides a reporting system that is remotely detectable and has a re-set capacity. Plant Biotechnol. J. 4, 605–622 (2006).
Samalova, M., Brzobohaty, B. & Moore, I. pOp6/LhGR: a stringently regulated and highly responsive dexamethasone-inducible gene expression system for tobacco. Plant J. 41, 919–935 (2005).
Sakuma, Y. et al. Functional analysis of an Arabidopsis transcription factor, DREB2A, involved in drought-responsive gene expression. Plant Cell 18, 1292–1309 (2006).
Yoo, S.D., Cho, Y.H. & Sheen, J. Arabidopsis mesophyll protoplasts: a versatile cell system for transient gene expression analysis. Nat. Protoc. 2, 1565–1572 (2007).
Clough, S.J. & Bent, A.F. Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J. 16, 735–743 (1998).
Acknowledgements
We thank D. McCarthy for valuable help in preparing the manuscript. Financial support from the US Department of Energy (Advanced Research Projects Agency–Energy 2012 grant DE-AR0000311) and the US Department of Defense (Defense Threat Reduction Agency grant W911NF-09-10526) is gratefully acknowledged.
Author information
Authors and Affiliations
Contributions
K.A.S. designed and performed experiments, as well as a significant part of the data analysis, and contributed to manuscript writing. M.S.A. designed and performed experiments and contributed to manuscript writing. T.K.K. engineered many of the constructs, designed and performed experiments and contributed to manuscript writing. C.S.Z. designed and performed experiments and contributed to manuscript writing. W.X. performed data analysis, developed the camera correction method and contributed to manuscript writing. J.I.M. designed experiments, contributed to manuscript writing and supervised the overall project. A.P. designed data analysis, supervised the computational part of the project, and contributed to and supervised the writing of the paper. All authors contributed to editing of the paper and read the final version.
Corresponding authors
Ethics declarations
Competing interests
J.I.M. is founder and president of the small startup company Phytodetectors, Inc.
Integrated supplementary information
Supplementary Figure 1 Plasmids used to test repressors, repressible promoters and promoter-repressor combinations in transient protoplast assays.
(a) Repressor module used to assemble synthetic repressors under control of dexamethasone (DEX). (b) Repressor module used to assemble synthetic repressors under control of 4-hydroxytamoxifen (OHT). BsaI restriction enzyme sites were included to exchange repressors. (c) Test plasmid used to assemble all DEX-inducible promoter-repressor combinations. (d) Test plasmid used to assemble all OHT-inducible promoter-repressor combinations. Repressors and repressible promoters were cloned into the test plasmids using the KpnI restriction site. Both test plasmids contain Firefly luciferase (F-luc) expressed under control of one of two inducible promoters, pOp6 and 10xN1. F-luc is used as a proxy for the amount of repressors produced in the system. (e) Sub-cloning plasmid used to generate synthetic repressible promoters containing repressor binding sites upstream (BsaI and HindIII) or downstream (MluI and AatII) of the promoter. (f) Promoter module used to assemble repressible promoters controlling expression of the reporter gene, Renilla luciferase (R-luc). BsaI restriction enzyme sites were included to exchange promoters. A PEST protein degradation sequence was added to R-luc to increase protein turnover and facilitate quantitative measurements of promoter repression. (g) Example of a complete DEX-inducible test plasmid used for protoplast assay. LEAR is a synthetic repressor composed of LexA DNA-binding domain and EAR repressor motif. 35S2xLexA is synthetic repressible promoter composed of constitutive CaMV 35S promoter and two copies of LexA binding elements placed downstream of the promoter scaffold. LhGR2, DEX-activated transcription factor; NEV, OHT-activated transcription factor; NOST, nopaline synthase terminator; E9T, pea rbcS-E9 terminator; TB, transcription block; 35S, Cauliflower Mosaic Virus 35S promoter; AmpR, ampicillin resistance gene for bacterial selection; KanR, kanamycin resistance gene for bacterial selection; ColE1, bacterial origin of replication.
Supplementary Figure 2 Camera correction.
(a) The camera collects one image (i.e., frame) every 1/30 seconds. Each image represents the sum of pixel intensity within each well for every frame. Upper graph shows the F-luc signal is stable over time; lower graph shows the R-luc signal decays over the same time. (b) Representative graph showing the distribution of luciferase pixel intensity values RLU/(area x sec) for each well for both a plate imaged with well position A1 in the top left hand corner of the camera (blue), and the same plate with A1 in the bottom left hand corner of the camera (green). Data show that amount of luminescence recorded is influenced by the well position and changes on plate rotation. (c) Representative images of the luminescence of individual wells for one 96-well plate experiment. Wells at the edges of the plate (blue and cyan outlines) show “new-moon-shaped” occluded areas, whereas wells at the center of the plate (green outline) do not have these same occluded areas. (d) Percent change in the luminescence of the wells after rotation of the plate is shown for the original data (blue) and after imaging correction (green). Image correction removes almost all of the positional bias in the data.
Supplementary Figure 3 Schematic geometric diagram of imaging correction method.
(a) Side view of the optical system and the well in microplate of interest. Part of the well is blocked from the sight of the camera by the nontransparent wall. (b) Top view of the well of interest with the upper rim shifted to the bottom along the sight direction shown in a. The overlapping area of the two circles O1 and O2 is the visible part of the well bottom. (c) Side view of well in microplate as part of an imaginary cone. r, radius; h, height; d, diameter; a, area.
Supplementary Figure 4 Sources of noise and standard curves.
(a) Standard deviation of the three different noise sources: within a 96 well plate (12 samples), between different transformations (3 samples), and between different batches of protoplasts (4 samples). (b and c). Standard curve of luminescence produced in RLU/(area x sec) as a function of total number of molecules for F-luc (b) and R-luc (c). Lines represent best fits.
Supplementary Figure 5 Testing our normalization method with simulated and experimental repressor-repressible promoter data.
(a) Coefficient of variation (COV) of the estimated parameter B with increasing noise levels in the distribution of the random multiplicative factor, α. Non-normalized (Raw) data shows increasing COV, but the normalized data (Norm) is able to adjust for the increase in noise in α, and shows no significant change in the COV. (b) COV of the estimated parameter H also increases with increasing noise for the raw data fits, but stays approximately constant for the normalized data. (c) COV of the estimates for n do not show a difference between the normalized and raw data.
Supplementary Figure 6 Representative curve fits to non-normalized Arabidopsis data.
Raw F-luc (Input) and R-luc (Output) luminescence values for six different promoter-repressor combinations, as indicated above the graphs. Solid lines represent fits to Hill function forms using the nonlinear least squares fitting package in MATLAB. Open circles represent experimental data.
Supplementary Figure 7 Experimental design to compare function of genetic circuits in transient-expression assays and stably transformed plants.
Schematic of experimental design comparing quantitative function of synthetic promoters and repressors analyzed with transient expression (left) or stable transgenic expression (right). For transient expression, the genetic circuit is introduced into protoplasts via PEG-mediated transformation. For stable expression, the genetic circuit is first integrated into the plant’s chromosome via Agrobacterium-mediated transformation. Protoplasts are then prepared from the stably transformed plants. Quantitative parameters for the promoter-repressor pair are compared from transient and stably transformed processes.
Supplementary Figure 8 Testing the normalization factor λ with simulated data.
(a) Mean levels of estimated parameter B with increasing absolute levels in both mean and standard deviation of random number,. Raw data (Raw) show decreases in mean values, but the normalized data (Norm) show insensitivity to changes in absolute levels. (b) Mean levels of estimated parameter H also show decreases in raw data and remain constant with increasing absolute levels. (c) Mean levels of estimated parameter values of n do not show a difference between the normalized and raw data and across different absolute values.
Supplementary Figure 9 Bootstrap results.
(a) Distribution of parameter values (B, H, n) obtained from bootstrapping fits. (b) Comparison of bootstrapped estimates of the parameter B for protoplasts from the stably transformed plants (Stable) and protoplasts from transient expression (Trans) for three promoter-repressor pairs. (c) Comparison of bootstrapped estimates of half-maximal expression, H. (d) Bootstrapped estimates of the Hill coefficient n are shown for the same three pairs.
Supplementary Figure 10 Analysis of variance and honest significant difference Tukey analysis.
(a) Arabidopsis log(B) values for all good performing gene circuits categorized first by inducer type (DEX or OHT), and then by binding site position. Red star indicates a significant difference of DEX-inducible promoters when binding sites are near the TATA-box compared with binding sites either upstream or downstream of the constitutive element (P < 0.01). Black star indicates the same for OHT promoters (P = 0.04 for upstream, and P = 0.01 for downstream). (b) Sorghum log(B) values for all good performing gene circuits categorized first by inducer and then by binding site position. Red and black stars have the same meaning as in a. (P-values for significance are P < 0.01 for both upstream and downstream for DEX, and P = 0.02 for upstream and P = 0.06 for downstream, respectively, for OHT.) (c) Sorghum log(B) values for all good performing gene circuits categorized first by inducer and then by constitutive element (CaMV 35S or ZmUbi1). Red star indicates a significant difference in DEX promoters with these two elements (P < 0.01). Black star indicates the same for OHT promoters (P < 0.01). TATA, promoters with binding sites just upstream of the TATA-box; Up, binding sites placed upstream of the constitutive scaffold; Down, binding sites placed downstream of the constitutive scaffold; 35S or ZmUbi1, refers to the constitutive promoter scaffold used. log(B), logarithm of the bulk promoter strength, B. Bars are standard deviations.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–10, Supplementary Tables 1 and 2, and Supplementary Notes 1–11 (PDF 2501 kb)
Rights and permissions
About this article
Cite this article
Schaumberg, K., Antunes, M., Kassaw, T. et al. Quantitative characterization of genetic parts and circuits for plant synthetic biology. Nat Methods 13, 94–100 (2016). https://doi.org/10.1038/nmeth.3659
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.3659
This article is cited by
-
An orthogonalized PYR1-based CID module with reprogrammable ligand-binding specificity
Nature Chemical Biology (2024)
-
Multifactorial analysis of terminator performance on heterologous gene expression in Physcomitrella
Plant Cell Reports (2024)
-
Design principles for synthetic control systems to engineer plants
Plant Cell Reports (2023)
-
Orthogonal control of gene expression in plants using synthetic promoters and CRISPR-based transcription factors
Plant Methods (2022)
-
Synthetic memory circuits for stable cell reprogramming in plants
Nature Biotechnology (2022)