Abstract
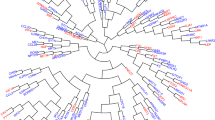
Brown adipose tissue (BAT) acts in mammals as a natural defense system against hypothermia, and its activation to a state of increased energy expenditure is believed to protect against the development of obesity. Even though the existence of BAT in adult humans has been widely appreciated1,2,3,4,5,6,7,8, its cellular origin and molecular identity remain elusive largely because of high cellular heterogeneity within various adipose tissue depots. To understand the nature of adult human brown adipocytes at single cell resolution, we isolated clonally derived adipocytes from stromal vascular fractions of adult human BAT from two individuals and globally analyzed their molecular signatures. We used RNA sequencing followed by unbiased genome-wide expression analyses and found that a population of uncoupling protein 1 (UCP1)-positive human adipocytes possessed molecular signatures resembling those of a recruitable form of thermogenic adipocytes (that is, beige adipocytes). In addition, we identified molecular markers that were highly enriched in UCP1-positive human adipocytes, a set that included potassium channel K3 (KCNK3) and mitochondrial tumor suppressor 1 (MTUS1). Further, we functionally characterized these two markers using a loss-of-function approach and found that KCNK3 and MTUS1 were required for beige adipocyte differentiation and thermogenic function. The results of this study present new opportunities for human BAT research, such as facilitating cell-based disease modeling and unbiased screens for thermogenic regulators.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
Primary accessions
ArrayExpress
Referenced accessions
Gene Expression Omnibus
References
Cypess, A.M. et al. Identification and importance of brown adipose tissue in adult humans. N. Engl. J. Med. 360, 1509–1517 (2009).
Ouellet, V. et al. Outdoor temperature, age, sex, body mass index, and diabetic status determine the prevalence, mass, and glucose-uptake activity of 18F-FDG-detected BAT in humans. J. Clin. Endocrinol. Metab. 96, 192–199 (2011).
van Marken Lichtenbelt, W.D. et al. Cold-activated brown adipose tissue in healthy men. N. Engl. J. Med. 360, 1500–1508 (2009).
Yoneshiro, T. et al. Age-related decrease in cold-activated brown adipose tissue and accumulation of body fat in healthy humans. Obesity (Silver Spring) 19, 1755–1760 (2011).
Saito, M. et al. High incidence of metabolically active brown adipose tissue in healthy adult humans: effects of cold exposure and adiposity. Diabetes 58, 1526–1531 (2009).
Nedergaard, J., Bengtsson, T. & Cannon, B. Unexpected evidence for active brown adipose tissue in adult humans. Am. J. Physiol. Endocrinol. Metab. 293, E444–E452 (2007).
Virtanen, K.A. et al. Functional brown adipose tissue in healthy adults. N. Engl. J. Med. 360, 1518–1525 (2009).
Lidell, M.E. et al. Evidence for two types of brown adipose tissue in humans. Nat. Med. 19, 631–634 (2013).
Seale, P. et al. PRDM16 controls a brown fat/skeletal muscle switch. Nature 454, 961–967 (2008).
Atit, R. et al. Beta-catenin activation is necessary and sufficient to specify the dorsal dermal fate in the mouse. Dev. Biol. 296, 164–176 (2006).
Timmons, J.A. et al. Myogenic gene expression signature establishes that brown and white adipocytes originate from distinct cell lineages. Proc. Natl. Acad. Sci. USA 104, 4401–4406 (2007).
Ohno, H., Shinoda, K., Ohyama, K., Sharp, L.Z. & Kajimura, S. EHMT1 controls brown adipose cell fate and thermogenesis through the PRDM16 complex. Nature 504, 163–167 (2013).
Harms, M. & Seale, P. Brown and beige fat: development, function and therapeutic potential. Nat. Med. 19, 1252–1263 (2013).
Kajimura, S. & Saito, M. A new era in brown adipose tissue biology: molecular control of brown fat development and energy homeostasis. Annu. Rev. Physiol. 76, 225–249 (2014).
Waldén, T.B., Hansen, I.R., Timmons, J.A., Cannon, B. & Nedergaard, J. Recruited vs. nonrecruited molecular signatures of brown, “brite,” and white adipose tissues. Am. J. Physiol. Endocrinol. Metab. 302, E19–E31 (2012).
Wu, J. et al. Beige adipocytes are a distinct type of thermogenic fat cell in mouse and human. Cell 150, 366–376 (2012).
Ohno, H., Shinoda, K., Spiegelman, B.M. & Kajimura, S. PPARgamma agonists induce a white-to-brown fat conversion through stabilization of PRDM16 protein. Cell Metab. 15, 395–404 (2012).
Sharp, L.Z. et al. Human BAT possesses molecular signatures that resemble beige/brite cells. PLoS ONE 7, e49452 (2012).
Lee, P., Werner, C.D., Kebebew, E. & Celi, F.S. Functional thermogenic beige adipogenesis is inducible in human neck fat. Int. J. Obes. (Lond). 38, 170–176 (2014).
Jespersen, N.Z. et al. A classical brown adipose tissue mRNA signature partly overlaps with brite in the supraclavicular region of adult humans. Cell Metab. 17, 798–805 (2013).
Cypess, A.M. et al. Anatomical localization, gene expression profiling and functional characterization of adult human neck brown fat. Nat. Med. 19, 635–639 (2013).
Villarroya, F. & Vidal-Puig, A. Beyond the sympathetic tone: the new brown fat activators. Cell Metab. 17, 638–643 (2013).
Tews, D. et al. Comparative gene array analysis of progenitor cells from human paired deep neck and subcutaneous adipose tissue. Mol. Cell. Endocrinol. 395, 41–50 (2014).
Long, J.Z. et al. A smooth muscle-like origin for beige adipocytes. Cell Metab. 19, 810–820 (2014).
Rajakumari, S. et al. EBF2 determines and maintains brown adipocyte identity. Cell Metab. 17, 562–574 (2013).
Schulz, T.J. et al. Brown-fat paucity due to impaired BMP signalling induces compensatory browning of white fat. Nature 495, 379–383 (2013).
Svensson, P.A. et al. Gene expression in human brown adipose tissue. Int. J. Mol. Med. 27, 227–232 (2011).
Søndergaard, E. et al. Chronic adrenergic stimulation induces brown adipose tissue differentiation in visceral adipose tissue. Diabet. Med. 32, e4–e8 (2015).
Nouet, S. et al. Trans-inactivation of receptor tyrosine kinases by novel angiotensin II AT2 receptor-interacting protein, ATIP. J. Biol. Chem. 279, 28989–28997 (2004).
Chondronikola, M. et al. Brown adipose tissue improves whole-body glucose homeostasis and insulin sensitivity in humans. Diabetes 63, 4089–4099 (2014).
van der Lans, A.A. et al. Cold acclimation recruits human brown fat and increases nonshivering thermogenesis. J. Clin. Invest. 123, 3395–3403 (2013).
Yoneshiro, T. et al. Recruited brown adipose tissue as an antiobesity agent in humans. J. Clin. Invest. 123, 3404–3408 (2013).
Lee, P. et al. Temperature-acclimated brown adipose tissue modulates insulin sensitivity in humans. Diabetes 63, 3686–3698 (2014).
Trapnell, C. et al. Differential analysis of gene regulation at transcript resolution with RNA-seq. Nat. Biotechnol. 31, 46–53 (2013).
Irizarry, R.A. et al. Exploration, normalization, and summaries of high density oligonucleotide array probe level data. Biostatistics 4, 249–264 (2003).
Li, C. & Wong, W.H. Model-based analysis of oligonucleotide arrays: expression index computation and outlier detection. Proc. Natl. Acad. Sci. USA 98, 31–36 (2001).
Saeed, A.I. et al. TM4: a free, open-source system for microarray data management and analysis. Biotechniques 34, 374–378 (2003).
Stöver, B.C. & Muller, K.F. TreeGraph 2: combining and visualizing evidence from different phylogenetic analyses. BMC Bioinformatics 11, 7 (2010).
Huang, W., Sherman, B.T. & Lempicki, R.A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 4, 44–57 (2009).
Inoue, M., Harada, K., Matsuoka, H., Sata, T. & Warashina, A. Inhibition of TASK1-like channels by muscarinic receptor stimulation in rat adrenal medullary cells. J. Neurochem. 106, 1804–1814 (2008).
Cohen, J. Statistical Power Analysis for the Behavioral Sciences (L. Erlbaum Associates, Hillsdale, N.J., 1988).
Acknowledgements
We acknowledge support from the National Institutes of Health (NIH) (DK087853 and DK097441), the UCSF Diabetes Research Center grant (DK63720), the UCSF Program for Breakthrough Biomedical Research program, the Pew Charitable Trust, and the Japan Science and Technology Agency (all to S.K.); and from the NIH (P50-GM60338) and the American Dental Association (1-14-TS-35) (both to L.S.S.). K.S. is supported by a fellowship from the Japan Society for the Promotion of Science. Y.H. is supported by the Manpei Suzuki Diabetes Foundation. I.H.N.L. is supported by the Dutch Heart Foundation.
Author information
Authors and Affiliations
Contributions
K.S. and S.K. designed the experiments. K.S., I.H.N.L., Y.H., H.H., S.B.S., M.K. and S.K. performed the cellular experiments and analyzed the data. M.C., A.M.C., L.S.S. and Y.-H.T. provided adipose tissue samples. K.S., Y.H., H.H., R.X. and S.K. analyzed adipose tissue samples. K.S. and S.K. wrote the manuscript. All authors contributed to editing the manuscript. S.K. conceived and managed the project.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–5 (PDF 864 kb)
Supplementary Table 1
Human brown adipocyte-enriched genes in a differentiated state. (PDF 1206 kb)
Supplementary Table 2
Human brown adipocyte-enriched genes in an undifferentiated state (PDF 1153 kb)
Supplementary Table 3
Expression level of smooth muscle lineage-selective genes in human brown and white adipocytes at undifferentiated and differentiated states. (PDF 79 kb)
Supplementary Table 4
List of core brown fat-selective genes conserved in mice and humans (group A). (PDF 125 kb)
Supplementary Table 5
Primer sequences used for quantitative RT-PCR. (PDF 115 kb)
Rights and permissions
About this article
Cite this article
Shinoda, K., Luijten, I., Hasegawa, Y. et al. Genetic and functional characterization of clonally derived adult human brown adipocytes. Nat Med 21, 389–394 (2015). https://doi.org/10.1038/nm.3819
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nm.3819
This article is cited by
-
The serotonin transporter sustains human brown adipose tissue thermogenesis
Nature Metabolism (2023)
-
Transcriptional repression of beige fat innervation via a YAP/TAZ-S100B axis
Nature Communications (2023)
-
Transcriptomic profiling of the telomerase transformed Mesenchymal stromal cells derived adipocytes in response to rosiglitazone
BMC Genomic Data (2022)
-
Browning of the white adipose tissue regulation: new insights into nutritional and metabolic relevance in health and diseases
Nutrition & Metabolism (2022)
-
Cold exposure prevents fat accumulation in striped hamsters refed a high-fat diet following food restriction
BMC Zoology (2022)