Abstract
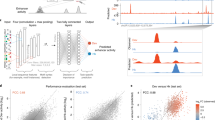
Genes are regulated by transcription factors that bind to regions of genomic DNA called enhancers. Considerable effort is focused on identifying transcription factor binding sites, with the goal of predicting gene expression from DNA sequence. Despite this effort, general, predictive models of enhancer function are currently lacking. Here we combine quantitative models of enhancer function with manipulations using engineered transcription factors to examine the extent to which enhancer function can be controlled in a quantitatively predictable manner. Our models, which incorporate few free parameters, can accurately predict the contributions of ectopic transcription factor inputs. These models allow the predictable 'tuning' of enhancers, providing a framework for the quantitative control of enhancers with engineered transcription factors.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Levo, M. & Segal, E. In pursuit of design principles of regulatory sequences. Nat. Rev. Genet. 15, 453–468 (2014).
Shlyueva, D., Stampfel, G. & Stark, A. Transcriptional enhancers: from properties to genome-wide predictions. Nat. Rev. Genet. 15, 272–286 (2014).
Crocker, J., Tamori, Y. & Erives, A. Evolution acts on enhancer organization to fine-tune gradient threshold readouts. PLoS Biol. 6, e263 (2008).
Erives, A. & Levine, M. Coordinate enhancers share common organizational features in the Drosophila genome. Proc. Natl. Acad. Sci. USA 101, 3851–3856 (2004).
Panne, D., Maniatis, T. & Harrison, S.C. An atomic model of the interferon-β enhanceosome. Cell 129, 1111–1123 (2007).
Panne, D. The enhanceosome. Curr. Opin. Struct. Biol. 18, 236–242 (2008).
Spitz, F. & Furlong, E.E.M. Transcription factors: from enhancer binding to developmental control. Nat. Rev. Genet. 13, 613–626 (2012).
Kulkarni, M.M. & Arnosti, D.N. Information display by transcriptional enhancers. Development 130, 6569–6575 (2003).
Lusk, R.W. & Eisen, M.B. Evolutionary mirages: selection on binding site composition creates the illusion of conserved grammars in Drosophila enhancers. PLoS Genet. 6, e1000829 (2010).
Swanson, C.I., Evans, N.C. & Barolo, S. Structural rules and complex regulatory circuitry constrain expression of a Notch- and EGFR-regulated eye enhancer. Dev. Cell 18, 359–370 (2010).
Turing, A.M. The chemical basis of morphogenesis. Philos. Trans. R. Soc. B Biol. Sci. 237, 37–72 (1952).
Jaeger, J. et al. Dynamic control of positional information in the early Drosophila embryo. Nature 430, 368–371 (2004).
Janssens, H. et al. Quantitative and predictive model of transcriptional control of the Drosophila melanogaster even skipped gene. Nat. Genet. 38, 1159–1165 (2006).
Kim, A.-R. et al. Rearrangements of 2.5 kilobases of noncoding DNA from the Drosophila even-skipped locus define predictive rules of genomic cis-regulatory logic. PLoS Genet. 9, e1003243 (2013).
Ay, A. & Arnosti, D.N. Mathematical modeling of gene expression: a guide for the perplexed biologist. Crit. Rev. Biochem. Mol. Biol. 46, 137–151 (2011).
Samee, M.A.H. & Sinha, S. Quantitative modeling of a gene's expression from its intergenic sequence. PLoS Comput. Biol. 10, e1003467 (2014).
Crocker, J. et al. Low affinity binding site clusters confer Hox specificity and regulatory robustness. Cell 160, 191–203 (2015).
Wong, D. et al. Extensive characterization of NF-κB binding uncovers non-canonical motifs and advances the interpretation of genetic functional traits. Genome Biol. 12, R70 (2011).
Badis, G. et al. Diversity and complexity in DNA recognition by transcription factors. Science 324, 1720–1723 (2009).
Zuo, Z. & Stormo, G.D. High-resolution specificity from DNA sequencing highlights alternative modes of Lac repressor binding. Genetics 198, 1329–1343 (2014).
Glass, L. & Kauffman, S.A. The logical analysis of continuous, non-linear biochemical control networks. J. Theor. Biol. 39, 103–129 (1973).
Reinitz, J., Mjolsness, E. & Sharp, D.H. Model for cooperative control of positional information in Drosophila by bicoid and maternal hunchback. J. Exp. Zool. 271, 47–56 (1995).
Reinitz, J. & Sharp, D.H. Mechanism of eve stripe formation. Mech. Dev. 49, 133–158 (1995).
Ilsley, G.R., Fisher, J., Apweiler, R., De Pace, A.H. & Luscombe, N.M. Cellular resolution models for even skipped regulation in the entire Drosophila embryo. eLife 2, e00522 (2013).
Staller, M.V. et al. Shadow enhancers enable Hunchback bifunctionality in the Drosophila embryo. Proc. Natl. Acad. Sci. USA 112, 785–790 (2015).
Crocker, J. & Stern, D.L. TALE-mediated modulation of transcriptional enhancers in vivo. Nat. Methods 10, 762–767 (2013).
Tracey, W.D. Jr., Ning, X., Klingler, M., Kramer, S.G. & Gergen, J.P. Quantitative analysis of gene function in the Drosophila embryo. Genetics 154, 273–284 (2000).
Fujioka, M., Emi-Sarker, Y., Yusibova, G.L., Goto, T. & Jaynes, J.B. Analysis of an even-skipped rescue transgene reveals both composite and discrete neuronal and early blastoderm enhancers, and multi-stripe positioning by gap gene repressor gradients. Development 126, 2527–2538 (1999).
Struffi, P., Corado, M., Kulkarni, M. & Arnosti, D.N. Quantitative contributions of CtBP-dependent and -independent repression activities of Knirps. Development 131, 2419–2429 (2004).
Langeland, J.A., Attai, S.F., Vorwerk, K. & Carroll, S.B. Positioning adjacent pair-rule stripes in the posterior Drosophila embryo. Development 120, 2945–2955 (1994).
Segal, E., Raveh-Sadka, T., Schroeder, M., Unnerstall, U. & Gaul, U. Predicting expression patterns from regulatory sequence in Drosophila segmentation. Nature 451, 535–540 (2008).
Clyde, D.E. et al. A self-organizing system of repressor gradients establishes segmental complexity in Drosophila. Nature 426, 849–853 (2003).
Small, S., Blair, A. & Levine, M. Regulation of two pair-rule stripes by a single enhancer in the Drosophila embryo. Dev. Biol. 175, 314–324 (1996).
Morán, E. & Jiménez, G. The tailless nuclear receptor acts as a dedicated repressor in the early Drosophila embryo. Mol. Cell. Biol. 26, 3446–3454 (2006).
Surkova, S. et al. Quantitative imaging of gene expression in Drosophila embryos. Cold Spring Harb. Protoc. 2013, 488–497 (2013).
Surkova, S. et al. Characterization of the Drosophila segment determination morphome. Dev. Biol. 313, 844–862 (2008).
Janssens, H. et al. Lack of tailless leads to an increase in expression variability in Drosophila embryos. Dev. Biol. 377, 305–317 (2013).
Jiang, J. & Levine, M. Binding affinities and cooperative interactions with bHLH activators delimit threshold responses to the dorsal gradient morphogen. Cell 72, 741–752 (1993).
Zinzen, R.P., Senger, K., Levine, M. & Papatsenko, D. Computational models for neurogenic gene expression in the Drosophila embryo. Curr. Biol. 16, 1358–1365 (2006).
Ip, Y.T., Park, R.E., Kosman, D., Bier, E. & Levine, M. The dorsal gradient morphogen regulates stripes of rhomboid expression in the presumptive neuroectoderm of the Drosophila embryo. Genes Dev. 6, 1728–1739 (1992).
Crocker, J., Potter, N. & Erives, A. Dynamic evolution of precise regulatory encodings creates the clustered site signature of enhancers. Nat. Commun. 1, 99 (2010).
Barolo, S. & Levine, M. hairy mediates dominant repression in the Drosophila embryo. EMBO J. 16, 2883–2891 (1997).
Davidson, E.H. The Regulatory Genome: Gene Regulatory Networks in Development And Evolution (Academic Press, 2006).
Arnosti, D.N., Barolo, S., Levine, M. & Small, S. The eve stripe 2 enhancer employs multiple modes of transcriptional synergy. Development 122, 205–214 (1996).
Ludwig, M.Z., Bergman, C., Patel, N.H. & Kreitman, M. Evidence for stabilizing selection in a eukaryotic enhancer element. Nature 403, 564–567 (2000).
Ludwig, M.Z. et al. Functional evolution of a cis-regulatory module. PLoS Biol. 3, e93 (2005).
Rastegar, S. et al. The words of the regulatory code are arranged in a variable manner in highly conserved enhancers. Dev. Biol. 318, 366–377 (2008).
Jin, H. et al. Genome-wide screens for in vivo Tinman binding sites identify cardiac enhancers with diverse functional architectures. PLoS Genet. 9, e1003195 (2013).
Brown, C.D., Johnson, D.S. & Sidow, A. Functional architecture and evolution of transcriptional elements that drive gene coexpression. Science 317, 1557–1560 (2007).
Menoret, D. et al. Genome-wide analyses of Shavenbaby target genes reveals distinct features of enhancer organization. Genome Biol. 14, R86 (2013).
Hare, E.E., Peterson, B.K., Iyer, V.N., Meier, R. & Eisen, M.B. Sepsid even-skipped enhancers are functionally conserved in Drosophila despite lack of sequence conservation. PLoS Genet. 4, e1000106 (2008).
Junion, G. et al. A transcription factor collective defines cardiac cell fate and reflects lineage history. Cell 148, 473–486 (2012).
Arnosti, D.N. & Kulkarni, M.M. Transcriptional enhancers: intelligent enhanceosomes or flexible billboards? J. Cell. Biochem. 94, 890–898 (2005).
Arnosti, D.N., Gray, S., Barolo, S., Zhou, J. & Levine, M. The gap protein knirps mediates both quenching and direct repression in the Drosophila embryo. EMBO J. 15, 3659–3666 (1996).
Deng, Q., Ramsköld, D., Reinius, B. & Sandberg, R. Single-cell RNA-seq reveals dynamic, random monoallelic gene expression in mammalian cells. Science 343, 193–196 (2014).
Shalek, A.K. et al. Single-cell RNA-seq reveals dynamic paracrine control of cellular variation. Nature 510, 363–369 (2014).
Jaitin, D.A. et al. Massively parallel single-cell RNA-seq for marker-free decomposition of tissues into cell types. Science 343, 776–779 (2014).
Treutlein, B. et al. Reconstructing lineage hierarchies of the distal lung epithelium using single-cell RNA-seq. Nature 509, 371–375 (2014).
Patel, A.P. et al. Single-cell RNA-seq highlights intratumoral heterogeneity in primary glioblastoma. Science 344, 1396–1401 (2014).
Achim, K. et al. High-throughput spatial mapping of single-cell RNA-seq data to tissue of origin. Nat. Biotechnol. 33, 503–509 (2015).
Satija, R., Farrell, J.A., Gennert, D., Schier, A.F. & Regev, A. Spatial reconstruction of single-cell gene expression data. Nat. Biotechnol. 33, 495–502 (2015).
Fowlkes, C.C. et al. A quantitative spatiotemporal atlas of gene expression in the Drosophila blastoderm. Cell 133, 364–374 (2008).
Acknowledgements
We thank M. Levine, J. Bothma, S. Barolo, J. Jaynes, N. Luscombe, the entire Stern laboratory and the members of the Janelia Transcriptional Imaging Consortium for discussions and advice throughout the project. We thank C. Standley, J. Cande, E. Preger–Ben Noon, J. Shaevitz and especially R. Mann for critically reviewing our manuscript. We thank D. Papatsenko and H. Janssens for supplying the dorsoventral rho and tll mutant data, respectively.
Author information
Authors and Affiliations
Contributions
J.C. and G.R.I. conceived the study, designed and executed the experiments, and analyzed the data, with mentorship from D.L.S. J.C., G.R.I. and D.L.S. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 eve stripe 3+7 TALE-GFP does not alter eve expression, and different levels of TALER expression do not alter the effect of TALER.
(a–d) Stage 5 embryos stained for Eve protein in either a wild-type background (a) or carrying a single ubiquitous nos::GAL4 driver (b,c) or two nos::GAL4 drivers (d) and the indicated UAS::stripe 3+7 TALE. (e–h) Profiles of average expression levels for the indicated genotype (n = 10 for each genotype). Bounding areas around experimental data indicate 1 s.d. AU, arbitrary units of fluorescence intensity.
Supplementary Figure 2 The logistic eve 4+6 model accurately predicts expression in different genetic backgrounds.
(a–c) Virtual Embryo predictions from the eve 4+6 model, for eve 4+6 expression in the indicated genetic background. The knirps (kni) heterozygote prediction has knirps at 75% of normal levels, which implies compensatory regulation; in the homozygote, knirps is not expressed. (d–f) Stage 5 embryos stained for Even-skipped protein (Eve) in either a wild-type background (d) or the indicated genetic background (e,f) with the third Eve stripe labeled in each panel. (g–i) Larval cuticle preparations for the indicated genomic backgrounds, with the A1 segment labeled in each panel. Red brackets highlight the modified larval segments.
Supplementary Figure 3 The logistic eve 4+6 model accurately predicts expression patterns resulting from input misexpression.
(a) Ventral view of the even-skipped (eve) expression pattern in a wild-type stage 5 embryo. (b) Ventral view of the Virtual Embryo eve 4+6 model prediction. (c) Expression pattern of an embryo carrying ectopic knirps (kni) expression driven from a snail (sna) enhancer. (d) Virtual embryo with ectopic knirps expression. (e,g) Ventral views showing stripe-specific changes in expression resulting from ectopic knirps expression, in embryos carrying one (e) or two (g) copies of the sna::kni transgene. (f,h) Ventral view of the eve 4+6 model predictions with ectopic expression of knirps in the snail domain (see also e and g). In situ images are reproduced from Figure 1 of Clyde et al.32.
Supplementary Figure 4 Hairy and Krüppel repressive domains display quantitatively different repressive strength.
(a–d) Stage 5 embryos stained for Eve protein in either a wild-type background (a) or carrying a ubiquitous nos::GAL4 driver and the indicated UAS::stripe 3+7–TALE (b–d). (e–h) Profiles of average expression levels for the indicated genotype (n = 10 for each genotype). In all plots, the solid blue line denotes model predictions; black, green, red and orange lines denote wild-type, enhancer-GFP, enhancer-TALER-hairy and enhancer-TALER-Krüppel constructs, respectively. Bounding areas around experimental data indicate 1 s.d. AU, arbitrary units of fluorescence intensity.
Supplementary Figure 5 The logistic eve 4+6 model accurately predicts expression in tailless (tll)-mutant embryos.
(a,b) Predicted expression levels for the eve 3+7 (pink) and eve 4+6 (blue) enhancers in wild-type (a) or tailless-mutant (b) embryos, based on previously published primary expression data37. Dashed lines indicate actual Even-skipped protein (Eve) expression levels for each background. (c,d) Stage 5 embryos stained for Eve protein in either a wild-type background (c) or the tailless-mutant background (d), with the third Eve stripe labeled in each panel. (e,f) Larval cuticle preparations for the indicated genomic backgrounds, with the A1 segment labeled in each panel.
Supplementary Figure 6 Cooperative interactions between Dorsal (Dl) and Twist (Twi) are combined with snail (sna) in a linear manner.
(a) A logistic model using linear combinations of previously published expression data for the known regulators of rhomboid (rho)39, indicating the model output (dashed black line), the regulatory inputs (blue, green and orange) and native rhomboid enhancer expression (pink). The cyan dashed line indicates the model prediction for three TALEAs. (b) Cooperativity between Dorsal and Twist is modeled by the addition of a nonlinear term to the model. Note the differences in the model predictions in a and b in the dorsal and ventral regions with the addition of TALEAs (blue dashed lines).
Supplementary Figure 7 The rhomboid (rho) enhancer model including Dorsal-Twist cooperativity accurately predicts expression patterns resulting from binding site modifications.
(a–d) rhomboid model predictions in wild-type (a), snail (sna)-deficient (b), twist-deficient (c) and dorsal-deficient backgrounds (d). (e–h) Lateral views of rhomboid enhancer expression with the indicated binding site modifications; see also Figure 4 of Ip et al.40.
Supplementary Figure 8 A balance of transcriptional activators and repressors at the rhomboid enhancer.
(a) Schematic of the rhomboid enhancer region, with known binding sites indicated40. (b–d) Stage 5 embryos from either wild-type rho-lacZ reporter (b), a reporter carrying a mutation in the Twist site of rho-lacZ (c) or the Twist-mutant rho-lacZ reporter together with UAS::rho-TALEA-VP64 (d) stained for lacZ (β-gal) RNA.
Supplementary Figure 9 TALEAs can drive increased activation of native enhancers in multiple developmental contexts.
(a,b) In the neuroectoderm, stage 5 embryos display increased expression of vndNEE-lacZ when targeted with vnd-enhancer-TALEA-VP64 by nos::GAL4 (b) as compared with a wild-type background (a). (c–e) In the wing imaginal disc, a TALEA increases enhancer expression. (c) A schematic of Decapentaplegic (dpp) regulation of optomotor blind (omb) in the larval imaginal wing disc. (d,e) Larval imaginal wing discs show stronger expression of omb-lacZ transgenes when targeted with omb-enhancer-TALEA-VP64 by Act5c::GAL4 (e) as compared with wild-type background (d).
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–9 and Supplementary Note. (PDF 4323 kb)
Rights and permissions
About this article
Cite this article
Crocker, J., Ilsley, G. & Stern, D. Quantitatively predictable control of Drosophila transcriptional enhancers in vivo with engineered transcription factors. Nat Genet 48, 292–298 (2016). https://doi.org/10.1038/ng.3509
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3509
This article is cited by
-
Reprogramming of regulatory network using expression uncovers sex-specific gene regulation in Drosophila
Nature Communications (2018)
-
Is a super-enhancer greater than the sum of its parts?
Nature Genetics (2017)