Abstract
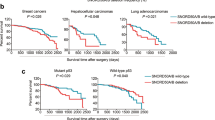
Small nucleolar RNAs (snoRNAs) are conserved noncoding RNAs best studied as ribonucleoprotein (RNP) guides in RNA modification1,2. To explore their role in cancer, we compared 5,473 tumor-normal genome pairs to identify snoRNAs with frequent copy number loss. The SNORD50A-SNORD50B snoRNA locus was deleted in 10–40% of 12 common cancers, where its loss was associated with reduced survival. A human protein microarray screen identified direct SNORD50A and SNORD50B RNA binding to K-Ras. Loss of SNORD50A and SNORD50B increased the amount of GTP-bound, active K-Ras and hyperactivated Ras-ERK1/ERK2 signaling. Loss of these snoRNAs also increased binding by farnesyltransferase to K-Ras and increased K-Ras prenylation, suggesting that KRAS mutation might synergize with SNORD50A and SNORD50B loss in cancer. In agreement with this hypothesis, CRISPR-mediated deletion of SNORD50A and SNORD50B in KRAS-mutant tumor cells enhanced tumorigenesis, and SNORD50A and SNORD50B deletion and oncogenic KRAS mutation co-occurred significantly in multiple human tumor types. SNORD50A and SNORD50B snoRNAs thus directly bind and inhibit K-Ras and are recurrently deleted in human cancer.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Matera, A.G., Terns, R.M. & Terns, M.P. Non-coding RNAs: lessons from the small nuclear and small nucleolar RNAs. Nat. Rev. Mol. Cell Biol. 8, 209–220 (2007).
Kiss, T. Small nucleolar RNA–guided post-transcriptional modification of cellular RNAs. EMBO J. 20, 3617–3622 (2001).
Falaleeva, M. & Stamm, S. Processing of snoRNAs as a new source of regulatory non-coding RNAs: snoRNA fragments form a new class of functional RNAs. Bioessays 35, 46–54 (2013).
Wang, C. & Meier, U.T. Architecture and assembly of mammalian H/ACA small nucleolar and telomerase ribonucleoproteins. EMBO J. 23, 1857–1867 (2004).
Reichow, S.L., Hamma, T., Ferre-D'Amare, A.R. & Varani, G. The structure and function of small nucleolar ribonucleoproteins. Nucleic Acids Res. 35, 1452–1464 (2007).
Sahoo, T. et al. Prader-Willi phenotype caused by paternal deficiency for the HBII-85 C/D box small nucleolar RNA cluster. Nat. Genet. 40, 719–721 (2008).
Bhartiya, D., Talwar, J., Hasija, Y. & Scaria, V. Systematic curation and analysis of genomic variations and their potential functional consequences in snoRNA loci. Hum. Mutat. 33, E2367–E2374 (2012).
Bellodi, C. et al. H/ACA small RNA dysfunctions in disease reveal key roles for noncoding RNA modifications in hematopoietic stem cell differentiation. Cell Rep. 3, 1493–1502 (2013).
Michel, C.I. et al. Small nucleolar RNAs U32a, U33, and U35a are critical mediators of metabolic stress. Cell Metab. 14, 33–44 (2011).
Mannoor, K., Liao, J. & Jiang, F. Small nucleolar RNAs in cancer. Biochim. Biophys. Acta 1826, 121–128 (2012).
Williams, G.T. & Farzaneh, F. Are snoRNAs and snoRNA host genes new players in cancer? Nat. Rev. Cancer 12, 84–88 (2012).
Yin, Q.F. et al. Long noncoding RNAs with snoRNA ends. Mol. Cell 48, 219–230 (2012).
Rebane, A., Roomere, H. & Metspalu, A. Locations of several novel 2′-O-methylated nucleotides in human 28S rRNA. BMC Mol. Biol. 3, 1 (2002).
Kretz, M. et al. Control of somatic tissue differentiation by the long non-coding RNA TINCR. Nature 493, 231–235 (2013).
Siprashvili, Z. et al. Identification of proteins binding coding and non-coding human RNAs using protein microarrays. BMC Genomics 13, 633 (2012).
Söderberg, O. et al. Direct observation of individual endogenous protein complexes in situ by proximity ligation. Nat. Methods 3, 995–1000 (2006).
Fernández-Medarde, A. & Santos, E. Ras in cancer and developmental diseases. Genes Cancer 2, 344–358 (2011).
Pritchard, A.L. & Hayward, N.K. Molecular pathways: mitogen-activated protein kinase pathway mutations and drug resistance. Clin. Cancer Res. 19, 2301–2309 (2013).
Lazarov, M. et al. CDK4 coexpression with Ras generates malignant human epidermal tumorigenesis. Nat. Med. 8, 1105–1114 (2002).
Serrano, M., Lin, A.W., McCurrach, M.E., Beach, D. & Lowe, S.W. Oncogenic ras provokes premature cell senescence associated with accumulation of p53 and p16INK4a. Cell 88, 593–602 (1997).
Weibrecht, I. et al. Proximity ligation assays: a recent addition to the proteomics toolbox. Expert Rev. Proteomics 7, 401–409 (2010).
Kandoth, C. et al. Mutational landscape and significance across 12 major cancer types. Nature 502, 333–339 (2013).
Mei, Y.P. et al. Small nucleolar RNA 42 acts as an oncogene in lung tumorigenesis. Oncogene 31, 2794–2804 (2012).
Tanaka, R. et al. Intronic U50 small-nucleolar-RNA (snoRNA) host gene of no protein-coding potential is mapped at the chromosome breakpoint t(3;6)(q27;q15) of human B-cell lymphoma. Genes Cells 5, 277–287 (2000).
Dong, X.Y. et al. SnoRNA U50 is a candidate tumor-suppressor gene at 6q14.3 with a mutation associated with clinically significant prostate cancer. Hum. Mol. Genet. 17, 1031–1042 (2008).
Karbstein, K., Jonas, S. & Doudna, J.A. An essential GTPase promotes assembly of preribosomal RNA processing complexes. Mol. Cell 20, 633–643 (2005).
Clementi, N. & Polacek, N. Ribosome-associated GTPases: the role of RNA for GTPase activation. RNA Biol. 7, 521–527 (2010).
Clementi, N., Chirkova, A., Puffer, B., Micura, R. & Polacek, N. Atomic mutagenesis reveals A2660 of 23S ribosomal RNA as key to EF-G GTPase activation. Nat. Chem. Biol. 6, 344–351 (2010).
Piccaluga, P.P. et al. Gene expression analysis of peripheral T cell lymphoma, unspecified, reveals distinct profiles and new potential therapeutic targets. J. Clin. Invest. 117, 823–834 (2007).
Flockhart, R.J. et al. BRAFV600E remodels the melanocyte transcriptome and induces BANCR to regulate melanoma cell migration. Genome Res. 22, 1006–1014 (2012).
Skrzypczak, M. et al. Modeling oncogenic signaling in colon tumors by multidirectional analyses of microarray data directed for maximization of analytical reliability. PLoS ONE 10.1371/journal.pone.0013091 (1 October 2010).
Acknowledgements
We thank J. Crabtree, J. Ferrell, S. Artandi, A. Oro, H. Chang and members of the Khavari laboratory for presubmission review and advice. This work was supported by the US Veterans Affairs Office of Research and Development and by US National Institutes of Health/National Cancer Institute grant CA142635 and by US National Institutes of Health/National Institute of Arthritis Musculoskeletal and Skin Diseases grant AR49737 to P.A.K.
Author information
Authors and Affiliations
Contributions
Z.S. designed and executed experiments, analyzed data and wrote the manuscript. D.E.W., D.J., A.J.U., A.B., R.F., B.J.Z., Y.C. and F.M. executed experiments, analyzed data and contributed to design of experiments. D.J. and R.M.S. executed experiments. J.D.P. helped design experiments and analyzed data. P.A.K. designed experiments, analyzed data and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7 and Supplementary Tables 1–5. (PDF 2969 kb)
Rights and permissions
About this article
Cite this article
Siprashvili, Z., Webster, D., Johnston, D. et al. The noncoding RNAs SNORD50A and SNORD50B bind K-Ras and are recurrently deleted in human cancer. Nat Genet 48, 53–58 (2016). https://doi.org/10.1038/ng.3452
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3452
This article is cited by
-
The association of UBAP2L and G3BP1 mediated by small nucleolar RNA is essential for stress granule formation
Communications Biology (2023)
-
SNORD6 promotes cervical cancer progression by accelerating E6-mediated p53 degradation
Cell Death Discovery (2023)
-
Involvement of the oncogenic small nucleolar RNA SNORA24 in regulation of p53 stability in colorectal cancer
Cell Biology and Toxicology (2023)
-
SNORD1C maintains stemness and 5-FU resistance by activation of Wnt signaling pathway in colorectal cancer
Cell Death Discovery (2022)
-
KRAS-related long noncoding RNAs in human cancers
Cancer Gene Therapy (2022)