Abstract
Cystic echinococcosis (hydatid disease), caused by the tapeworm E. granulosus, is responsible for considerable human morbidity and mortality. This cosmopolitan disease is difficult to diagnose, treat and control. We present a draft genomic sequence for the worm comprising 151.6 Mb encoding 11,325 genes. Comparisons with the genome sequences from other taxa show that E. granulosus has acquired a spectrum of genes, including the EgAgB family, whose products are secreted by the parasite to interact and redirect host immune responses. We also find that genes in bile salt pathways may control the bidirectional development of E. granulosus, and sequence differences in the calcium channel subunit EgCavβ1 may be associated with praziquantel sensitivity. Our study offers insights into host interaction, nutrient acquisition, strobilization, reproduction, immune evasion and maturation in the parasite and provides a platform to facilitate the development of new, effective treatments and interventions for echinococcosis control.
Similar content being viewed by others
Main
The dog tapeworm E. granulosus is one of a group of medically important parasitic helminths of the family Taeniidae (Platyhelminthes; Cestoda; Cyclophyllidea) that infect at least 50 million people globally1. Its life cycle involves two mammals, including an intermediate host, usually a domestic or wild ungulate (humans are accidental intermediate hosts) and a canine-definitive host, such as the domestic dog. The larval (metacestode) stage causes hydatidosis (cystic hydatid disease; cystic echinococcosis), a chronic cyst-forming disease in the intermediate (human) host. Currently, up to 3 million people are infected with E. granulosus2,3, and, in some areas, 10% of the population has detectable hydatid cysts by abdominal ultrasound and chest X-ray4,5.
E. granulosus has no gut, circulatory or respiratory organs. It is monoecious, producing diploid eggs that give rise to ovoid embryos, the oncospheres. Strobilization is a notable feature of cestode biology, whereby proglottids bud distally from the anterior scolex, resulting in the production of tandem reproductive units exhibiting increasing degrees of development. A unique characteristic of the larvae (protoscoleces, PSCs) within the hydatid cyst is an ability to develop bidirectionally into an adult worm in the dog gastrointestinal tract or into a secondary hydatid cyst in the intermediate (human) host, a process triggered by bile acids6. Another distinct feature of E. granulosus is its capacity to infect and adapt to a large number of mammalian species as intermediate hosts, which has contributed to its cosmopolitan global distribution.
Here we report the sequence and analysis of the E. granulosus genome. Comprising nine pairs of chromosomes7, it is one of the first cestode genomes to be sequenced and complements the recent publication by Tsai et al.8 of a high-quality genome for Echinococcus multilocularis (the cause of alveolar echinococcosis), together with draft genomes of three other tapeworm species including E. granulosus. Our study provides insights into the biology, development, differentiation, evolution and host interaction of E. granulosus and has identified a range of drug and vaccine targets that can facilitate the development of new intervention tools for hydatid treatment and control.
Results
Genome sequencing and annotation
We sequenced 2.8 Gb of 454 GS FLX shotgun sequences and 20.8 Gb of Solexa paired-end or mate-paired sequences using DNA extracted from a single E. granulosus cyst (G1 genotype; common sheepdog strain)9 and obtained 967 scaffolds totaling 110.86 Mb (Supplementary Tables 1 and 2). We validated genome sequence quality and assembly through comparisons with the E. granulosus mitochondrial genome, fosmid clones and EST sequences (Supplementary Fig. 1, Supplementary Table 3 and Supplementary Note). Of the 22,340 contigs, 13,158 were identified as repeats, with the total size reaching 45.86 Mb when taking into account repeat copies (Supplementary Tables 4 and 5 and Supplementary Note). We estimated the genome size to be 151.61 Mb, including 105.75 Mb of unique contigs and 45.86 Mb of repeats. This genome size is consistent with that calculated on the basis of k-mer frequencies (Supplementary Fig. 2).
We predicted a total of 11,325 protein-coding genes spanning 10.4% of the genome (Table 1, Supplementary Figs. 3 and 4, and Supplementary Tables 6, 7, 8); 4,569 of the encoded proteins were annotated with gene ontology (GO) terms, and 2,949 proteins were assigned KO (KEGG (Kyoto Encyclopedia of Genes and Genomes) Orthology) identifiers, with 1,928 involved in at least 1 pathway. There were 158 tRNA genes representing 20 amino acids and 5 18S, 3 5.8S and 1 28S rRNA genes. Of the 13,158 repeats, only 933 matched known sequences (Supplementary Table 9). No complete retrotransposons were found, in contrast with the schistosome genomes, which have 20% retrotransposons10,11. Although the E. granulosus genome was sequenced with material from a single cyst (originating from a single egg and thus a clone), we found 145,534 SNP sites, with a density of 0.96 SNPs/kb (Supplementary Tables 10, 11, 12).
E. granulosus had the highest GC content in both its genome (42.1%) and coding regions (49.3%) of the four parasitic helminths and the two free-living nematode taxa with which we compared it. The CpG/expected CpG ratio in E. granulosus genes (0.83) was similar to that of other worms (0.80–1.00) but was much higher than in mammals (0.44 for humans and 0.48 for the domestic dog) (Supplementary Fig. 5). Cytosine methylation in the schistosome genome regulates oviposition12, but the higher CpG content in worms might hint that such methylation occurs much less frequently in these organisms than in mammals. We found only one DNA (cytosine-5)-methyltransferase gene (gene symbol, DNMT3B; gene ID: EG_07014; KO identifier, K00558) and one methyl-CpG–binding domain gene (MBD; EG_02905; K11590) in the E. granulosus genome (Supplementary Table 13), whereas ten DNMT genes are present in humans.
Comparative genomics and features associated with parasitism
We compared the protein domain profiles of E. granulosus with those of six other worms and two mammalian hosts. A total of 6,428 Pfam domains were found in the 9 taxa, with 3,405 identified in E. granulosus, similar to the number of domains in the other 4 parasites but fewer than in the 2 free-living nematodes (Supplementary Figs. 6 and 7 and Supplementary Table 14).
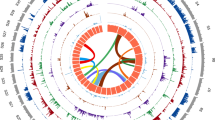
KEGG analysis showed that E. granulosus has complete pathways for glycolysis, the tricarboxylic acid (TCA) cycle and the pentose phosphate pathway (Supplementary Table 15). It lacks the capability for the de novo synthesis of pyrimidines, purines and most amino acids (except for alanine, aspartic acid and glutamic acid) (Fig. 1 and Supplementary Fig. 8), thus relying on the host for these essential nutrients. This feature is supported by its loss of 495 Pfam domains compared with the free-living nematode and mammalian species (Supplementary Fig. 9 and Supplementary Table 16).
Major pathways present, including lipid, nucleotide and amino acid metabolism, are shown with different colored backgrounds. Black solid arrows indicate the presence of enzymes, whereas red dashed arrows indicate their absence. The right upper section shows the transport of bile acids (BA) and the pathways regulated by bile acids. Bile acids are imported by the bile acid transporter (BAT) and may bind and activate the FXR–retinoid X receptor (RXR), which leads to transcriptional activation of the nuclear receptor. Furthermore, bile acids may stimulate the dimerization and activation of VDR-RXR. This binding and activation regulates the expression of genes involved in differentiation, development, homeostasis and metabolism.
Comparing the KEGG ontology terms of E. granulosus and the other parasites with those of Caenorhabditis elegans and Pristionchus pacificus, we found that the former group had 500–550 KEGG ontology terms associated with metabolism, fewer than in the free-living worms (Supplementary Table 17). Enrichment analysis showed that the lost KEGG ontology terms were significantly enriched in 13 pathways (false discovery rate (FDR) < 0.01), all related to metabolism, including, among others, amino acid metabolism, lipid metabolism, biosynthesis of other secondary metabolites, xenobiotic biodegradation and the peroxisome pathway (Supplementary Table 18), further emphasizing the dependence of E. granulosus on its host for key metabolites.
The genome encoded 219 proteases or peptidase-like proteins, including 25 extracellular proteases, 38 cell membrane–associated proteases, 156 intracellular proteases and/or peptidases (Supplementary Table 19), and 68 protein transporters and amino acid transporters (Supplementary Table 20), further implying that E. granulosus complements its lost ability to synthesize important amino acids by obtaining them from the host.
We identified 18 lipases, 10 low-density lipoprotein (LDL) receptors, 1 long-chain fatty acid transport protein and 2 ATP-binding-cassette transporters encoded in the E. granulosus genome sequence (Supplementary Table 21). Similar to in schistosomes11, the genome sequence indicates that E. granulosus cannot generate cholesterol de novo, owing to the absence of several enzymes such as squalene synthase (enzyme code (EC): 2.5.1.21), squalene monooxygenase (EC: 1.14.13.132) (Supplementary Fig. 10). Consequently, cholesterol ester provided by the host would be the only source of cholesterol (Fig. 1), and this notion is supported by the presence of sequences encoding cytoplasmic sterol O-acyltransferase (EG_03337) and transmembrane cholesterol esterase (EG_10760 and EG_11290) in E. granulosus.
Domain families gained in E. granulosus
Only one E. granulosus–specific protein domain family was confirmed—antigen B (EgAgB), a complex of antigens comprising seven members. Expanded domain families in E. granulosus included the heat shock protein 70 (Hsp70), universal stress protein (USP), poly-(ADP-ribose) polymerase (PARP) and prothymosin families (Supplementary Table 22a). Hsp70 proteins have important roles in protein folding and in protecting cells from stress, and expansion of this family in E. granulosus has been reported13. Expression of Hsp70 genes was substantially different in the four life-cycle stages; for example, EG_09650 was only expressed in adult worms, and EG_10561 was highly expressed in the hydatid cyst membrane (Supplementary Table 23). Furthermore, we identified 13 USP genes in E. granulosus compared with the 7–8 identified in schistosomes, whereas none occurred in nematodes (Supplementary Table 22b). USPs are small cytoplasmic proteins associated with stress responses. USP genes are found in urochordates, cnidarians and the Lophotrochozoa (including the Platyhelminthes) but not in the non-urochordate deuterostomes (including mammals) and ecdysozoans (including Nematoda)14, suggesting that the Platyhelminthes have evolved different mechanisms from nematodes to overcome stress. Hsp70 and USP proteins may be involved in stress responses associated with the extremely harsh host environment of the intestinal tract, which has reactive oxygen species (ROS), variable pH and numerous highly active proteases.
There were more dynein light chain (DLC), dynein heavy chain (DHC) and cadherin family members in E. granulosus and schistosomes than in nematodes. Dyneins are motor proteins that act in the force-generating eukaryotic cilia and flagella and in the intracellular retrograde motility of vesicles and organelles, and DLC proteins are associated with transforming growth factor (TGF)-β signaling15. The E. granulosus genome had 48 DLC members (compared to 35 in Schistosoma japonicum and 29 in Schistosoma mansoni), whereas 4–7 were found in the 4 nematodes. We identified 49 cadherins in the E. granulosus genome, similar to the numbers found in schistosomes (51–65) but more than in nematodes (12–18) (Supplementary Table 22b). Cadherins belong to a class of type 1 transmembrane proteins that have important roles in cell recognition and adhesion. They localize to the cell membrane, interacting with another cadherin subtype on an adjacent cell in a zipper-like fashion, and may have a role in invasion16,17. Overall, expansions in these protein domain in E. granulosus seem to represent another adaptation to parasitism.
Orthologs in parasitic helminths as potential intervention targets
Comparing the protein domains present in both E. granulosus and other sequenced parasitic helminths, we did not find any shared functional annotations suggesting a common association with parasitism. However, using ortholog analysis, we found 33 ortholog groups (represented by 42 E. granulosus genes) uniquely present in parasitic helminths (Supplementary Fig. 11 and Supplementary Tables 24, 25, 26). These genes may be of interest as potential drug, vaccine or diagnostic targets.
E. granulosus and the other parasitic worms encode a special orthologous group of prenylcysteine oxidases (EG_06057), which may catalyze the final step in the degradation of prenylated proteins. Protein prenylation has been studied extensively in parasites, with prenyltransferase inhibitors found to inhibit the differentiation and growth of several species18. Prenyltransferase is a key enzyme in the biosynthesis of prenylated proteins and, along with prenylcysteine oxidase, may represent a novel drug target.
Another parasite-specific ortholog group encoded proteins similar to the CD151 antigen (tetraspanin). Notably, two tetraspanins from S. mansoni are protective when used in vaccines in mice, and there is a strong IgG-mediated response to tetraspanin in individuals naturally resistant to S. mansoni19. Further, RNA interference studies suggest that tetraspanins have important structural roles in S. mansoni tegument development, maturation or stability20, which may also be the case in E. granulosus.
Bidirectional development
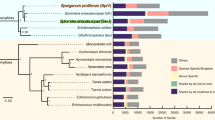
The gene expression profile of E. granulosus suggests that upregulated genes have important roles in controlling and maintaining stage-specific features of the parasite during its life cycle (Fig. 2, Supplementary Fig. 12, Supplementary Tables 27–29 and Supplementary Note). A striking feature of the biology of E. granulosus is that the PSC has the potential to develop in either of two directions. Larvae ingested by a dog will develop in a sexual direction to form adult tapeworms in the gut. In contrast, if a hydatid cyst ruptures within the intermediate or human host, each released PSC is capable of differentiating asexually into a new hydatid cyst, meaning that 'secondary' hydatidosis results (Fig. 2).
(a) Percentage of reads of the top 50 upregulated genes and stage-specific genes in the 4 developmental stages of E. granulosus. Genes were considered as upregulated (fold change > 2; P < 0.00001) in one stage compared with in two other stages (adult versus PSC and oncosphere; oncosphere versus adult and hydatid cyst; hydatid cyst versus oncosphere and PSC; PSC versus adult and hydatid cyst). The percentage of expressed genes upregulated in each stage is equal to the total reads of upregulated genes or specific genes divided by the total transcript reads in each stage times 100%. (b) Distribution of transcripts in the PSC, adult and hydatid cyst putatively involved in the bidirectional development of the PSC into either an adult worm or a secondary hydatid cyst. (c) Genes transcribed in the adult, oncosphere and hydatid cyst stages. Numbers in blue indicate the number of genes transcribed in each stage. The number with an asterisk indicates the number of genes significantly upregulated in each stage compared with in the immediately linked stage(s) in the life cycle (P < 0.00001). The number in red represents uniquely expressed upregulated genes in each stage compared with in immediately linked stage(s). The statistical value was calculated by the MA-plot-based method with a random sampling model.
It has been shown that bile acids have a crucial role in the differentiation of PSCs into adult worms6, and E. granulosus may express bile acid receptors and transporters to stimulate the relevant pathways (Fig. 1). The two major kinds of bile acid signaling receptors are represented by TGR5, a G protein–coupled receptor (GPCR)21, and members of the nuclear hormone receptor superfamily, including the farnesoid X receptor (FXR)22. Although we identified several downstream signal transduction components of the bile acid pathways, we found no TGR5-like receptors in the E. granulosus genome or transcriptome. We identified four genes as candidate nuclear hormone receptors for bile acid signals (EG_00119, EG_00780, EG_04405 and EG_08428), which encode proteins that have more than 20% amino acid identity with FXR and the vitamin D receptor (VDR) and contain a DNA-binding domain and a ligand-binding domain. We also found genes encoding five sodium–bile acid cotransporters and seven multidrug resistance proteins (MRPs)23, as well as genes associated with bile acid metabolism, including sterol regulatory element–binding protein 1 and bile acid β-glucosidase–related proteins (Supplementary Table 30). Ecdysone or other sex steroids might regulate molting in parasitic nematodes through nuclear hormone receptors24,25, and it is reasonable to assume that exogenous bile acid from the host has an important role in the development of E. granulosus in a similar fashion.
The nuclear receptors FXR and VDR are less sensitive to their physiological ligands (EC50 (half-maximal effective concentration) of ∼10 μM) than the membrane receptor TGR5 (EC50 of ∼300–600 nM)26, and this difference in sensitivity supports the observation that E. granulosus only develops into an adult worm in the presence of high concentrations of bile acid, such as are found in the dog intestine. Although current knowledge does not exclude the possibility of an unknown, novel GPCR bile acid receptor in E. granulosus, FXR-like nuclear receptors, which are more conserved among species, likely have a role in its bile acid signaling process.
The PSC is more complex in structure than the thin hydatid cyst, and secondary cyst development from the PSC is likely a process of dedifferentiation6. We identified 356 genes upregulated in the PSC compared with in the hydatid cyst (Fig. 2), including 45 associated with signal transmission. In addition, 28 genes upregulated in the PSC were completely silenced in the hydatid cyst (Supplementary Table 31).
Strobilization and reproduction
E. granulosus undergoes both sexual and asexual reproduction. Adult worms sexually produce eggs in each gravid proglottid, which in turn are replicated through strobilization6. Hsp90-like protein (EG_10560) was highly expressed in adults and hydatid cysts (Supplementary Table 32). Hsp90-mediated homeostasis controls stage differentiation in Leishmania donovani27, suggesting that Hsp90 may have a role in strobilization in adult E. granulosus, along with other proteins such as that encoded by the segmentation gene fushi tarazu (ftz-f1; EG_10234).
The E. granulosus genome contains a range of genes associated with segmentation, including, among others, Hox genes, arm/catenin, nanos homolog 1, pair-rule genes and tailless (Supplementary Table 33). Homologs of genes involved in sexual reproduction in C. elegans28,29 were also identified, including ones associated with meiosis, spermatogenesis or oogenesis, fertilization and egg development (Supplementary Table 34).
Gene ontology and KEGG pathway analyses showed that E. granulosus possesses most of the key molecules involved in the meiotic pathway. We identified 20 meiosis-associated components, including early meiotic induction protein (EG_00791), meiotic recombination protein rec8 (mre8; EG_04509), an mre11 homolog (EG_07425) and a meiotic nuclear division protein 1 homolog (EG_01539) (Supplementary Table 35). Egg surface LDL receptor repeat–containing protein (EGG) is a member of a protein family localized to the plasma membrane of the oocyte that is necessary for fertilization30. We identified a gene (EG_08608) homologous to EGG encoding a type II transmembrane molecule with the extracellular domains including eight LDL receptor repeats that are known to function as receptors for a variety of ligands and to mediate multiple cellular responses30.
Signaling pathways
Wed found that E. granulosus possesses several complete signaling pathways, including for mitogen-activated protein kinase (MAPK), ErbB, Wnt, Notch, Hedgehog, TGF-β, Jak-Stat and insulin signaling (Supplementary Table 36). The genome encoded fibroblast growth factor receptor (FGFR; EG_04208), epidermal growth factor receptor (EGFR; EG_03329) and insulin-like growth factor receptor (IGFR; EG_02146 and EG_02635) (Table 2), but no genes for FGF, EGF or IGF were identified. The Lin-3 gene encodes EGF in C. elegans31, but we did not find a homolog of Lin-3 in E. granulosus. The identities of the E. granulosus components acting on these receptors are unclear, and the parasite may use host signaling proteins, sharing the same common signaling components and pathways as its host, to signal cell proliferation, differentiation and even cell death during organogenesis and tissue development. We found 11 cytokine receptors, 12 nuclear receptors and 66 GPCRs involved in regulating numerous cellular processes (Supplementary Table 37). E. granulosus expressed many proteins (n = 192) responsible for cell interaction (Supplementary Table 38), which may function in host-parasite cross-talk (Supplementary Note).
Neuroendocrine and nervous systems
Most hormones and receptors associated with the classical neuroendocrine hypothalamus-pituitary–peripheral endocrine gland axis were absent in E. granulosus (Supplementary Table 39 and Supplementary Note), although two putative receptors of the hypothalamus-pituitary-thyroid axis were found. One gene (EG_07666) had high sequence similarity with the pituitary thyrotropin-releasing hormone receptors (TRHRs) in humans and domestic dogs; another (EG_08053) was similar to the thyroid hormone receptor (THR) α isoform 1 of these mammalian hosts, but thyroid-stimulating hormone receptor (TSHR) was absent. The neuroendocrine system of E. granulosus is simple and incomplete compared with that of S. japonicum11. In addition, we identified 92 genes encoding sensory system elements (Supplementary Table 40), including homologs associated with taste and smell, such as olfactory receptor, G protein and adenylate cyclase type 3.
Evading immune recognition and regulation of host immunological responses
A hallmark of E. granulosus is its prolonged survival—up to 53 years in humans32—in many mammalian host species, indicating that it selectively produces components to moderate the host immune response, thereby enabling escape from host attack33. E. granulosus–specific EgAgB is likely to be a key factor in the process of immune evasion, as the antigen is secreted and variable34. Encoded by a gene family, we found seven EgAgB genes in the genome, similar to in a previous report13.
Other evasion strategies include the production and release of proteases (Supplementary Table 19) to digest host proteins and protease inhibitors to avoid host protease digestion. We found 1,965 predicted proteins in the genome with signal peptides, of which 809 were extracellular or secreted (Supplementary Table 41). These proteins likely serve as messengers for cross-talk between E. granulosus and its hosts, having key roles in regulating host immune responses. Secreted products from E. granulosus, expressed highly in the adult, oncosphere and hydatid cyst (Supplementary Table 42), may affect the host immune system by influencing the cytokine network and signal transduction pathways or by inhibiting essential enzymes, resulting in immune regulation through suppression, diversion or alteration of the host immune response. This process may provide an anti-inflammatory environment that is favorable for parasite survival35.
Unlike in schistosomes, no genes associated with the Toll-like receptor pathway were identified in E. granulosus. However, we found that E. granulosus possesses a range of genes encoding molecules associated with innate immune defense mechanisms. These included leukotriene A4 hydrolase (LTA4H; EG_02555), a chemoattractant (EG_09475), cytokines, macroglobulin, tumor necrosis factor (TNF) receptor, glycan, mannosyltransferase, prostaglandins, lipooxygenase, beige beach and histone H2A (Supplementary Table 43). In addition, the genome contained 25 leucine-rich repeat (LRR)-containing proteins and lectin-like proteins (EG_07089, EG_04345 and EG_10148), which have the potential to act in the recognition and clearance of microbes36, an important feature, given that the adult worms of E. granulosus live in the canine intestine, which harbors countless number of microorganisms.
New intervention targets
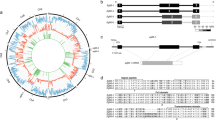
Praziquantel is a highly effective drug in killing adult worms of schistosomes and E. granulosus. It is hypothesized to act directly or indirectly on calcium channel β (Cavβ) subunits in schstosomes37. Given that humans, mice, rats and rabbits can accommodate high quantities of praziquantel with only mild side effects38, putative sites related to praziquantel sensitivity in worms may be located in conserved functional domains that have different sequences than in the mammalian hosts. We found EgCavβ1 (EG_10568) and EgCavβ2 (EG_04487), the homologs of schistosome SmCavβ1 and SmCavβ2, which contain a Src-homology 3 domain (SH3), a guanylate kinase domain (GK) and a β-interaction domain (BID). The BID domain has been suggested to be the region whereby the Cavβ and Cavα subunits interact39. Additionally, two non-functional putative serine phosphorylation sites in this region of wild-type SmCavβ1 have been linked to praziquantel action, as mutations affecting either of the two wild-type sites result in the elimination of sensitivity37. Phylogenetic analysis confirmed in detail the previous suggestion that the E. granulosus Cavβ1 proteins are encoded by different orthologs, whereas Cavβ2 and the host Cavβ proteins represent one orthologous group (Fig. 3a). The conserved sequences of the E. granulosus BID phosphorylation sites further support the hypothesis of non-functional phosphorylation sites (Fig. 3b). However, the crystal structure of rat Cavβ indicates that the BID phosphorylation sites have very low or no relative accessibility and thus are unlikely to interact with protein kinases under the proposed protein conformation. Instead of BID, a Cavα-binding pocket (ABP) has been suggested as the region whereby the Cavβ and Cavα subunits interact39.
(a) SH3 domain, BID and GK domain sequences encoded by Cavβ genes from ten species (gene identification and ortholog information are provided in the Supplementary Note) were used to generate a neighbor-joining phylogenetic tree. The numbers at the branches are confidence values (percentage) calculated based on the bootstrap method. The two E. granulosus proteins EgCavβ1 and EgCavβ2 are in two groups with their schistosome homologs. EgCavβ1 is more distant from the Cavβ genes of mammals and the Nematoda, which have low or no sensitivity to praziquantel. (b) Two non-functional putative serine phosphorylation sites in the BID of Cavβ are denoted by asterisks. Neither of the two serine residues occurs in EgCavβ1 or its homologs SmCavβ1 and SjCavβ1 in the schistosomes. Although the two serine residues are highly conserved in SmCavβ2 and SjCavβ2 as well as in the human, rabbit, bovine, dog and C. elegans Cavβ subunits, there is one serine residue transformed to threonine in EgCavβ2. (c) Another phosphorylation site (indicated by an asterisk) near the ABP of Cavβ might have a role in the interaction of the Cavβ and Cavα subunits. The isoleucine and arginine residues downstream of this serine residue in EgCavβ1 are similar in SmCavβ1 and SjCavβ1 but are different in the other species.
We noticed another reported putative phosphorylation site near the ABP with a relative accessibility of 35.4%, which is reduced to 12% when Cavβ binds to the α-interaction domain (AID) of Cavα. It is noteworthy that the region containing this phosphorylation site may also be dysfunctional in the Cavβ1 subunit of E. granulosus and schistosomes, being recognized by protein kinases other than the common kinases of Cavβ2 in worms and the host Cavβ subunits (Fig. 3c). This finding implies that other praziquantel-sensitive sites may exist, possibly representing a different mechanism of indirect interaction between Cavβ and praziquantel. It is of note that, unlike in schistosomes, praziquantel is poorly effective or ineffective40 against the liver fluke Faciola hepatica, which has no Cavβ homologs present in its transcriptome41.
Current treatment of hydatid disease involves surgery and the use of benzimidazole drugs, but the results are far from satisfactory, and new compounds for the treatment of cystic echinococcosis are urgently needed. By examining the genome, we identified a number of possible new druggable targets for echinococcosis (n = 72) (Supplementary Tables 44 and 45), including GPCRs, serine-threonine and tyrosine protein kinases, serine proteases and nuclear hormones, which are the targets of successful new drugs discovered in recent years42.
Ion channels may prove to be additional attractive targets for future anthelmintic development43. We identified genes encoding 29 ligand-gated ion channels, 39 voltage-gated cation channels, 5 chloride channels and 9 other types of channels in the E. granulosus genome (Supplementary Table 46). The ligand-gated ion channels included 13 Cys-loop superfamily proteins, 6 glutamate-activated cation channels, 2 epithelial sodium channels and 2 ATP-gated ion channels. Among these, seven nicotinic acetylcholine receptors of the Cys-loop superfamily constituted the largest subfamily.
The EG95 vaccine has been shown to induce almost complete protection in sheep against E. granulosus challenge infection, and homologs from other taeniid worms induce similar levels of protective efficacy44. We identified seven genes encoding EG95 and others, such as protease inhibitors and tetraspanins, that were highly and specifically expressed in oncospheres and likely represent additional vaccine candidates for echinococcosis (Supplementary Table 47). In addition, the most relevant diagnostic target molecules in E. granulosus were secreted proteins, including EgAgB, antigen 5, EG10 and TPx, which have already shown some promise in serodiagnosis33 (Supplementary Table 48).
Discussion
The genome of E. granulosus is one of the first tapeworm genomes to be sequenced. The work presented here provides a model for future studies on evolution, genomic architecture in general and the biology of bidirectional development-differentiation and segmentation-strobilization. E. granulosus has lost a range of genes associated with lipid and amino acid synthesis but has acquired others with potential for biasing the host immune response and for inducing host production of cytokines and other factors that are beneficial for parasite growth and survival.
The genome and transcriptome data will provide a platform, not only for deeper understanding of the molecular biology and physiology of E. granulosus and for illuminating mechanisms of pathogenesis in echinococcosis, but also for developing new public health interventions against hydatid disease, given the inefficiencies of currently available drugs, the lack of appropriate diagnostic procedures and the current difficulties in treatment and control45.
We obtained similar findings for E. granulosus to those reported by Tsai et al.8, including expansion of the Hsp70-domain family and the lack of some key metabolic pathways for the synthesis of several amino acids, fatty acids and cholesterol. However, in addition to assembled scaffolds, we identified more repetitive sequences from unassembled contigs and obtained a higher ratio of repeats (30.25% or 45.86 Mb) and a larger genome size for E. granulosus. As well as providing an invaluable resource to facilitate the development of much needed new treatments and tools for the control of echinococcosis, these two independent reports will provide a comprehensive basis for exploring basic questions on the biology and evolution of the Gestoda.
Methods
Parasites.
All E. granulosus materials were collected from the Xinjiang Uyghur Autonomous Region, China. For genome sequencing, we collected a large unilocular cyst of 11 cm in diameter from a sheep liver and completely removed the cyst. After rinsing ten times with PBS, the cyst was opened to collect internal materials. We obtained 9 ml of precipitated PSCs and brood capsules. We also stirred the cyst wall to release germinal cells and membranes. All cyst materials were combined, mixed and used for direct extraction of genomic DNA for sequencing.
For transcriptome sequencing, we prepared four stages of E. granulosus, including PSCs, cyst germinal cells and membranes, adult worms and oncospheres. PSCs and brood capsules were aspirated from hydatid cysts. Capsules and PSCs were washed once with PBS and then treated with 0.025% pepsin (Sigma) in Hanks' balanced salt solution (HBSS, pH 2.0) for 15 min. Pepsin-activated PSCs were washed three times with PBS and then soaked in 10 volumes of RNAlater (Sigma).
In preparing cyst germinal cells and membranes, we washed the cyst wall ten times with PBS after PSCs and brood capsules were aspirated from the cyst, checking that there were no PSCs in the wash buffer. The cyst wall was then cut into small pieces that were stirred quickly in a flask with PBS and a glass bar to release germinal membranes and cells from the cyst wall. After sedimentation at 4 °C for 3 min, the suspension was transferred into centrifuge tubes. After centrifugation, pellets were resuspended immediately in 10 volumes of RNAlater for RNA extraction.
Mature adult worms were collected 62 d after infection from dogs experimentally infected with PSCs. The use of dogs was approved by the Animal Ethics Committee of the Xinjiang Academy of Animal Science. Worms were washed with PBS and then soaked in 10 volumes of RNAlater before extraction of total RNA.
In preparing activated larval oncospheres, we released eggs by homogenizing the mature adult worms in an electric blender. Homogenate was passed through a 132-μm sieve, and sheared worm material was discarded after we thoroughly rinsed worm tissues through the sieve. Eggs were further washed and retained on 20-μm mesh. Washed eggs were stored in PBS containing 100 IU/ml benzyl penicillin and 100 μg/ml streptomycin sulfate at 4 °C.
Eggs were incubated in 50-ml screw-cap tubes at 37 °C for 45 min in a sterile solution of 1% pepsin (Sigma) and 1% HCl in 0.85% NaCl. After centrifugation (500g for 5 min), the pepsin solution was decanted. Eggs were washed once with PBS and incubated in a sterile solution of 1% pancreatin (Sigma, 4× US Pharmacopeia), 1% NaHCO3 and 5% sterile sheep bile. Oncospheres were checked every 2 min with a microscope until all had been released from embryonic membranes. Oncospheres were pelleted by centrifugation (1,000g for 5 min). The supernatant was discarded, and oncospheres were washed twice with HBSS.
Oncospheres were further purified by density-gradient separation in 100% Percoll (Sigma)46. After oncospheres were washed three times with PBS, the supernatant was discarded, and pelleted oncospheres were stored with 10 volumes of RNAlater for total RNA extraction.
DNA isolation, library construction and sequencing.
Genomic DNA was isolated using a standard phenol-chloroform method. A shotgun library of fragments of 300–800 bp in length was prepared from 5 μg of DNA using a standard GS FLX shotgun library protocol. A total of 7,503,355 reads with an average length of 378 bp were produced by Roche 454 GS FLX, providing 18.7-fold coverage of the E. granulosus genome. The 300-bp paired-end library was constructed using a standard Solexa paired-end protocol, and 55,012,872 pairs of 120-bp reads were produced on the Illumina Genome Analyzer platform, providing 83.5-fold genome coverage. The 3-kb mate-pair library was constructed combining the GS FLX and Solexa mate-pair protocol, with an adaptor sequence inserted between the mate-pair reads. A total of 116,732,731 mate-pair reads of 35 bp in length were generated on the Illumina Genome Analyzer platform, providing 53.9-fold genome coverage.
Genome assembly.
Roche 454 reads were first assembled using Newbler v2.3, and 22,340 contigs with an average length of 5,004 bp were produced. Solexa paired-end reads were then mapped to the contigs to increase sequencing quality. Solexa mate-pair reads (insert size of 3 kb) were used to establish genome scaffolds. There were more than 23 million pairs for which the reads mapped to different contigs. A simple greedy algorithm was used to optimize the order of the contigs and to provide a feasible heuristic solution for scaffold construction47. The 9,974 contigs were then linked and used to generate 967 genome scaffolds with a maximum length of 3,893,204 bp and a total size of 110,862,006 bp.
Identification of repeats.
Repeat families were identified using RepeatScout48 with default parameters. Tandem repeats were identified using TANDEM REPEATS FINDER (version 4.07b)49 and categorized using TRAP (version 1.1)50. Microsatellites, minisatellites and satellites were classically defined as repeat units of 2–6 bp, 7–100 bp and more than 100 bp, respectively.
Gene prediction and annotation.
Exonhunter51, Genemark (v2.3a)52 and Augustus (v2.5)53 were used to predict genes. A gene model was then produced by combining the three prediction results with the help of the 'GLAD' program, which was developed in house, a tool that creates consensus gene lists by integrating evidence from homology, de novo prediction and RNA sequencing and/or EST data. Finally, a total of 11,325 protein-encoding genes were predicted from the genome. Annotation was performed by comparing predicted proteins with the non-redundant protein database (nr), UniProt and the KEGG database. Pathway construction and functional classification were preformed with the KEGG database54. Blast2GO55 and InterProScan56 were used separately to assign preliminary GO terms and domains to predicted gene models. GPCRs were identified by searching the IPR domain (IPR000276) in addition to KEGG classification. Proteases and protease inhibitors were identified using BLASTP against the MEROPS database57 with E value < 1 × 10−5.
Calcium channel β subunit analysis.
The SH3, BID and GK domain sequences of the calcium channel β (Cavβ) subunits from ten species were used to generate a neighbor-joining phylogenetic tree with MEGA 5 (1,000 bootstrap replications)58. These subunits included EgCavβ1 and EgCavβ2 from E. granulosus; BmCavβ1 (XP_001902270) from Brugia malayi; BtCavβ1 (NP_787013), BtCavβ2 (NP_786983), BtCavβ3 (NP_776934) and BtCavβ4 (NP_001179033) from Bos taurus; CeCavβ1 (NP_491193) and CeCavβ2 (NP_491055) from C. elegans; ClfCavβ1 (XP_548150), ClfCavβ2 (XP_855770), ClfCavβ3 (XP_543689) and ClfCavβ4 (XP_851697) from Canis lupus familiaris; HsCavβ1 (NP_954855), HsCavβ2 (NP_963890), HsCavβ3 (NP_001193846) and HsCavβ4 (NP_000717) from Homo sapiens; OcCavβ1 (NP_001075748), OcCavβ2 (NP_001075865) and OcCavβ3 (NP_001095185) from Oryctolagus cuniculus; SjCavβ1 (AAK51116) and SjCavβ2 (CAX82734) from S. japonicum; SmCavβ1 (AAK51117) and SmCavβ2 (AAK51118) from S. mansoni; and TsCavβ1 (EFV52876) and TsCavβ2 (EFV54005) from Trichinella spiralis. On the basis of OrthoMCL DB classification59, EgCavβ1, SjCavβ1 and SmCavβ1 were clustered into OG5_246029, CeCavβ1 was clustered into OG5_217851, and EgCavβ2 and all other Cavβ subunits of mammalian, nematode and schistosome origin were clustered into the same ortholog OG5_128949. Multiple-sequence alignment was performed using ClustalX 2.1 (ref. 60).
RNA extraction.
Total RNA was extracted from E. granulosus materials using TRIzol Reagent (Life Technologies). mRNA was extracted from total RNA using an Oligotex mRNA Mini kit (Qiagen). mRNA was then precipitated by adding 0.1 volumes of 3 M sodium acetate (pH 5.2) and 0.8 volumes of isopropanol. Tubes were kept at −20 °C overnight and then centrifuged at 12,000g for 30 min at 4 °C. Pellets were washed with 70% ethanol, air dried at room temperature for 10–15 min and dissolved in 20 μl of DEPC-treated water. The resulting mRNA was used for cDNA library construction.
EST sequencing.
Double-stranded cDNA was synthesized according to the full-length cDNA synthesis protocol of Ng et al.61 and then fragmented to 300–800 bp for Roche 454 sequencing. A total of 561,998 ESTs with an average length of 300 bp were produced from the 4 constructed cDNA libraries.
Transcriptome data analysis.
To validate the expression of predicted genes, 561,998 ESTs representing the 4 different life-cycle stages were compared with the predicted genes. After trimming low-quality (Q < 20) bases from both ends, we compared reads with predicted genes using BLASTN with criteria settings as 50% coverage and 95% identity. Finally, 8,336 genes (73.6%) were determined to have EST expression in the 4 stages (at least 2 ESTs observed in 1 gene). The expressed read number for each gene was first transformed into RPKM (reads per kilobase per million reads)62, and differently expressed contigs were then identified by the DEGseq package using MARS (MA plot–based method with Random Sampling model)63. Enrichment of KEGG pathways for significantly expressed genes (or a given gene list) was calculated using a classical hypergeometric distribution statistical comparison of the query gene list against all predicted E. granulosus genes (or a reference gene list). Calculated P values were subjected to FDR correction (Benjamini and Hochberg), taking a corrected P value of <0.01 as the threshold for significance.
URLs.
E. granulosus genome assembly contigs and scaffolds, complete sequences from 19 fosmids and transcriptome sequences can be downloaded from ftp sites of the Chinese National Human Genome Center at Shanghai (CHGCS; http://chgc.sh.cn/Eg).
Accession codes.
E. granulosus genome assembly contigs and scaffolds, complete sequences for 19 fosmids and transcriptome sequences have been deposited in GenBank or the Sequence Read Archive (SRA) (E. granulosus genome, APAU00000000; fosmids, KC585039, KC585040, KC585041, KC585042, KC585043, KC585044, KC585045, KC585046, KC585047, KC585048, KC585049, KC585050, KC585051, KC585052, KC585053, KC585054, KC585055, KC585056, KC585057; ESTs, SRA066120). Sequences and functional annotations of E. granulosus protein-encoding genes, including of predicted genes, are available from the NCBI and CHGCS websites.
References
Garcia, H.H. & Del Brutto, O.H. Neurocysticercosis: updated concepts about an old disease. Lancet Neurol. 4, 653–661 (2005).
McManus, D.P., Zhang, W., Li, J. & Bartley, P.B. Echinococcosis. Lancet 362, 1295–1304 (2003).
Craig, P.S. et al. Prevention and control of cystic echinococcosis. Lancet Infect. Dis. 7, 385–394 (2007).
Li, T. et al. Post-treatment follow-up study of abdominal cystic echinococcosis in tibetan communities of northwest Sichuan Province, China. PLoS Negl. Trop. Dis. 5, e1364 (2011).
Moro, P.L. et al. Human hydatidosis in the central Andes of Peru: evolution of the disease over 3 years. Clin. Infect. Dis. 29, 807–812 (1999).
Smyth, J.D. in In Vitro Cultivation of Parasitic Helminths (ed. Smyth, J.D.) 77–154 (CRC Press, Boca Raton, FL, 1990).
Fiori, P.L. et al. Establishment of cell cultures from hydatid cysts of Echinococcus granulosus. Int. J. Parasitol. 18, 297–305 (1988).
Tsai, I.J. et al. The genomes of four tapeworm species reveal adaptations to parasitism. Nature 496, 57–63 (2013).
McManus, D.P. Molecular discrimination of taeniid cestodes. Parasitol. Int. 55 (suppl.), S31–S37 (2006).
Berriman, M. et al. The genome of the blood fluke Schistosoma mansoni. Nature 460, 352–358 (2009).
Schistosoma japonicum Genome Sequencing and Functional Analysis Consortium. The Schistosoma japonicum genome reveals features of host-parasite interplay. Nature 460, 345–351 (2009).
Geyer, K.K. et al. Cytosine methylation regulates oviposition in the pathogenic blood fluke Schistosoma mansoni. Nat. Commun. 2, 424 (2011).
Olson, P.D., Zarowiecki, M., Kiss, F. & Brehm, K. Cestode genomics—progress and prospects for advancing basic and applied aspects of flatworm biology. Parasite Immunol. 34, 130–150 (2012).
Forêt, S. et al. Phylogenomics reveals an anomalous distribution of USP genes in metazoans. Mol. Biol. Evol. 28, 153–161 (2011).
Tang, Q. et al. A novel transforming growth factor-β receptor–interacting protein that is also a light chain of the motor protein dynein. Mol. Biol. Cell 13, 4484–4496 (2002).
Lauwaet, T., Oliveira, M.J., Mareel, M. & Leroy, A. Molecular mechanisms of invasion by cancer cells, leukocytes and microorganisms. Microbes Infect. 2, 923–931 (2000).
Bouchut, A., Roger, E., Coustau, C., Gourbal, B. & Mitta, G. Compatibility in the Biomphalaria glabrata/Echinostoma caproni model: potential involvement of adhesion genes. Int. J. Parasitol. 36, 175–184 (2006).
Jortzik, E., Wang, L. & Becker, K. Thiol-based posttranslational modifications in parasites. Antioxid. Redox Signal. 17, 657–673 (2012).
Tran, M.H. et al. Tetraspanins on the surface of Schistosoma mansoni are protective antigens against schistosomiasis. Nat. Med. 12, 835–840 (2006).
Tran, M.H. et al. Suppression of mRNAs encoding tegument tetraspanins from Schistosoma mansoni results in impaired tegument turnover. PLoS Pathog. 6, e1000840 (2010).
Keitel, V. & Haussinger, D. Perspective: TGR5 (Gpbar-1) in liver physiology and disease. Clin. Res. Hepatol. Gastroenterol. 36, 412–419 (2012).
Lefebvre, P., Cariou, B., Lien, F., Kuipers, F. & Staels, B. Role of bile acids and bile acid receptors in metabolic regulation. Physiol. Rev. 89, 147–191 (2009).
Hirohashi, T., Suzuki, H., Takikawa, H. & Sugiyama, Y. ATP-dependent transport of bile salts by rat multidrug resistance–associated protein 3 (Mrp3). J. Biol. Chem. 275, 2905–2910 (2000).
Hernández-Bello, R. et al. Sex steroids effects on the molting process of the helminth human parasite Trichinella spiralis. J. Biomed. Biotechnol. 2011, 625380 (2011).
Tzertzinis, G. et al. Molecular evidence for a functional ecdysone signaling system in Brugia malayi. PLoS Negl. Trop. Dis. 4, e625 (2010).
Fiorucci, S., Mencarelli, A., Palladino, G. & Cipriani, S. Bile-acid-activated receptors: targeting TGR5 and farnesoid-X-receptor in lipid and glucose disorders. Trends Pharmacol. Sci. 30, 570–580 (2009).
Wiesgigl, M. & Clos, J. Heat shock protein 90 homeostasis controls stage differentiation in Leishmania donovani. Mol. Biol. Cell 12, 3307–3316 (2001).
White-Cooper, H. & Bausek, N. Evolution and spermatogenesis. Phil. Trans. R. Soc. Lond. B 365, 1465–1480 (2010).
Nishimura, H. & L'Hernault, S.W. Spermatogenesis-defective (spe) mutants of the nematode Caenorhabditis elegans provide clues to solve the puzzle of male germline functions during reproduction. Dev. Dyn. 239, 1502–1514 (2010).
Kadandale, P. et al. The egg surface LDL receptor repeat–containing proteins EGG-1 and EGG-2 are required for fertilization in Caenorhabditis elegans. Curr. Biol. 15, 2222–2229 (2005).
Liu, G., Rogers, J., Murphy, C.T. & Rongo, C. EGF signalling activates the ubiquitin proteasome system to modulate C. elegans lifespan. EMBO J. 30, 2990–3003 (2011).
Spruance, S.L. Latent period of 53 years in a case of hydatid cyst disease. Arch. Intern. Med. 134, 741–742 (1974).
Zhang, W., Li, J. & McManus, D.P. Concepts in immunology and diagnosis of hydatid disease. Clin. Microbiol. Rev. 16, 18–36 (2003).
Kamenetzky, L. et al. High polymorphism in genes encoding antigen B from human infecting strains of Echinococcus granulosus. Parasitology 131, 805–815 (2005).
Sher, A., Wynn, T.A. & Sacks, D.L. in The Immune Response to Parasites (ed. Paul, W.E.) 1171–1200 (Lippincott, Williams & Wilkins, Philadelphia, 2003).
Zugasti, O. & Ewbank, J.J. Neuroimmune regulation of antimicrobial peptide expression by a noncanonical TGF-β signaling pathway in Caenorhabditis elegans epidermis. Nat. Immunol. 10, 249–256 (2009).
Greenberg, R.M. Are Ca2+ channels targets of praziquantel action? Int. J. Parasitol. 35, 1–9 (2005).
Xiao, S. et al. Early treatment with artemether and praziquantel in rabbits repeatedly infected with Schistosoma japonicum cercariae. Zhongguo Ji Sheng Chong Xue Yu Ji Sheng Chong Bing Za Zhi 12, 252–256 (1994).
Van Petegem, F., Clark, K.A., Chatelain, F.C. & Minor, D.L. Jr. Structure of a complex between a voltage-gated calcium channel β-subunit and an α-subunit domain. Nature 429, 671–675 (2004).
Patrick, D.M. & Isaac-Renton, J. Praziquantel failure in the treatment of Fasciola hepatica. Can. J. Infect. Dis. 3, 33–36 (1992).
Young, N.D., Hall, R.S., Jex, A.R., Cantacessi, C. & Gasser, R.B. Elucidating the transcriptome of Fasciola hepatica—a key to fundamental and biotechnological discoveries for a neglected parasite. Biotechnol. Adv. 28, 222–231 (2010).
Hopkins, A.L. & Groom, C.R. The druggable genome. Nat. Rev. Drug Discov. 1, 727–730 (2002).
Robertson, A.P. & Martin, R.J. Ion-channels on parasite muscle: pharmacology and physiology. Invert. Neurosci. 7, 209–217 (2007).
Gauci, C., Heath, D., Chow, C. & Lightowlers, M.W. Hydatid disease: vaccinology and development of the EG95 recombinant vaccine. Expert Rev. Vaccines 4, 103–112 (2005).
McManus, D.P., Gray, D.J., Zhang, W. & Yang, Y. Diagnosis, treatment, and management of echinococcosis. Br. Med. J. 344, e3866 (2012).
Rajasekariah, G.R., Rickard, M.D. & Mitchell, G.F. Density-gradient separation of Taenia pisiformis oncospheres. J. Parasitol. 66, 355–356 (1980).
Kim, P.G., Cho, H.G. & Park, K. A scaffold analysis tool using mate-pair information in genome sequencing. J. Biomed. Biotechnol. 2008, 675741 (2008).
Price, A.L., Jones, N.C. & Pevzner, P.A. De novo identification of repeat families in large genomes. Bioinformatics 21 (suppl. 1), i351–i358 (2005).
Benson, G. Tandem repeats finder: a program to analyze DNA sequences. Nucleic Acids Res. 27, 573–580 (1999).
Sobreira, T.J., Durham, A.M. & Gruber, A. TRAP: automated classification, quantification and annotation of tandemly repeated sequences. Bioinformatics 22, 361–362 (2006).
Brejová, B. et al. Finding genes in Schistosoma japonicum: annotating novel genomes with help of extrinsic evidence. Nucleic Acids Res. 37, e52 (2009).
Lomsadze, A., Ter-Hovhannisyan, V., Chernoff, Y.O. & Borodovsky, M. Gene identification in novel eukaryotic genomes by self-training algorithm. Nucleic Acids Res. 33, 6494–6506 (2005).
Stanke, M. & Waack, S. Gene prediction with a hidden Markov model and a new intron submodel. Bioinformatics 19 (suppl. 2), ii215–ii225 (2003).
Kanehisa, M., Goto, S., Kawashima, S., Okuno, Y. & Hattori, M. The KEGG resource for deciphering the genome. Nucleic Acids Res. 32, D277–D280 (2004).
Conesa, A. & Gotz, S. Blast2GO: a comprehensive suite for functional analysis in plant genomics. Int. J. Plant Genomics 2008, 619832 (2008).
Quevillon, E. et al. InterProScan: protein domains identifier. Nucleic Acids Res. 33, W116–W120 (2005).
Rawlings, N.D., Barrett, A.J. & Bateman, A. MEROPS: the database of proteolytic enzymes, their substrates and inhibitors. Nucleic Acids Res. 40, D343–D350 (2012).
Tamura, K. et al. MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol. Biol. Evol. 28, 2731–2739 (2011).
Chen, F., Mackey, A.J., Stoeckert, C.J. Jr. & Roos, D.S. OrthoMCL-DB: querying a comprehensive multi-species collection of ortholog groups. Nucleic Acids Res. 34, D363–D368 (2006).
Larkin, M.A. et al. Clustal W and Clustal X version 2.0. Bioinformatics 23, 2947–2948 (2007).
Ng, P. et al. Gene identification signature (GIS) analysis for transcriptome characterization and genome annotation. Nat. Methods 2, 105–111 (2005).
Mortazavi, A., Williams, B.A., McCue, K., Schaeffer, L. & Wold, B. Mapping and quantifying mammalian transcriptomes by RNA-Seq. Nat. Methods 5, 621–628 (2008).
Wang, L., Feng, Z., Wang, X. & Zhang, X. DEGseq: an R package for identifying differentially expressed genes from RNA-seq data. Bioinformatics 26, 136–138 (2010).
Acknowledgements
We thank Z. Ning of the Wellcome Trust Sanger Institute for assisting in genome assembly. The Shanghai Supercomputer Center and the Fudan University High-End Computing Center kindly provided computation facilities for aspects of the data analysis. This work was funded by grants from the National Basic Research Program (973) of China (2010CB534906 and 2011CB111610), the National Natural Science Foundation of China (30760185, 81271868 and 31071158), the Shanghai Municipal Commission for Science and Technology (10XD1403200 and 11DZ2292600), the National High-Tech R&D Program (863) of China (2012AA020409) and the Program for Changjiang Scholars and Innovative Teams in Universities in China, the Ministry of Education (IRT1181). Support from the National Health and Medical Research Council (NHMRC) of Australia is also gratefully acknowledged. D.P.M. is an NHMRC of Australia Senior Principal Research Fellow and a Senior Scientist at the Queensland Institute of Medical Research.
Author information
Authors and Affiliations
Contributions
S.W. and W.Z. coordinated the project. W.Z., Z.Z., L. Zhao, B.S. and R.L. collected the E. granulosus samples and extracted DNA and RNA. S.W., H.Z., Y. Zhu, H.K., J.C., L.W., A.C., S.Y., Z.G., L.J. and W.G. directed and performed the genome and transcriptome sequencing. S.W., Y. Zhou, G.L., Y.H., H.Z. and Y. Zhu assembled the genome and EST sequence data. Y. Zhou, W.Z., H.Z., L. Zhang, Y.W., Y.H., J. Li, L. Zhao, J. Liu, Z.W., B.B., T.V. and S.W. performed gene prediction, annotation and genomic analysis. Z.C., G.Z., D.P.M., H.W. and R.L. commented on the genome sequencing and analysis. W.Z., Y. Zhou, H.Z., L. Zhang, M.K.J., D.P.M., Z.C. and S.W. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–12, Supplementary Tables 1–3, 5–12, 14–26, 30 and 32–48, and Supplementary Note (PDF 5300 kb)
Supplementary Table 4
Coverage statistics of repetitive contigs (XLSX 723 kb)
Supplementary Table 13
Distribution of genes in KEGG orthologs (XLSX 292 kb)
Supplementary Table 27
Significantly regulated genes in adult worms (Adult), oncospheres (Onc), protoscoleces (PSC) and hydatid cyst membrane (Cyst) of E. granulosus (XLSX 95 kb)
Supplementary Table 28
Profiles of transcription and annotation of genes from adult, oncosphere (Onc), protoscolex (PSC) and hydatid cyst membrane (Cyst) of E. granulosus (XLSX 3404 kb)
Supplementary Table 29
Genes up- or down-regulated in adult worms (Adult), oncospheres (Onc), protoscoleces (PSC) and hydatid cyst membrane (Cyst) of E. granulosus (XLSX 124 kb)
Supplementary Table 31
Genes up- or down-regulated in hydatid cyst membrane (Cyst) and adult worms (Adult) of E.granulosus (XLSX 210 kb)
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/.
About this article
Cite this article
Zheng, H., Zhang, W., Zhang, L. et al. The genome of the hydatid tapeworm Echinococcus granulosus. Nat Genet 45, 1168–1175 (2013). https://doi.org/10.1038/ng.2757
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.2757
This article is cited by
-
Integrated transcriptomic and proteomic analyses of plerocercoid and adult Spirometra mansoni reveal potential important pathways in the development of the medical tapeworm
Parasites & Vectors (2023)
-
Characterization of protein cargo of Echinococcus granulosus extracellular vesicles in drug response and its influence on immune response
Parasites & Vectors (2023)
-
Dimerization, host-parasite communication and expression studies of an Echinococcus granulosus 2DBD nuclear receptor
Parasitology Research (2023)
-
Bioinformatics analysis and experimental verification of Notch signalling pathway-related miRNA–mRNA subnetwork in extracellular vesicles during Echinococcus granulosus encystation
Parasites & Vectors (2022)
-
Proteomic profiling of hydatid fluid from pulmonary cystic echinococcosis
Parasites & Vectors (2022)