Abstract
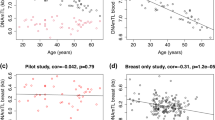
TERT-locus SNPs and leukocyte telomere measures are reportedly associated with risks of multiple cancers. Using the Illumina custom genotyping array iCOGs, we analyzed ∼480 SNPs at the TERT locus in breast (n = 103,991), ovarian (n = 39,774) and BRCA1 mutation carrier (n = 11,705) cancer cases and controls. Leukocyte telomere measurements were also available for 53,724 participants. Most associations cluster into three independent peaks. The minor allele at the peak 1 SNP rs2736108 associates with longer telomeres (P = 5.8 × 10−7), lower risks for estrogen receptor (ER)-negative (P = 1.0 × 10−8) and BRCA1 mutation carrier (P = 1.1 × 10−5) breast cancers and altered promoter assay signal. The minor allele at the peak 2 SNP rs7705526 associates with longer telomeres (P = 2.3 × 10−14), higher risk of low-malignant-potential ovarian cancer (P = 1.3 × 10−15) and greater promoter activity. The minor alleles at the peak 3 SNPs rs10069690 and rs2242652 increase ER-negative (P = 1.2 × 10−12) and BRCA1 mutation carrier (P = 1.6 × 10−14) breast and invasive ovarian (P = 1.3 × 10−11) cancer risks but not via altered telomere length. The cancer risk alleles of rs2242652 and rs10069690, respectively, increase silencing and generate a truncated TERT splice variant.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
McEachern, M.J., Krauskopf, A. & Blackburn, E.H. Telomeres and their control. Annu. Rev. Genet. 34, 331–358 (2000).
Palm, W. & de Lange, T. How shelterin protects mammalian telomeres. Annu. Rev. Genet. 42, 301–334 (2008).
Baird, D.M. Telomeres. Exp. Gerontol. 41, 1223–1227 (2006).
Moyzis, R.K. et al. A highly conserved repetitive DNA sequence, (TTAGGG)n, present at the telomeres of human chromosomes. Proc. Natl. Acad. Sci. USA 85, 6622–6626 (1988).
Allsopp, R.C. et al. Telomere length predicts replicative capacity of human fibroblasts. Proc. Natl. Acad. Sci. USA 89, 10114–10118 (1992).
Harley, C.B. Telomere loss: mitotic clock or genetic time bomb? Mutat. Res. 256, 271–282 (1991).
Levy, M.Z. et al. Telomere end-replication problem and cell aging. J. Mol. Biol. 225, 951–960 (1992).
Counter, C.M. et al. Telomere shortening associated with chromosome instability is arrested in immortal cells which express telomerase activity. EMBO J. 11, 1921–1929 (1992).
Hiyama, E. et al. Telomerase activity in human breast tumors. J. Natl. Cancer Inst. 88, 116–122 (1996).
Stampfer, M.R. & Yaswen, P. Human epithelial cell immortalization as a step in carcinogenesis. Cancer Lett. 194, 199–208 (2003).
Alter, B.P. et al. Telomere length is associated with disease severity and declines with age in dyskeratosis congenita. Haematologica 97, 353–359 (2012).
Njajou, O.T. et al. Telomere length is paternally inherited and is associated with parental lifespan. Proc. Natl. Acad. Sci. USA 104, 12135–12139 (2007).
Slagboom, P.E., Droog, S. & Boomsma, D.I. Genetic determination of telomere size in humans: a twin study of three age groups. Am. J. Hum. Genet. 55, 876–882 (1994).
Mirabello, L. et al. The association of telomere length and genetic variation in telomere biology genes. Hum. Mutat. 31, 1050–1058 (2010).
Pooley, K.A. et al. No association between TERT-CLPTM1L single nucleotide polymorphism rs401681 and mean telomere length or cancer risk. Cancer Epidemiol. Biomarkers Prev. 19, 1862–1865 (2010).
Mocellin, S. et al. Telomerase reverse transcriptase locus polymorphisms and cancer risk: a field synopsis and meta-analysis. J. Natl. Cancer Inst. 104, 840–854 (2012).
Soerensen, M. et al. Genetic variation in TERT and TERC and human leukocyte telomere length and longevity: a cross-sectional and longitudinal analysis. Aging Cell 11, 223–227 (2012).
McGrath, M. et al. Telomere length, cigarette smoking, and bladder cancer risk in men and women. Cancer Epidemiol. Biomarkers Prev. 16, 815–819 (2007).
Pooley, K.A. et al. Telomere length in prospective and retrospective cancer case-control studies. Cancer Res. 70, 3170–3176 (2010).
Shen, J. et al. Telomere length, oxidative damage, antioxidants and breast cancer risk. Int. J. Cancer 124, 1637–1643 (2009).
Wentzensen, I.M. et al. The association of telomere length and cancer: a meta-analysis. Cancer Epidemiol. Biomarkers Prev. 20, 1238–1250 (2011).
De Vivo, I. et al. A prospective study of relative telomere length and postmenopausal breast cancer risk. Cancer Epidemiol. Biomarkers Prev. 18, 1152–1156 (2009).
Zee, R.Y. et al. Mean telomere length and risk of incident colorectal carcinoma: a prospective, nested case-control approach. Cancer Epidemiol. Biomarkers Prev. 18, 2280–2282 (2009).
Baird, D.M. Variation at the TERT locus and predisposition for cancer. Expert Rev. Mol. Med. 12, e16 (2010).
Beesley, J. et al. Functional polymorphisms in the TERT promoter are associated with risk of serous epithelial ovarian and breast cancers. PLoS ONE 6, e24987 (2011).
Kote-Jarai, Z. et al. Seven prostate cancer susceptibility loci identified by a multi-stage genome-wide association study. Nat. Genet. 43, 785–791 (2011).
Landi, M.T. et al. A genome-wide association study of lung cancer identifies a region of chromosome 5p15 associated with risk for adenocarcinoma. Am. J. Hum. Genet. 85, 679–691 (2009).
Rafnar, T. et al. Sequence variants at the TERT-CLPTM1L locus associate with many cancer types. Nat. Genet. 41, 221–227 (2009).
Shete, S. et al. Genome-wide association study identifies five susceptibility loci for glioma. Nat. Genet. 41, 899–904 (2009).
Stacey, S.N. et al. New common variants affecting susceptibility to basal cell carcinoma. Nat. Genet. 41, 909–914 (2009).
Turnbull, C. et al. Variants near DMRT1, TERT and ATF7IP are associated with testicular germ cell cancer. Nat. Genet. 42, 604–607 (2010).
Wang, Y. et al. Common 5p15.33 and 6p21.33 variants influence lung cancer risk. Nat. Genet. 40, 1407–1409 (2008).
Johnatty, S.E. et al. Evaluation of candidate stromal epithelial cross-talk genes identifies association between risk of serous ovarian cancer and TERT, a cancer susceptibility “hot-spot”. PLoS Genet. 6, e1001016 (2010).
Haiman, C.A. et al. A common variant at the TERT-CLPTM1L locus is associated with estrogen receptor–negative breast cancer. Nat. Genet. 43, 1210–1214 (2011).
1000 Genomes Project Consortium. A map of human genome variation from population-scale sequencing. Nature 467, 1061–1073 (2010).
Michailidou, K. et al. Large-scale genotyping identifies 41 new loci associated with breast cancer risk. Nat. Genet. published online; doi:10.1038/ng.2563 (27 March 2013).
Pharoah, P.D.P. et al. GWAS meta-analysis and replication identifies three new common susceptibility loci for ovarian cancer. Nat. Genet. published online; doi:10.1038/ng.2564 (27 March 2013).
Garcia-Closas, M. et al. Genome-wide association studies identify four ER negative–specific breast cancer risk loci. Nat. Genet. published online; doi:10.1038/ng.2561 (27 March 2013).
Rosenbloom, K.R. et al. ENCODE whole-genome data in the UCSC Genome Browser: update 2012. Nucleic Acids Res. 40, D912–D917 (2012).
Giresi, P.G. et al. FAIRE (Formaldehyde-Assisted Isolation of Regulatory Elements) isolates active regulatory elements from human chromatin. Genome Res. 17, 877–885 (2007).
Taylor, J. et al. ESPERR: learning strong and weak signals in genomic sequence alignments to identify functional elements. Genome Res. 16, 1596–1604 (2006).
Lim, E. et al. Aberrant luminal progenitors as the candidate target population for basal tumor development in BRCA1 mutation carriers. Nat. Med. 15, 907–913 (2009).
Chen, Y.J. et al. PAX8 regulates telomerase reverse transcriptase and telomerase RNA component in glioma. Cancer Res. 68, 5724–5732 (2008).
Colgin, L.M. et al. The hTERTα splice variant is a dominant negative inhibitor of telomerase activity. Neoplasia 2, 426–432 (2000).
Saebøe-Larssen, S., Fossberg, E. & Gaudernack, G. Characterization of novel alternative splicing sites in human telomerase reverse transcriptase (hTERT): analysis of expression and mutual correlation in mRNA isoforms from normal and tumour tissues. BMC Mol. Biol. 7, 26 (2006).
Kilian, A. et al. Isolation of a candidate human telomerase catalytic subunit gene, which reveals complex splicing patterns in different cell types. Hum. Mol. Genet. 6, 2011–2019 (1997).
Wick, M., Zubov, D. & Hagen, G. Genomic organization and promoter characterization of the gene encoding the human telomerase reverse transcriptase (hTERT). Gene 232, 97–106 (1999).
Desmet, F.O. et al. Human Splicing Finder: an online bioinformatics tool to predict splicing signals. Nucleic Acids Res. 37, e67 (2009).
Cancer Genome Atlas Research Network. Integrated genomic analyses of ovarian carcinoma. Nature 474, 609–615 (2011).
Weischer, M. et al. Short telomere length, myocardial infarction, ischemic heart disease, and early death. Arterioscler. Thromb. Vasc. Biol. 32, 822–829 (2012).
Codd, V. et al. Common variants near TERC are associated with mean telomere length. Nat. Genet. 42, 197–199 (2010).
Levy, D. et al. Genome-wide association identifies OBFC1 as a locus involved in human leukocyte telomere biology. Proc. Natl. Acad. Sci. USA 107, 9293–9298 (2010).
Willeit, P. et al. Telomere length and risk of incident cancer and cancer mortality. J. Am. Med. Assoc. 304, 69–75 (2010).
Kote-Jarai, Z. et al. Fine-mapping identifies multiple prostate cancer risk loci on 5p15, one of which associates with TERT expression. Hum. Mol. Genet. published online; doi:10.1093/hmg/ddt086 (27 March 2013).
Krtolica, A. et al. Senescent fibroblasts promote epithelial cell growth and tumorigenesis: a link between cancer and aging. Proc. Natl. Acad. Sci. USA 98, 12072–12077 (2001).
Lawrenson, K. et al. Senescent fibroblasts promote neoplastic transformation of partially transformed ovarian epithelial cells in a three-dimensional model of early stage ovarian cancer. Neoplasia 12, 317–325 (2010).
McKay, J.D. et al. Lung cancer susceptibility locus at 5p15.33. Nat. Genet. 40, 1404–1406 (2008).
Shen, J. et al. Multiple genetic variants in telomere pathway genes and breast cancer risk. Cancer Epidemiol. Biomarkers Prev. 19, 219–228 (2010).
Hu, Z. et al. A genome-wide association study identifies two new lung cancer susceptibility loci at 13q12.12 and 22q12.2 in Han Chinese. Nat. Genet. 43, 792–796 (2011).
Mushiroda, T. et al. A genome-wide association study identifies an association of a common variant in TERT with susceptibility to idiopathic pulmonary fibrosis. J. Med. Genet. 45, 654–656 (2008).
Truong, T. et al. Replication of lung cancer susceptibility loci at chromosomes 15q25, 5p15, and 6p21: a pooled analysis from the International Lung Cancer Consortium. J. Natl. Cancer Inst. 102, 959–971 (2010).
Zou, P. et al. The TERT rs2736100 polymorphism and cancer risk: a meta-analysis based on 25 case-control studies. BMC Cancer 12, 7 (2012).
Zhao, Y.M. et al. Fine-mapping of a region of chromosome 5p15.33 (TERT-CLPTM1L) suggests a novel locus in TERT and a CLPTM1L haplotype are associated with glioma susceptibility in a Chinese population. Int. J. Cancer 129, 2463–2472 (2011).
Takubo, K. et al. Telomere lengths are characteristic in each human individual. Exp. Gerontol. 37, 523–531 (2002).
Gadalla, S.M. et al. Telomere length in blood, buccal cells, and fibroblasts from patients with inherited bone marrow failure syndromes. Aging (Albany. NY) 2, 867–874 (2010).
Thomas, P., O' Callaghan, N.J. & Fenech, M. Telomere length in white blood cells, buccal cells and brain tissue and its variation with ageing and Alzheimer's disease. Mech. Ageing Dev. 129, 183–190 (2008).
Greider, C.W. & Blackburn, E.H. Identification of a specific telomere terminal transferase activity in Tetrahymena extracts. Cell 43, 405–413 (1985).
Choi, J. et al. TERT promotes epithelial proliferation through transcriptional control of a Myc- and Wnt-related developmental program. PLoS Genet. 4, e10 (2008).
Park, J.I. et al. Telomerase modulates Wnt signalling by association with target gene chromatin. Nature 460, 66–72 (2009).
Mukherjee, S. et al. Separation of telomerase functions by reverse genetics. Proc. Natl. Acad. Sci. USA 108, E1363–E1371 (2011).
Gaudet, M.M. et al. Identification of a BRCA2-specific modifier locus at 6p24 related to breast cancer risk. PLoS Genet. 9, e1003173 (2013).
Azzato, E.M. et al. Association between a germline OCA2 polymorphism at chromosome 15q13.1 and estrogen receptor–negative breast cancer survival. J. Natl. Cancer Inst. 102, 650–662 (2010).
Bojesen, S.E., Tybjaerg-Hansen, A. & Nordestgaard, B.G. Integrin β3 Leu33Pro homozygosity and risk of cancer. J. Natl. Cancer Inst. 95, 1150–1157 (2003).
Allin, K.H., Bojesen, S.E. & Nordestgaard, B.G. Baseline C-reactive protein is associated with incident cancer and survival in patients with cancer. J. Clin. Oncol. 27, 2217–2224 (2009).
Allin, K.H. et al. C-reactive protein and the risk of cancer: a mendelian randomization study. J. Natl. Cancer Inst. 102, 202–206 (2010).
Zacho, J. et al. Genetically elevated C-reactive protein and ischemic vascular disease. N. Engl. J. Med. 359, 1897–1908 (2008).
Sankararaman, S. et al. Estimating local ancestry in admixed populations. Am. J. Hum. Genet. 82, 290–303 (2008).
Terry, K.L. et al. Telomere length and genetic variation in telomere maintenance genes in relation to ovarian cancer risk. Cancer Epidemiol. Biomarkers Prev. 21, 504–512 (2012).
Cawthon, R.M. Telomere measurement by quantitative PCR. Nucleic Acids Res. 30, e47 (2002).
Cawthon, R.M. Telomere length measurement by a novel monochrome multiplex quantitative PCR method. Nucleic Acids Res. 37, e21 (2009).
Barnes, D.R. et al. Evaluation of association methods for analysing modifiers of disease risk in carriers of high-risk mutations. Genet. Epidemiol. 36, 274–291 (2012).
Antoniou, A.C. et al. A locus on 19p13 modifies risk of breast cancer in BRCA1 mutation carriers and is associated with hormone receptor–negative breast cancer in the general population. Nat. Genet. 42, 885–892 (2010).
Antoniou, A.C. et al. A weighted cohort approach for analysing factors modifying disease risks in carriers of high-risk susceptibility genes. Genet. Epidemiol. 29, 1–11 (2005).
Eirew, P. et al. A method for quantifying normal human mammary epithelial stem cells with in vivo regenerative ability. Nat. Med. 14, 1384–1389 (2008).
Acknowledgements
We thank all the individuals who took part in these studies and all the researchers, clinicians, technicians and administrative staff who have enabled this work to be carried out. COGS is funded through a grant from the European Commission's Seventh Framework Programme (agreement 223175–HEALTH-F2-2009-223175). BCAC is funded by Cancer Research UK (C1287/A10118 and C1287/A12014). BCAC meetings have been funded by the European Union Cooperation in Science and Technology (COST) programme (BM0606). Telomere length measurement and analysis were funded by Cancer Research UK project grant C1287/A9540 and Chief Physician Johan Boserup and Lise Boserup's Fund. CIMBA data management and analysis were supported by Cancer Research UK grants C12292/A11174 and C1287/A10118. OCAC is supported by a grant from the Ovarian Cancer Research Fund thanks to the family and friends of Kathryn Sladek Smith (PPD/RPCI.07). Genotyping of the iCOGS array was funded by the European Union (HEALTH-F2-2009-223175), Cancer Research UK (C1287/A10710), the Canadian Institutes of Health Research (CIHR) for the CIHR Team in Familial Risks of Breast Cancer program (J.S. and D.E.) and the Ministry of Economic Development, Innovation and Export Trade of Quebec (grant PSR-SIIRI-701; J.S., D.E. and P. Hall). Scientific development and funding of the OCAC portion of this project were supported by Genetic Associations and Mechanisms in Oncology (GAME-ON; U19-CA148112). CIMBA genotyping was supported by US National Institutes of Health (NIH) grant CA128978, a National Cancer Institute (NCI) Specialized Program of Research Excellence (SPORE) in Breast Cancer (CA116201), a US Department of Defense Ovarian Cancer Idea award (W81XWH-10-1-0341) and grants from the Breast Cancer Research Foundation and the Komen Foundation for the Cure. This study made use of data generated by The Wellcome Trust Case Control Consortium (funding was provided by Wellcome Trust award 076113) and the TCGA Pilot Project established by NCI and the National Human Genome Research Institute.
Author information
Authors and Affiliations
Consortia
Contributions
Manuscript writing group: S.E.B., K.A.P., S.E.J., J. Beesley, K. Michailidou, S.L.E., H.A.P., P.L.M., M.H.G., H.C.S., K.L., A.N.A.M., S.A.G., A. Berchuck, P.D.P.P., E.L.G., R.R.R., D.F.E., A.C.A., G.C.-T. and A.M.D. Locus SNP selection: S.E.B., K.A.P., M.H.G., E. Dicks and A.M.D. Preparation of OCC samples for genotyping: S.J.R. and C.L.P. iCOGS genotyping, calling and quality control: S.E.B., S.F.N., A.G.-N., M.A. Rossing, J. Beesley, D.C.T., D.V., F.B., A. Swerdlow, M.J.M., C.L., C. Baynes, J.M. Cunningham, J.A.D., A.M.D., G.C.-T., K.A.P., D.F.E., P. Soucy and J.S. Imputation: K. Michailidou, K.B.K., J.P.T., A.C.A. and D.F.E. Telomere length determination and analysis: S.E.B., K.A.P., M.W., A.M.D. and D.F.E. Statistical analyses and programming: K. Michailidou, K.B.K., S.E.B., K.A.P., A.C.A., D.F.E., S.E.J. and Y. Lu. Functional analysis and bioinformatics: S.L.E., J.D.F., K.M.H., H.A.P., R.R.R., H.C.S., K.L., S.A.G., A.N.A.M., B.L.F., E.L.G., S.J.R., M.C.L., J. Beesley, M.D.S., K.L., C.E.S., R.L.J., S.R.L. and G.C.-T. COGS coordination: P. Hall, D.F.E., J. Benitez and A.M.D. BCAC coordination: D.F.E., G.C.-T. and P.D.P.P. BCAC data management: M.K.B. and Q.W. CIMBA coordination: A.C.A., G.C.-T. and F.J.C. OCAC coordination: P.D.P.P., S.J.R. and C.M.P. CIMBA data management: L.M. and D.B. Provided participant samples and phenotype information and read and approved the manuscript: S.E.B., K.A.P., S.E.J., J. Beesley, K. Michailidou, J.P.T., S.L.E., H.A.P., H.C.S., C.E.S., K.M.H., P.L.M., K.L., M.D.S., Y. Lu, R.K., N. Woods, R.L.J., J.D.F., X.C., M.W., S.F.N., M.J.M., M. Ghoussaini, S.A., C. Baynes, M.K.B., Q.W., J.D., L.M., D.B., A. Lee, S. Healey, M.L., D.C.T., D.V., F.B., I.V., S.L., E. Despierre, H.A.R., A.G.-N., M.A. Rossing, G.P., J.A.D., N. Álvarez, M.C.L., B.L.F., N. Schoof, J.C.-C., M.S.C., J. Peto, K.R.K., A. Broeks, S.M.A., M.K.S., L.M.B., B. Winterhoff, H.N., G.E.K., D.L., L.R., P.G., A.T., R.L.M., J.J.G., A.C., V.S., B. Burwinkel, F.M., R.H., E.J.S., C.A.H., S.W.-G., I.L.A., K.B.M., J.L.H., K. Odunsi, A. Lindblom, G.G.G., H. Brenner, J.S., G.L., P.A.F., M.E.C., P.R., L.R.W., A. Swerdlow, M.T.G., H. Brauch, M.G.-C., P. Hillemanns, R.W., M. Dürst, P.D., I.R., A. Jakubowska, J. Lubinski, A. Mannermaa, R. Butzow, N.V.B., T.D., L.M.P., W.Z., A. Leminen, H.A.-C., C.H.B., V. Kristensen, R.B.N., K. Muir, R.E., A. Meindl, F.H., K. Matsuo, A.d.B., A.H.W., P. Harter, S.-H.T., I.S., X.-O.S., W.B., S. Hosono, D.K., T.N., M. Hartman, Y.Y., U.H., B.Y.K., S. Sangrajrang, S.K.K., V.G., A. Jensen, D.E., E.H., C.-Y.S., J. Brown, Y.L.W., M. Shah, M.A.N.A., R.L., S.Z.O., K.C., R.A.V., B.G.N., H.F., C.V., J.E.O., X.W., D.A.L., A.R., R.P.W., D.F.-J., E.I., S.N., J.M.S., I.D.S.S., D.W.C., L.G., K.L.T., O.F., A.F.V., C.E.v.d.S., E.M.P., F.B.L.H., S.S.T., J. Liu, E.V.B., J. Li, S.H.O., K.H., I.O., C. Blomqvist, L.R.-R., K.A., H.B.S., T.A.M., E. Wik, B. Brouwers, C.K., E. Wauters, M.K.H., H.W., L.A.K., C.M., K.K.A., P.L.-P., A.M.v.A., T.T., L.F.A.G.M., J. Benitez, T.P., J.I.A.P., M. Hoatlin, M.P.Z., L.S.C., S.P.B., L.E.K., A. Schneeweiss, N.D.L., C. Sohn, A.B.-W., I.T., M.J. Kerin, N.M., C.C., B.E.H., J. Menkiszak, F.S., N. Wentzensen, L.L.M., H.P.Y., A.M.M., G. Glendon, S.A.E., J.A.K., C.K.H., C.A., M. Gore, H.T., H.S., M.C.S., A. Jager, A.M.W.v.d.O., R. Brown, J.W.M.M., J.M.F., M.K., J. Paul, S. Margolin, N. Siddiqui, G.S., A.S.W., L. Baglietto, V.M., C. Stegmaier, W.S., H. Müller, V.A., F.L., Y.-T.G., M.S.G., G.Y., M. Dumont, J.R.M., A. Hartmann, A.B.E., M.W.B., C.M.P., M.P.L., J.P.-W., B.P., T.A.S., F.F., M. Barile, A.Z., A.A., A.G.-M., M.J., S.J.R., N.O., U.M., C.L.P., T. Brüning, M.C.P., Y.-D.K., J. Lissowska, J.F., J.K., S.J.C., A.D.-M., A.J.-V., I.K.R., K.P., M. Bidzinski, S.K., A. Hollestelle, C. Seynaeve, R.A.E.M.T., K.D., K.J., J.M.H., V.-M.K., V. Kataja, N.N.A., J. Long, M. Shrubsole, S.D.-H., A. Lophatananon, P. Siriwanarangsan, S.S.-B., N.D., P.L., R.K.S., H. Ito, H. Iwata, K.T., C.-C.T., D.O.S., D.v.d.B., C.H.Y., M.K.I., Y.-C.T., H.C., W.L., L.B.S., Q.C., D.-Y.N., K.-Y.Y., H. Miao, P.T.-C.I., Y.Y.T., J. McKay, C. Shapiro, F.A., G.F., C.-N.H., J.-C.Y., M.-F.H., C.S.H., C.L., S.P., D.S.-L., P.P., T.R.R., M.P., C.F.S., E. Friedman, M.T., K. Offit, T.V.O.H., S.L.N., C.I.S., I.B., J. Garber, S.A.N., J.N.W., M.M., E.O., A.K.G., D.Y., D.E.G., T. Caldes, E.N.I., L.T., B.K.A., I.C., A.R.M., C.J.v.A., K.E.P.v.R., H.M.-H., J.M. Collée, J.C.O., M.J.H., M.A. Rookus, R.B.v.d.L., T.A.M.v.O., D.G.E., D.F., E. Fineberg, J. Barwell, L.W., M.J. Kennedy, R.P., R. Davidson, S.D.E., T. Cole, B.B.-d.P., B. Buecher, F.D., L. Faivre, M.F., O.M.S., O.C., S.G., S. Mazoyer, V.B., V.C.-M., A.T.-G., J. Gronwald, T. Byrski, A.B.S., B. Bonanni, D.Z., G. Giannini, L. Bernard, R. Dolcetti, S. Manoukian, N. Arnold, C.E., H.D., K.R., D.N., H.P., C. Sutter, B. Wappenschmidt, Å. Borg, B.M., J.R., M. Soller, K.L.N., S.M.D., G.C.R., R.S., D.G.K., M.-K.T., S.S.P., Y. Laitman, A.-B.S., T.A.K., U.B.J., M.R., A.-M.G., B.E., L. Foretova, S.A.S., J. Lester, P. Soucy, K.B.K., C.O., J.M. Cunningham, S. Slager, V.S.P., E. Dicks, S.R.L., F.J.C., P. Hall, A.N.A.M., S.A.G., P.D.P.P., R.R.R., E.L.G., M.H.G., D.F.E., A. Berchuck, A.C.A., G.C.-T. and A.M.D.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
A list of members is provided in the Supplementary Note.
A list of members is provided in the Supplementary Note.
A list of members is provided in the Supplementary Note.
A list of members is provided in the Supplementary Note.
A list of members is provided in the Supplementary Note.
A list of members is provided in the Supplementary Note.
A list of members is provided in the Supplementary Note.
A list of members is provided in the Supplementary Note.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7, Supplementary Tables 1–11 and Supplementary Note (PDF 2042 kb)
Rights and permissions
About this article
Cite this article
Bojesen, S., Pooley, K., Johnatty, S. et al. Multiple independent variants at the TERT locus are associated with telomere length and risks of breast and ovarian cancer. Nat Genet 45, 371–384 (2013). https://doi.org/10.1038/ng.2566
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.2566
This article is cited by
-
Combining old and new concepts in targeting telomerase for cancer therapy: transient, immediate, complete and combinatory attack (TICCA)
Cancer Cell International (2023)
-
Telomere length and hTERT genetic variants as potential prognostic markers in multiple myeloma
Scientific Reports (2023)
-
Genetics of human telomere biology disorders
Nature Reviews Genetics (2023)
-
DNA methylation QTL mapping across diverse human tissues provides molecular links between genetic variation and complex traits
Nature Genetics (2023)
-
Identification of h-TERT Promoter Mutations in Germline DNA from North Indian Lung Carcinoma Patients
Indian Journal of Clinical Biochemistry (2023)