Abstract
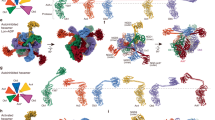
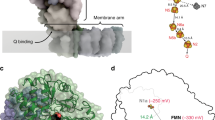
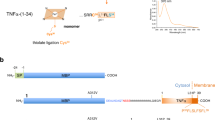
The Lon AAA+ protease degrades damaged or misfolded proteins in its intramolecular chamber. Its activity must be precisely controlled, but the mechanism by which Lon is regulated in response to different environments is not known. Facultative anaerobes in the Enterobacteriaceae family, mostly symbionts and pathogens, encounter both anaerobic and aerobic environments inside and outside the host′s body, respectively. The bacteria characteristically have two cysteine residues on the Lon protease (P) domain surface that unusually form a disulfide bond. Here we show that the cysteine residues act as a redox switch of Lon. Upon disulfide bond reduction, the exit pore of the P-domain ring narrows by ∼30%, thus interrupting product passage and decreasing activity by 80%; disulfide bonding by oxidation restores the pore size and activity. The redox switch (E°′ = −227 mV) is appropriately tuned to respond to variation between anaerobic and aerobic conditions, thus optimizing the cellular proteolysis level for each environment.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Goldberg, A.L., Moerschell, R.P., Chung, C.H. & Maurizi, M.R. ATP-dependent protease La (Lon) from Escherichia coli. Methods Enzymol. 244, 350–375 (1994).
Van Melderen, L. & Aertsen, A. Regulation and quality control by Lon-dependent proteolysis. Res. Microbiol. 160, 645–651 (2009).
Sauer, R.T. & Baker, T.A. AAA+ proteases: ATP-fueled machines of protein destruction. Annu. Rev. Biochem. 80, 587–612 (2011).
Waxman, L. & Goldberg, A.L. Selectivity of intracellular proteolysis: protein substrates activate the ATP-dependent protease (La). Science 232, 500–503 (1986).
Goff, S.A. & Goldberg, A.L. An increased content of protease La, the lon gene product, increases protein degradation and blocks growth in Escherichia coli. J. Biol. Chem. 262, 4508–4515 (1987).
Christensen, S.K. et al. Overproduction of the Lon protease triggers inhibition of translation in Escherichia coli: involvement of the yefM-yoeB toxin-antitoxin system. Mol. Microbiol. 51, 1705–1717 (2004).
Phillips, T.A., VanBogelen, R.A. & Neidhardt, F.C. lon gene product of Escherichia coli is a heat-shock protein. J. Bacteriol. 159, 283–287 (1984).
Fukuda, R. et al. HIF-1 regulates cytochrome oxidase subunits to optimize efficiency of respiration in hypoxic cells. Cell 129, 111–122 (2007).
Rotanova, T.V. et al. Slicing a protease: structural features of the ATP-dependent Lon proteases gleaned from investigations of isolated domains. Protein Sci. 15, 1815–1828 (2006).
Cha, S.-S. Crystal structure of Lon protease: molecular architecture of gated entry to a sequestered degradation chamber. EMBO J. 29, 3520–3530 (2010).
Maurizi, M.R. Degradation in vitro of bacteriophage λN protein by Lon protease from Escherichia coli. J. Biol. Chem. 262, 2696–2703 (1987).
Van Melderen, L. et al. ATP-dependent degradation of CcdA by Lon protease. Effects of secondary structure and heterologous subunit interactions. J. Biol. Chem. 271, 27730–27738 (1996).
Nishii, W., Maruyama, T., Matsuoka, R., Muramatsu, T. & Takahashi, K. The unique sites in SulA protein preferentially cleaved by ATP-dependent Lon protease from Escherichia coli. Eur. J. Biochem. 269, 451–457 (2002).
Nishii, W. et al. Cleavage mechanism of ATP-dependent Lon protease toward ribosomal S2 protein. FEBS Lett. 579, 6846–6850 (2005).
Botos, I. et al. Crystal structure of the AAA+ α domain of E. coli Lon protease at 1.9 Å resolution. J. Struct. Biol. 146, 113–122 (2004).
Botos, I. et al. The catalytic domain of Escherichia coli Lon protease has a unique fold and a Ser-Lys dyad in the active site. J. Biol. Chem. 279, 8140–8148 (2004).
García-Nafría, J. et al. Structure of the catalytic domain of the human mitochondrial Lon protease: proposed relation of oligomer formation and activity. Protein Sci. 19, 987–999 (2010).
Li, M. et al. Crystal structure of the N-terminal domain of E. coli Lon protease. Protein Sci. 14, 2895–2900 (2005).
Gilbert, H.F. Molecular and cellular aspects of thiol-disulfide exchange. Adv. Enzymol. 63, 69–172 (1990).
Waxman, L. & Goldberg, A.L. Protease La, the lon gene product, cleaves specific fluorogenic peptides in an ATP-dependent reaction. J. Biol. Chem. 260, 12022–12028 (1985).
Kim, S.O. et al. OxyR: a molecular code for redox-related signaling. Cell 109, 383–396 (2002).
Goldberg, A.L. & Waxman, L. The role of ATP hydrolysis in the breakdown of proteins and peptides by protease La from Escherichia coli. J. Biol. Chem. 260, 12029–12034 (1985).
Thomas-Wohlever, J. & Lee, I. Kinetic characterization of the peptidase activity of Escherichia coli Lon reveals the mechanistic similarities in ATP-dependent hydrolysis of peptide and protein substrates. Biochemistry 41, 9418–9425 (2002).
Lee, I., Berdis, A.J. & Suzuki, C.K. Recent developments in the mechanistic enzymology of the ATP-dependent Lon protease from Escherichia coli: highlights from kinetic studies. Mol. Biosyst. 2, 477–483 (2006).
Fischer, H. & Glockshuber, R. ATP hydrolysis is not stoichiometrically linked with proteolysis in the ATP-dependent protease La from Escherichia coli. J. Biol. Chem. 268, 22502–22507 (1993).
Kroh, H.E. & Simon, L.D. Increased ATP-dependent proteolytic activity in lon-deficient Escherichia coli strains lacking the DnaK protein. J. Bacteriol. 173, 2691–2695 (1991).
Tomoyasu, T., Mogk, A., Langen, H., Goloubinoff, P. & Bukau, B. Genetic dissection of the roles of chaperones and proteases in protein folding and degradation in the Escherichia coli cytosol. Mol. Microbiol. 40, 397–413 (2001).
Howard-Flanders, P., Simson, E. & Theriot, L. A locus that controls filament formation and sensitivity to radiation in Escherichia coli K-12. Genetics 49, 237–246 (1964).
Wilson, M. Bacteriology of Humans: an Ecological Perspective (Wiley-Blackwell, 2008).
Aslund, F., Berndt, K.D. & Holmgren, A. Redox potentials of glutaredoxins and other thiol-disulfide oxidoreductases of the thioredoxin superfamily determined by direct protein-protein redox equilibria. J. Biol. Chem. 272, 30780–30786 (1997).
Zheng, M., Aslund, F. & Storz, G. Activation of the OxyR transcription factor by reversible disulfide bond formation. Science 279, 1718–1721 (1998).
Jakob, U., Muse, W., Eser, M. & Bardwell, J.C. Chaperone activity with a redox switch. Cell 96, 341–352 (1999).
Miller, R.A. & Britigan, B.E. Role of oxidants in microbial pathophysiology. Clin. Microbiol. Rev. 10, 1–18 (1997).
Choi, H. et al. Structural basis of the redox switch in the OxyR transcription factor. Cell 105, 103–113 (2001).
Green, J. & Paget, M.S. Bacterial redox sensors. Nat. Rev. Microbiol. 2, 954–966 (2004).
Ilbert, M. et al. The redox-switch domain of Hsp33 functions as dual stress sensor. Nat. Struct. Mol. Biol. 14, 556–563 (2007).
Antelmann, H. & Helmann, J.D. Thiol-based redox switches and gene regulation. Antioxid. Redox Signal. 14, 1049–1063 (2011).
Wedemeyer, W.J., Welker, E., Narayan, M. & Scheraga, H.A. Disulfide bonds and protein folding. Biochemistry 39, 4207–4216 (2000).
Schmidt, B., Ho, L. & Hogg, P.J. Allosteric disulfide bonds. Biochemistry 45, 7429–7433 (2006).
Suzuki, C.K. et al. ATP-dependent proteases that also chaperone protein biogenesis. Trends Biochem. Sci. 22, 118–123 (1997).
Harpaz, Y., Gerstein, M. & Chothia, C. Volume changes on protein folding. Structure 2, 641–649 (1994).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
Adams, P.D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Emsley, P., Lohkamp, B., Scott, W.G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D Biol. Crystallogr. 66, 486–501 (2010).
Vaguine, A.A., Richelle, J. & Wodak, S.J. SFCHECK: a unified set of procedures for evaluating the quality of macromolecular structure-factor data and their agreement with the atomic model. Acta Crystallogr. D Biol. Crystallogr. 55, 191–205 (1999).
Collaborative Computational Project, No. 4. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D Biol. Crystallogr. 50, 760–763 (1994).
DeLano, W.L. The PyMOL Molecular Graphics System. (2002).
Kornberg, A., Scott, J.F. & Bertsch, L.L. ATP utilization by rep protein in the catalytic separation of DNA strands at a replicating fork. J. Biol. Chem. 253, 3298–3304 (1978).
Frieden, C. Kinetic aspects of regulation of metabolic processes. The hysteretic enzyme concept. J. Biol. Chem. 245, 5788–5799 (1970).
Acknowledgements
We thank T. Nishii for assistance with the enzyme preparation and F. Amano (Osaka University of Pharmaceutical Science) for providing the E. coli AB1157 and JK405 strains. We would like to thank the beamline staff at BL26B1 and BL26B2 of SPring-8 for assistance during data collection. We wish to thank T. Nakagawa (UNISOKU, Co., Ltd.) for assistance with stopped-flow experiments. We thank A. Ishii, K. Ake, T. Imada and T. Nakayama for assistance with manuscript preparation. This work was supported in part by a Grant-in-Aid for Scientific Research (17770116) from the Ministry of Education, Culture, Sports, Science and Technology (MEXT), Japan (to W.N.). This work was also supported in part by the Targeted Proteins Research Program (TPRP) and the Platform for Drug Discovery, Informatics, and Structural Life Science, both from MEXT (to S.Y.).
Author information
Authors and Affiliations
Contributions
W.N. and S.Y. designed this study, interpreted the data and wrote the manuscript. W.N. performed biochemical and physiological experiments. M.K.-N. carried out the crystallographic study. T.T. and M.S. supported crystallization. T.M. contributed to plasmid construction. M.K. and H.K. contributed to stopped-flow analysis.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Results, Supplementary Figures 1–17 and Supplementary Tables 1–4. (PDF 11968 kb)
Supplementary Video
Redox switch of Lon. Structural changes in the exit pore of Lon between the oxidized and reduced forms are shown. (MOV 6554 kb)
Rights and permissions
About this article
Cite this article
Nishii, W., Kukimoto-Niino, M., Terada, T. et al. A redox switch shapes the Lon protease exit pore to facultatively regulate proteolysis. Nat Chem Biol 11, 46–51 (2015). https://doi.org/10.1038/nchembio.1688
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.1688